Abstract
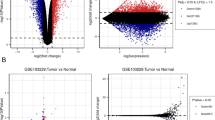
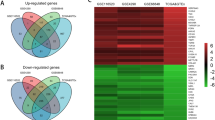
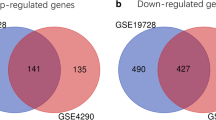
Glioblastoma (GBM) is one of the most common malignancies of the central nervous system, and the Isocitrate Dehydrogenase (IDH) mutation status of GBM has been recognized as a critical prognostic indicator. However, the molecular mechanism underlying the GBM with different IDH mutation status is still not unclear. In this study, a total of 353 DEGs including 207 up-regulated and 146 down-regulated were screened from multiple GBM data sets. Moreover, the biological processes and pathways enriched by DEGs were mainly associated with tumor progression, especially invasion and migration. Then, eight hub genes, including SDC4, SERPINE1, TNC, THBS1, COL1A1, CXCL8, TIMP1 and VEGFA, were selected from a PPI network. Finally, core genes, SERPINE1 and TIMP1, were identified from hub genes by survival analysis and sample validation. Overall, in this study, we revealed underlying molecular mechanisms in GBMs with different IDH mutation status and identified core genes that could be potential markers and targets for diagnosis and treatment of GBMs.







Similar content being viewed by others
References
Wen PY, Kesari S (2008) Malignant gliomas in adults. N Engl J Med 359:492–507. https://doi.org/10.1056/NEJMra0708126
Perry JR, Laperriere N, O’Callaghan CJ et al (2017) Short-course radiation plus temozolomide in elderly patients with glioblastoma. N Engl J Med 376:1027–1037. https://doi.org/10.1056/NEJMoa1611977
Stupp R, Mason WP, van den Bent MJ et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352:987–996. https://doi.org/10.1056/NEJMoa043330
Wechsler-Reya R, Scott MP (2001) The developmental biology of brain tumors. Annu Rev Neurosci 24:385–428. https://doi.org/10.1146/annurev.neuro.24.1.385
Huse JT, Holland E, DeAngelis LM (2013) Glioblastoma: molecular analysis and clinical implications. Annu Rev Med 64:59–70. https://doi.org/10.1146/annurev-med-100711-143028
Patel AP, Tirosh I, Trombetta JJ et al (2014) Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344:1396–1401. https://doi.org/10.1126/science.1254257
Sottoriva A, Spiteri I, Piccirillo SGM et al (2013) Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc Natl Acad Sci 110:4009–4014. https://doi.org/10.1073/pnas.1219747110
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820. https://doi.org/10.1007/s00401-016-1545-1
Eckel-Passow JE, Lachance DH, Molinaro AM et al (2015) Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N Engl J Med 372:2499–2508. https://doi.org/10.1056/NEJMoa1407279
Cohen AL, Holmen SL, Colman H (2013) IDH1 and IDH2 mutations in gliomas. Curr Neurol Neurosci Rep 13:345. https://doi.org/10.1007/s11910-013-0345-4
Turkalp Z, Karamchandani J, Das S (2014) IDH mutation in glioma. JAMA Neurol 71:1319. https://doi.org/10.1001/jamaneurol.2014.1205
Berghoff AS, Kiesel B, Widhalm G et al (2017) Correlation of immune phenotype with IDH mutation in diffuse glioma. Neuro Oncol 19:1460–1468. https://doi.org/10.1093/neuonc/nox054
Pan Y-B, Zhang C-H, Wang S-Q et al (2018) Transforming growth factor beta induced (TGFBI) is a potential signature gene for mesenchymal subtype high-grade glioma. J Neurooncol 137:395–407. https://doi.org/10.1007/s11060-017-2729-9
Han M-Z, Wang S, Zhao W-B et al (2019) Immune checkpoint molecule herpes virus entry mediator is overexpressed and associated with poor prognosis in human glioblastoma. EBioMedicine 43:159–170. https://doi.org/10.1016/j.ebiom.2019.04.002
Reon BJ, Anaya J, Zhang Y et al (2016) Expression of lncRNAs in low-grade gliomas and glioblastoma multiforme: an in silico analysis. PLOS Med 13:e1002192. https://doi.org/10.1371/journal.pmed.1002192
Wang Q, Hu B, Hu X et al (2017) Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32:42–56.e6. https://doi.org/10.1016/j.ccell.2017.06.003
Yao K, Wu B, Xi M et al (2015) Distant dissemination of mixed low-grade astroblastoma-arteriovenous malformation after initial operation: a case report. Int J Clin Exp Pathol 8:7450–7456
Kun Y, Duan Z, Mei X et al (2015) A rare case of malignant pediatric ectomesenchymoma arising from the cerebrum. Int J Clin Exp Pathol 8:8545–8550
Pickup MW, Mouw JK, Weaver VM (2014) The extracellular matrix modulates the hallmarks of cancer. EMBO Rep 15:1243–1253. https://doi.org/10.15252/embr.201439246
Fruman DA, Rommel C (2014) PI3K and cancer: lessons, challenges and opportunities. Nat Rev Drug Discov 13:140–156. https://doi.org/10.1038/nrd4204
Bray SJ (2006) Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol 7:678–689. https://doi.org/10.1038/nrm2009
Ohkawara B, Glinka A, Niehrs C (2011) Rspo3 binds syndecan 4 and induces Wnt/PCP signaling via clathrin-mediated endocytosis to promote morphogenesis. Dev Cell 20:303–314. https://doi.org/10.1016/j.devcel.2011.01.006
Chen LL, Gao GX, Shen FX et al (2018) SDC4 gene silencing favors human papillary thyroid carcinoma cell apoptosis and inhibits epithelial mesenchymal transition via wnt/β-catenin pathway. Mol Cells 41:853–867. https://doi.org/10.14348/molcells.2018.0103
Kosmopoulos M, Christofides A, Drekolias D et al (2018) Critical role of IL-8 targeting in gliomas. Curr Med Chem 25:1954–1967. https://doi.org/10.2174/0929867325666171129125712
Raychaudhuri B, Vogelbaum MA (2011) IL-8 is a mediator of NF-κB induced invasion by gliomas. J Neurooncol 101:227–235. https://doi.org/10.1007/s11060-010-0261-2
Brat DJ, Bellail AC, Van Meir EG (2005) The role of interleukin-8 and its receptors in gliomagenesis and tumoral angiogenesis. Neuro Oncol 7:122–133. https://doi.org/10.1215/S1152851704001061
Jiang Y, He J, Guo Y et al (2019) Identification of genes related to low-grade glioma progression and prognosis based on integrated transcriptome analysis. J Cell Biochem. https://doi.org/10.1002/jcb.29577
Hayashi M, Nomoto S, Hishida M et al (2014) Identification of the collagen type 1 alpha 1 gene (COL1A1) as a candidate survival-related factor associated with hepatocellular carcinoma. BMC Cancer 14:1–10. https://doi.org/10.1186/1471-2407-14-108
Wang Q, Yu J (2018) MiR-129-5p suppresses gastric cancer cell invasion and proliferation by inhibiting COL1A1. Biochem Cell Biol 96:19–25. https://doi.org/10.1139/bcb-2016-0254
Sun S, Wang Y, Wu Y et al (2018) Identification of COL1A1 as an invasion-related gene in malignant astrocytoma. Int J Oncol 53:2542–2554. https://doi.org/10.3892/ijo.2018.4568
Jones FS, Jones PL (2000) The tenascin family of ECM glycoproteins: structure, function, and regulation during embryonic development and tissue remodeling. Dev Dyn 218:235–259. https://doi.org/10.1002/(SICI)1097-0177(200006)218:2<235:AID-DVDY2>3.3.CO;2-7
Qi J, Esfahani DR, Huang T et al (2019) Tenascin-C expression contributes to pediatric brainstem glioma tumor phenotype and represents a novel biomarker of disease. Acta Neuropathol Commun 7:75. https://doi.org/10.1186/s40478-019-0727-1
Xia S, Lal B, Tung B et al (2016) Tumor microenvironment tenascin-C promotes glioblastoma invasion and negatively regulates tumor proliferation. Neuro Oncol 18:507–517. https://doi.org/10.1093/neuonc/nov171
Sarkar S, Mirzaei R, Zemp FJ et al (2017) Activation of NOTCH signaling by tenascin-C promotes growth of human brain tumor-initiating cells. Cancer Res 77:3231–3243. https://doi.org/10.1158/0008-5472.CAN-16-2171
Roberts DD (1996) Regulation of tumor growth and metastasis by thrombospondin-1. FASEB J 10:1183–1191. https://doi.org/10.1096/fasebj.10.10.8751720
Tuszynski GP, Nicosia RF (1996) The role of thrombospondin-1 in tumor progression and angiogenesis. BioEssays 18:71–76. https://doi.org/10.1002/bies.950180113
Seliger C, Leukel P, Moeckel S et al (2013) Lactate-modulated induction of THBS-1 activates transforming growth factor (TGF)-beta2 and migration of glioma cells in vitro. PLoS One 8:e78935. https://doi.org/10.1371/journal.pone.0078935
Liu Q, Liao F, Wu H et al (2016) Different expression of miR-29b and VEGFA in glioma. Artif Cells Nanomed Biotechnol 44:1927–1932. https://doi.org/10.3109/21691401.2015.1111237
Ye J, Zhu J, Chen H et al (2020) A novel lncRNA-LINC01116 regulates tumorigenesis of glioma by targeting VEGFA. Int J Cancer 146:248–261. https://doi.org/10.1002/ijc.32483
Hu Y, Li Y, Wu C et al (2017) MicroRNA-140-5p inhibits cell proliferation and invasion by regulating VEGFA/MMP2 signaling in glioma. Tumor Biol 39:101042831769755. https://doi.org/10.1177/1010428317697558
Lu P, Takai K, Weaver VM, Werb Z (2011) Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb Perspect Biol 3:a005058. https://doi.org/10.1101/cshperspect.a005058
Frantz C, Stewart KM, Weaver VM (2010) The extracellular matrix at a glance. J Cell Sci 123:4195–4200. https://doi.org/10.1242/jcs.023820
Ulisse S, Baldini E, Sorrenti S, D’Armiento M (2009) The urokinase plasminogen activator system: a target for anti-cancer therapy. Curr Cancer Drug Targets 9:32–71. https://doi.org/10.2174/156800909787314002
Harris L, Fritsche H, Mennel R et al (2007) American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol 25:5287–5312. https://doi.org/10.1200/JCO.2007.14.2364
Strojan P, Budihna M, Šmid L et al (1998) Urokinase-type plasminogen activator (uPA) and plasminogen activator inhibitor type 1 (PAI-1) in tissue and serum of head and neck squamous cell carcinoma patients. Eur J Cancer 34:1193–1197. https://doi.org/10.1016/S0959-8049(98)00029-X
Chin D, Boyle GM, Williams RM et al (2005) Novel markers for poor prognosis in head and neck cancer. Int J Cancer 113:789–797. https://doi.org/10.1002/ijc.20608
Wu D-M, Wang S, Wen X et al (2018) MircoRNA-1275 promotes proliferation, invasion and migration of glioma cells via SERPINE1. J Cell Mol Med 22:4963–4974. https://doi.org/10.1111/jcmm.13760
Seker F, Cingoz A, Sur-Erdem İ et al (2019) Identification of SERPINE1 as a regulator of glioblastoma cell dispersal with transcriptome profiling. Cancers (Basel) 11:1651. https://doi.org/10.3390/cancers11111651
Holten-Andersen M, Christensen IJ, Nilbert M et al (2004) Association between preoperative plasma levels of tissue inhibitor of metalloproteinases 1 and rectal cancer patient survival. Eur J Cancer 40:64–72. https://doi.org/10.1016/j.ejca.2003.09.019
Birgisson H, Nielsen HJ, Christensen IJ et al (2010) Preoperative plasma TIMP-1 is an independent prognostic indicator in patients with primary colorectal cancer: a prospective validation study. Eur J Cancer 46:3323–3331. https://doi.org/10.1016/j.ejca.2010.06.009
Cheng G, Fan X, Hao M et al (2016) Higher levels of TIMP-1 expression are associated with a poor prognosis in triple-negative breast cancer. Mol Cancer 15:30. https://doi.org/10.1186/s12943-016-0515-5
Schrohl A-S (2004) Tumor tissue levels of tissue inhibitor of metalloproteinase-1 as a prognostic marker in primary breast cancer. Clin Cancer Res 10:2289–2298. https://doi.org/10.1158/1078-0432.CCR-03-0360
Yoshikawa T, Cho H, Tsuburaya A, Kobayashi O (2009) Impact of plasma tissue inhibitor of metalloproteinase-1 on long-term survival in patients with gastric cancer. Gastric Cancer 12:31–36. https://doi.org/10.1007/s10120-008-0494-3
Rauvala M, Puistola U, Turpeenniemihujanen T (2005) Gelatinases and their tissue inhibitors in ovarian tumors; TIMP-1 is a predictive as well as a prognostic factor. Gynecol Oncol 99:656–663. https://doi.org/10.1016/j.ygyno.2005.07.009
Song G, Xu S, Zhang H et al (2016) TIMP1 is a prognostic marker for the progression and metastasis of colon cancer through FAK-PI3K/AKT and MAPK pathway. J Exp Clin Cancer Res 35:1–12. https://doi.org/10.1186/s13046-016-0427-7
Aaberg-Jessen C, Christensen K, Offenberg H et al (2009) Low expression of tissue inhibitor of metalloproteinases-1 (TIMP-1) in glioblastoma predicts longer patient survival. J Neurooncol 95:117–128. https://doi.org/10.1007/s11060-009-9910-8
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest exists in the submission of this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guo, Y., Wang, X., Ning, W. et al. Identification of two core genes in glioblastomas with different isocitrate dehydrogenase mutation status. Mol Biol Rep 47, 7477–7488 (2020). https://doi.org/10.1007/s11033-020-05804-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05804-w