Abstract
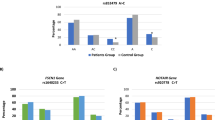
SMAD2 is a critical signal transducer molecule in the TGFβ- SMAD pathway which is also known for its tumor suppressor role. Genetic variations in SMAD2 render cells insensitive to its anti-proliferative signals leading to tumor formation. In this study, we demonstrate the impact of single nucleotide polymorphisms (SNPs) of SMAD2 (rs4940086 and rs8085335) on cervical cancer risk development in Bangladeshi population. 132 cervical cancer patients and 98 control volunteers were enrolled in the study and genotyped utilizing polymerase chain reaction–restriction fragment length polymorphism (PCR–RFLP) method. The association between cervical cancer susceptibility and the chosen SNPs were evaluated through multiple logistic regression. SMAD2 rs4940086 heterozygous genotype (T/C) was associated with a 3.89 times higher risk of cervical cancer development (P = 0.001, AOR 3.89, 95% CI 1.777–8.513). The T/C and C/C genotypes in combination also significantly elevated cervical cancer risk (P = 0.035, AOR 1.876, 95% CI 1.047–3.364). Urban cancer patients had a significantly higher chance of carrying the rs4940086 polymorphism as compared to rural cancer patients (P = 0.045, OR 2.59 95% CI 1.02–6.59). SMAD2 rs8085335 heterozygous variant (A/G) demonstrated modest effects in increasing cervical cancer susceptibility (P = 0.594, AOR 1.247, 95% CI 0.554–2.809). Our results suggest that polymorphic variations in SMAD2, particularly rs4940086, can potentially elevate cervical cancer susceptibility in Bangladeshi women.

Similar content being viewed by others
References
Ferlay J, Soerjomataram I, Ervik M, Dikshit R, Eser S, Mathers CGLOBOCAN et al (2012) Estimated cancer incidence, mortality and prevalence worldwide in 2012. IARC, Lyon
Hussain SA, Sullivan R (2013) Cancer control in Bangladesh. Jpn J Clin Oncol 43:1159–1169. https://doi.org/10.1093/jjco/hyt140
Au WW, Sierra-Torres CH, Tyring SK (2003) Acquired and genetic susceptibility to cervical cancer. Mutat Res 544:361–364
Hildesheim A, Wang SS (2002) Host and viral genetics and risk of cervical cancer: a review. Virus Res 89:229–240
Chen X, Jiang J, Shen H, Hu Z (2011) Genetic susceptibility of cervical cancer. J Biomed Res 25:155–164. https://doi.org/10.1016/S1674-8301(11)60020-1
Pasche B (2001) Role of transforming growth factor beta in cancer. J Cell Physiol 186:153–168. https://doi.org/10.1002/1097-4652(200002)186:2<153:AID-JCP1016>3.0.CO;2-J
Kleeff J, Ishiwata T, Maruyama H et al (1999) The TGF-beta signaling inhibitor Smad7 enhances tumorigenicity in pancreatic cancer. Oncogene 18:5363–5372. https://doi.org/10.1038/sj.onc.1202909
Massagué J, Blain SW, Lo RS (2000) TGFbeta signaling in growth control, cancer, and heritable disorders. Cell 103:295–309
Horbelt D, Denkis A, Knaus P (2012) A portrait of transforming growth factor β superfamily signalling: background matters. Int J Biochem Cell Biol 44:469–474. https://doi.org/10.1016/j.biocel.2011.12.013
Xie W, Mertens JC, Reiss DJ et al (2002) Alterations of Smad signaling in human breast carcinoma are associated with poor outcome: a tissue microarray study. Cancer Res 62:497–505
Fleming NI, Jorissen RN, Mouradov D et al (2013) SMAD2, SMAD3 and SMAD4 mutations in colorectal cancer. Cancer Res 73:725–735. https://doi.org/10.1158/0008-5472.CAN-12-2706
Qiu W, Schönleben F, Li X, Su GH (2007) Disruption of transforming growth factor beta-Smad signaling pathway in head and neck squamous cell carcinoma as evidenced by mutations of SMAD2 and SMAD4. Cancer Lett 245:163–170. https://doi.org/10.1016/j.canlet.2006.01.003
Schutte M, Hruban RH, Hedrick L et al (1996) DPC4 gene in various tumor types. Cancer Res 56:2527–2530
MacGrogan D, Pegram M, Slamon D, Bookstein R (1997) Comparative mutational analysis of DPC4 (Smad4) in prostatic and colorectal carcinomas. Oncogene 15:1111–1114. https://doi.org/10.1038/sj.onc.1201232
Slattery ML, Herrick JS, Lundgreen A, Wolff RK (2011) Genetic variation in the TGF-β signaling pathway and colon and rectal cancer risk. Cancer Epidemiol Biomark Prev 20:57–69. https://doi.org/10.1158/1055-9965.EPI-10-0843
Ki K-D, Tong S-Y, Huh C-Y et al (2009) Expression and mutational analysis of TGF-β/Smads signaling in human cervical cancers. J Gynecol Oncol 20:117–121. https://doi.org/10.3802/jgo.2009.20.2.117
Yakicier MC, Irmak MB, Romano A et al (1999) Smad2 and Smad4 gene mutations in hepatocellular carcinoma. Oncogene 18:4879–4883. https://doi.org/10.1038/sj.onc.1202866
Chen T, de Vries EG, Hollema H et al (1999) Structural alterations of transforming growth factor-beta receptor genes in human cervical carcinoma. Int J Cancer 82:43–51
Iancu IV, Botezatu A, Goia-Ruşanu CD et al (2010) TGF-beta signalling pathway factors in HPV-induced cervical lesions. Roum Arch Microbiol Immunol 69:113–118
Maliekal TT, Antony M-L, Nair A et al (2003) Loss of expression, and mutations of Smad 2 and Smad 4 in human cervical cancer. Oncogene 22:4889–4897. https://doi.org/10.1038/sj.onc.1206806
Kloth JN, Kenter GG, Spijker HS et al (2008) Expression of Smad2 and Smad4 in cervical cancer: absent nuclear Smad4 expression correlates with poor survival. Mod Pathol 21:866–875. https://doi.org/10.1038/modpathol.2008.62
World Medical Association (2013) World medical association declaration of Helsinki: ethical principles for medical research involving human subjects. JAMA 310:2191–2194. https://doi.org/10.1001/jama.2013.281053
Daly AK, Monkman SC, Smart J et al (1998) Analysis of cytochrome P450 polymorphisms. Methods Mol Biol Clifton NJ 107:405–422. https://doi.org/10.1385/0-89603-519-0:405
Takebayashi S, Ogawa T, Jung KY et al (2000) Identification of new minimally lost regions on 18q in head and neck squamous cell carcinoma. Cancer Res 60:3397–3403
Samanta D, Datta PK (2012) Alterations in the Smad pathway in human cancers. Front Biosci Landmark Ed 17:1281–1293
Prunier C, Ferrand N, Frottier B et al (2001) Mechanism for mutational inactivation of the tumor suppressor Smad2. Mol Cell Biol 21:3302–3313. https://doi.org/10.1128/MCB.21.10.3302-3313.2001
Tian F, DaCosta BS, Parks WT et al (2003) Reduction in Smad2/3 signaling enhances tumorigenesis but suppresses metastasis of breast cancer cell lines. Cancer Res 63:8284–8292
Kuchenbaecker KB, Hopper JL, Barnes DR et al (2017) Risks of breast, ovarian, and contralateral breast cancer for BRCA1 and BRCA2 mutation carriers. JAMA 317:2402–2416. https://doi.org/10.1001/jama.2017.7112
Deng N, Zhou H, Fan H, Yuan Y (2017) Single nucleotide polymorphisms and cancer susceptibility. Oncotarget 8:110635–110649. https://doi.org/10.18632/oncotarget.22372
Chen J, Wu X, Huang Y et al (2016) Identification of genetic variants predictive of early onset pancreatic cancer through a population science analysis of functional genomic datasets. Oncotarget 7:56480–56490. https://doi.org/10.18632/oncotarget.10924
Yang IA, Fong KM, Zimmerman PV et al (2009) Genetic susceptibility to the respiratory effects of air pollution. Postgrad Med J 85:428–436. https://doi.org/10.1136/thx.2007.079426
Funding
None.
Author information
Authors and Affiliations
Contributions
PSH: performed all the experiments and drafted the manuscript; MNHA: conceived research design, performed statistical analyses, drafted and revised the manuscript; NAN, FI: assisted in experiments, revised manuscript; MRI: revised manuscript; AH: conceptualized the research; MSI: principal investigator.
Corresponding author
Ethics declarations
Conflict of interest
The authors disclose no competing interests.
Ethical approval
The current study was in accordance with the declaration of Helsinki and its further amendments and the ethical committee of the participating hospital approved the study protocol.
Informed consent
Each patient and control subject signed an informed consent document after they were informed of the study objectives.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Haque, P.S., Apu, M.N.H., Nahid, N.A. et al. SMAD2 rs4940086 heterozygosity increases the risk of cervical cancer development among the women in Bangladesh. Mol Biol Rep 47, 5033–5040 (2020). https://doi.org/10.1007/s11033-020-05572-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05572-7