Abstract
In mammary gland development, normal stem cell activity occurs in the embryonic stage and postnatally. Research supports that certain breast cancers contain a small sub-population of cells that mimic stem-like activity. It is believed stem cell activation in the mutated mature human mammary tissue is what drives quiescent epithelial cells to convert to mesenchymal states initiating migration, invasion, and metastasis in breast cancer. The goal of the work reported herein was to investigate early mammary development gene expression in the postnatal pig using fine needle biopsy methods in order to establish a reliable model for human breast cancer detection. Tissue samples were collected from pig mammary glands beginning at Day 11 of age through Day 39 in order to capture early postnatal-growth gene expression. Based on the initial clustering analysis, two distinct clusters of gene expression profiles occurred before and after Day 25 of mammary development. Gene set enrichment analysis (GSEA) ontology indicated the cellular processes that changed after Day 25, and many of these processes were implicated in epithelial–mesenchymal transition (EMT) signaling events. Gene expression in the postnatal pig was compared with the Epithelial–Mesenchymal Transition gene database (dbEMT) confirming the presence of EMT activity in this early developmental program. Information from this study will provide insight into early postnatal mammary gland development. In addition, mechanisms exploited by mutated mammary epithelial cells leading to cancer initiation and growth may be detected considering that mutated mammary epithelial cells can reactivate early developmental signals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Due to ethical considerations and minimal tissue acquisition from young human mammary glands, our current knowledge of human breast development has been limited. Therefore, interpretation of developing human breast morphology is based on analogies in the mouse. Additionally, cancer research progress has been hampered by lack of anatomically and physiologically relevant animal models, because large animal models more closely recapitulate human cancer and diseases [1]. Transitional animal models could bridge the gap between diagnostic discoveries and human clinical trials. The pig is currently advancing as a transitional model bridging the gap from mouse studies to human trials.
Comparing the morphology of developing mammary glands, humans and pigs develop terminal ductal lobular units (TDLU) and epithelial proliferation is concentrated in the endbuds or TDLU's. The mouse morphology has sparse ducts and alveolar tissue and epithelial proliferation occurs in ductal endbuds and alveoli [2].
This research report promotes the use of an innovative procedure (Dunn Biopsy) by collecting mammary tissue from the pig weekly beginning at 11 days of age and ending at 39 days of age. This methodology is described in the pending U.S. Provisional Patent No. 62/256,416 titled Dunn Biopsy: A method of fine needle biopsy of mammalian tissue over time as a diagnostic tool. This biopsy technique provides a representative sample of tissue that was not altered due to genetic modifications, introduction of exogenous cells, or surgical techniques. This technique allows for repeated sampling from the same subject over time eliminating genetic variability from different individuals. Due to this naturally occurring state, our model provides an alternative analogy of breast development morphology and gene expression.
Previous studies indicate that stem cell activity which occurs in early postnatal development may be reactivated in the mutated human adult mammary gland as a trigger to initiate cell proliferation in early breast cancer [3,4,5]. These mechanisms involved in early postnatal growth period could provide insight into the initiation of quiescent cells in the mature human adult. In developing tissue, epithelial cells undergo a transdifferentiation process known as the epithelial–mesenchymal transition (EMT) where they alter cell states from epithelial to mesenchymal [6]. The reverse program, mesenchymal–epithelial transition (MET), occurs during development to allow the formation of distinct tissues [6, 7].
Early-stage tumors exhibit epithelial phenotypes, and highly aggressive tumors display mesenchymal features; therefore, the EMT–MET phenomenon can be linked to tumor progression and malignancy. Since cancer cells can interconvert between phenotypes of epithelial or mesenchymal states [8] design of research strategies have been limited. Given the similar phenotypic attributes of the EMT–MET program in developing tissue and epithelial-based cancers, it is plausible that understanding biological mechanisms in natural, transdifferentiating states may provide critical information regarding initiation and metastasis.
By evaluating cellular mechanisms in their natural environment, the Dunn Biopsy technique could establish a comprehensive database allowing investigators to disentangle neo-natal mammary developmental processes. The focus of this research was to evaluate gene expression patterns of mammary development in postnatal pigs using a fine needle biopsy method. This information may provide insight into mechanisms exploited by mutated mammary epithelial cells leading to cancer initiation and growth.
Materials and methods
Animals
Animals were maintained in certified Association for Assessment and Accreditation of Laboratory Animal Care facilities (AAALAC), and animal handling adhered to the Institutional Animal Care and Use Committee (IACUC) approved protocols and guidelines. Three Yorkshire/Berkshire cross-bred gilts were used for this study and housed with sow and litter mates until weaning. At 21 days of age, the three animals were moved to a nursery where they remained through the final sampling.
Dunn biopsy procedure
Tissue samples were collected from pig mammary glands using the Dunn Biopsy method beginning at 11 days of age continuing weekly through Day 39. Mammary glands chosen for each biopsy procedure were randomized to prevent selection bias and remove differences in the mammary chain of pigs. Animals were manually restrained for preparation and biopsy procedure by holding the animal in a prone position on a cushioned surface. The mammary gland was triple washed with chlorhexidine and iodopovidone prior to administering 2 ml of lidocaine at the sample site. Mammary tissue was collected with a 14 g biopsy needle using the Dunn Biopsy technique (U.S. patent application number 62/256,416). Tissue samples were cut in half and stored in 10% neutral buffered formalin for immunohistochemistry and RNAlater (Sigma Aldrich) for RNA isolation and sequencing.
Histology preparation
Samples placed in 10% neutral buffered formalin remained at room temperature overnight followed by ethanol dehydration, xylene clearing and paraffin embedding (FFPE). Blocks were sectioned at 6 μm for hematoxylin and eosin (H&E) staining in order to confirm samples collected were representative of mammary parenchyma and to evaluate tissue morphology.
RNA isolation, library preparation, sequencing
Samples in RNAlater (Sigma Aldrich) were stored at − 20 °C until RNA isolation. Biopsy samples were prepared using Qiagen TissueRupter, treated with Qiagen on-column DNase, and RNA was isolated using Qiagen RNeasy kit. Samples were individually normalized and an Illumina compatible sequencing library was prepared robotically on a Microlab STAR with Illumina’s TruSeq stranded total RNA library prep kit. The libraries were assessed for size on a 2100 Agilent Bioanalyzer and sequence data collected on an Illumina HiSeq2500. Each sample was sequenced to a depth of at least 22 million reads. Quality metrics, using FastQC [9] were run on all samples. Trimming of low-quality bases was carried out with Trimmomatic [10]. Trimmed reads were aligned to the Sus scrofa 11.1 genome using STAR, a splice aware RNA-seq aligner [11]. Transcripts were then quantified using the FeatureCounts [12] tool in the subreads [13] package in R [14]. DESeq [15] was then used for differential expression analysis and gene expression normalization. Gene ontology analysis was performed using GSEA [16, 17] software and association with compared to the dbEMT database [18] for a more targeted analysis.
Results
A change in morphology occurs after 25 days of pig mammary gland development
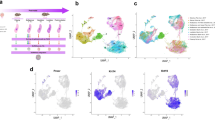
Postnatal time points from Day 11 through Day 39 of age were selected for data collection of mammary gland tissue in order to evaluate early postnatal gene expression. Tissue was collected using the Dunn Biopsy fine needle methodology which allows for genetic expression comparisons from the same animal's tissues at different time points (Fig. 1a). This methodology removes many of the confounding effects that can be caused by collecting tissue from different animals to perform developmental analyses over time. In order to analyze gene expression changes of early pig mammary gland development, RNA-seq was performed on the biopsied mammary tissue (Fig. 1b). RNA extraction quality for all time-points and the percent mapped after trimming are illustrated in Fig. 2a and b respectively.
Methods for data collection and analyses: a schematic illustration of the Dunn Biopsy methodology sampling from the same animal over time. Biopsy tissue was collected for each time point in RNAlater and 10% formalin for downstream applications. b RNA Seq was mapped using STAR to the Sus scrofa 11 reference genome and quantified using FeatureCounts using the Sus scrofa 11 transcriptome annotation file. The data was then processed for ontologies and changes in expression profile over time
Clustering of whole transcriptome gene expression analysis indicated a profound change in global gene expression that occurs after Day 25 of pig mammary gland development (Fig. 3a). This is consistent with histological examination where the appearance of TDLU's that were identified in Day 32 and Day 39 (Fig. 3b). Due to this substantial change of genetic expression and morphology prior to and after Day 25 of age, we decided to further probe what processes were altered after this point in development.
Patterns of gene expression and morphological changes: a gene expression clustering of all time points collected. Shows distinct changes occur in gene expression after day 25 of mammary gland development. b Morphological changes in the developing mammary gland. The presence of TDLU's begin to appear after day 25
Gene expression changes after 25 days of development are related to the epithelial mesenchymal transition
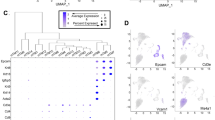
Investigations of mammary gene expression patterns led to the discovery of highly implicated EMT–MET developmental program events. In order to delineate what processes were altered due to noticeable gene expression changes, we performed a Gene Set Enrichment Analysis (GSEA). This analysis revealed several processes which significantly changed after Day 25 of gland development. Noticeably, we found processes down regulated after Day 25 associated with actin cytoskeleton, actin mediated cell contraction, regulation of calcium ion import, and contractile fiber (Fig. 4a). These processes are all implicated in different aspects of breast cancer progression.
Correlation of EMT events in the developing mammary gland: a GSEA analysis comparing gene expression before and after day 25 of development (p < 0.01, FDR < 0.1). b Gene expression clustering of genes known to be involved in EMT. c Gene expression of transcription factors known to be involved in EMT, showing expression before and after day 25 of gland development
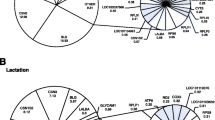
In addition to the gene expression changes known to be implicated in breast cancer, there are substantial changes to pathways involved in EMT. For this reason, we decided to further probe genes involved in EMT by analyzing their expression within our dataset. We identified EMT genes by examining those found in the dbEMT. This analysis (Fig. 4b) revealed the overall expression of EMT associated genes replicates results determined from the total gene set (Fig. 3a) where samples after Day 25 are more similar to each other than those earlier in development. This confirmed that changes occurring in the developing pig mammary gland included EMT associated events, which may be involved in driving other gene expression changes.
Gene expression changes in EMT-inducing transcription factors were analyzed to identify drivers of EMT related processes. The analyzed genes included CDH1, TWIST, TEAD, SNAI, VEGF, and SOX (Fig. 4c). The two genes with highest levels of upregulation in later time points included PRRX1 and TEAD1. PRRX1 has been shown to promote EMT in cancer, and TEAD1 enhances proliferation in cancer. The two genes with the strongest downregulation included CDH1 and ZEB2. CDH1 encodes E-cadherin which is highly implicated in the EMT pathway. Mutation of this gene leads to predisposition of certain cancers. ZEB2 has been shown to maintain stemness and is associated as an E-cadherin repressor. EMT related transcription identified through this pig development model indicate the pig mammary gland is an ideal candidate as a model system for mammary gland and cancer development.
Discussion
The focus of this research was to evaluate gene expression patterns of mammary gland development in postnatal pigs using the Dunn Biopsy method. To our knowledge, this is the first study that reports genetic expression patterns in young pig mammary glands from the same animals over time. This information may provide insight into mechanisms exploited by mutated mammary epithelial cells leading to cancer initiation and growth.
Puberty in pigs is defined as the first observed standing reflex in the presence of a boar [19]. The pig industry determines a pig has reached puberty as the age when the animal can be successfully bred, which can vary based on the physical composition of the animal [20]. Considering body size and physical attributes contribute to sexual maturation, the average pig would reach puberty between 120 and 150 days of age. Pig growth and development including the estrous cycle have been established, however mammary gland development prior to the onset of puberty has not been reported. Based on the results from this study, there are genetic and morphological changes that occur beginning as young as 25 days of age. The data support early pre-pubertal influences occur in the pig mammary gland based on the presence of TDLU's in animals at 32 and 39 days of age.
Whole transcriptome gene expression clustering analysis indicated a profound change in global gene expression after Day 25 of age which was consistent with the histological appearance of TDLU's after Day 25. This indicates the importance of EMT-MET mediated events in early mammary gland development. Processes that are highly implicated in EMT and cancer progression were identified by GSEA in the pig model, with significant downregulation after Day 25. These processes included actin cytoskeleton, actin-mediated cell contraction, regulation of calcium ion transport, and contractile fiber. The actin cytoskeleton is a primary contributor to cell shape, however remodeling of the actin cytoskeleton is associated with loss of cell–cell contact and linked to poor prognosis in breast cancer [21,22,23]. Actin-mediated cell contraction when downregulated decreases intercellular adhesion usually as a result of the loss of E-cadherin (CDH1). These events increase the opportunity for remodeling and carcinoma migration due to alterations in cell structure and adhesion [24]. The down regulation of calcium ion transport has been implicated in cancer progression due to the imperative need for calcium signaling in normal cellular behavior [25]. Additionally, EMT processes are associated with altered calcium transport [26]. The motile forces in cell migration are dependent on contractile fibers and the deregulation of these fibers are necessary for cell plasticity and motility in EMT and cancer progression [27].
The overall expression of EMT associated genes replicates the results determined from the total gene set where samples collected up to Day 25 are more similar to each other than those later in development after Day 25. These confirmed EMT associated events occurred in a similar pattern when compared with global gene expression in the early postnatal pig mammary model. EMT-related transcription factors implicated in human breast cancer were identified in our model including CDH1, TWIST, TEAD, SNAI, VEGF, and SOX [28,29,30,31,32,33,34]. The two genes with highest levels of upregulation in later time points include PRRX1 and TEAD1. PRRX1 has been shown to promote EMT and is involved in stem-like activities associated in cancer progression [35, 36]. TEAD1 is involved in apoptotic resistance in cancer and mediates cell growth and EMT [29, 30]. The two genes with the most significant downregulation include CDH1 and ZEB2. CDH1 encodes E-cadherin which is highly implicated in the EMT pathway. Mutation of this gene leads to predisposition of certain cancers [28] and increased EMT mediated events [37]. In addition, the loss of E-cadherin is associated with metastasis and poor prognosis in invasive breast cancer [38]. ZEB2 downregulation has been linked to stem cell properties and EMT events [39] and is known to be an E-cadherin repressor [40]. Given these findings, we report the gene expression patterns in the postnatal pig correlate to EMT mediated events in the human adult. The Dunn Biopsy sampled pig mammary gland serves as a translational model to evaluate EMT and EMT-processes that are implicated in cancer.
Considering 80% of life-threatening cancers are derived from epithelial tissue [41] the information reported herein warrants further investigation. Early stage tumors exhibit epithelial phenotypes and highly aggressive tumors display mesenchymal features [42], therefore the EMT-MET phenomenon can be linked to tumor progression and malignancy. Investigating aggressive subtypes of breast cancer indicates a positive association with mesenchymal features [43]. The EMT process in breast cancer is also linked to potential drug resistance [44] and disease recurrence [45]. Given the similar phenotypic attributes of the EMT–MET program in developing tissue and epithelial-based cancers, it’s plausible that understanding biological mechanisms through repetitive biopsied samples in natural transdifferentiating states of mammary development may provide critical information regarding initiation and metastasis.
Experimental models provide worthwhile insight to many human physiological processes including the progression of cancer, however no individual animal model can truly mimic all complex mechanisms. Therefore, different animal models need to be studied to provide additional information regarding development, disease progression, and response to treatment [46]. The pig mammary gland is an ideal alternative model system to study global changes of mammary development in order to evaluate the process of breast cancer progression.
Conclusion
Using the Dunn Biopsy method to evaluate tissue changes over time has provided a new methodology to evaluate gene expression changes in postnatal mammary glands. Based on the high relevance of EMT related activity identified in this study, these results indicate the pig mammary gland is an ideal candidate as a model system for investigating mammary gland development and breast cancer progression. Further studies will continue to evaluate the pig model for gene expression and how mutated cells can reactivate early developmental processes. Application of this technique may provide insight of the initiating steps involved in cancer progression.
References
Schachtschneider KM et al (2017) The oncopig cancer model: an innovative large animal translational oncology platform. Front Oncol 7:190
Rowson AR, Daniels KM, Ellis SE, Hovey RC (2012) Growth and development of the mammary glands of livestock: a veritable barnyard of opportunities. Semin Cell Dev Biol 23(5):557–566
Schmucker HS, Park JP, Coissieux M-M, Bentires-Alj M, Feltus FA, Booth BW (2017) RNA expression profiling reveals differentially regulated growth factor and receptor expression in redirected cancer cells. Stem Cells Dev 26(9):646–655
Hong D et al (2018) Epithelial-to-mesenchymal transition and cancer stem cells contribute to breast cancer heterogeneity. J Cell Physiol 233(12):9136–9144
Guen VJ, Chavarria TE, Kröger C, Ye X, Weinberg RA, Lees JA (2017) EMT programs promote basal mammary stem cell and tumor-initiating cell stemness by inducing primary ciliogenesis and Hedgehog signaling. Proc Natl Acad Sci USA 114(49):E10532–E10539
Nieto MA (2013) Epithelial plasticity: a common theme in embryonic and cancer cells. Science 342(6159):1234850–1234850
Thiery JP, Acloque H, Huang RYJ, Nieto MA (2009) Epithelial–mesenchymal transitions in development and disease. Cell 139(5):871–890
Shibue T, Weinberg RA (2017) EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat Rev Clin Oncol 14(10):611–629
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120
Dobin A et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7):923–930
Liao Y, Smyth GK, Shi W (2013) The Subread aligner: fast, accurate and scalable read mapping by seed-and- vote. Nucleic Acids Res 41(10):e108–e108
Liao Y, Smyth GK, Shi W (2018) The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Bioinformatics 47:e47
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11(10):R106
Subramanian A et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome- wide expression profiles. Proc Natl Acad Sci USA 102(43):15545–15550
Mootha VK et al (2003) PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34(3):267–273
Zhao M, Kong L, Liu Y, Qu H (2015) dbEMT: an epithelial–mesenchymal transition associated gene resource. Sci Rep 5(1):11459
See GM, Knauer MT (2017) Selection for age at puberty in swine: correlated response in sow productivity. J Anim Sci 95:14
Whittemore AT, Yang H (1989) Physical and chemical composition of the body of breeding sows with differing body subcutaneous fat depth at parturition, differing nutrition during lactation and differing litter size. Br Soc Anim Sci 48(1):203–212
Shankar J, Nabi IR (2015) Actin cytoskeleton regulation of epithelial mesenchymal transition in metastatic cancer cells. PLoS ONE 10(3):e0119954
da Silva PL et al (2015) Prolactin promotes breast cancer cell migration through actin cytoskeleton remodeling. Front Endocrinol 6:186
Al Absi A et al (2018) Actin cytoskeleton remodeling drives breast cancer cell escape from natural killer-mediated cytotoxicity. Cancer Res 78(19):5631–5643
Gari HH, DeGala GD, Ray R, Lucia MS, Lambert JR (2016) PRL-3 engages the focal adhesion pathway in triple-negative breast cancer cells to alter actin structure and substrate adhesion properties critical for cell migration and invasion. Cancer Lett 380(2):505–512
Prevarskaya N, Ouadid-Ahidouch H, Skryma R, Shuba Y (2014) Remodelling of Ca2+ transport in cancer: how it contributes to cancer hallmarks? Philos Trans R Soc Lond B 369(1638):20130097
Stewart TA, Yapa KTDS, Monteith GR (2015) Altered calcium signaling in cancer cells. Biochimica et Biophysica Acta (BBA) 1848(10):2502–2511
Mierke B, Rosel D, Fabry B, Brabek J (2008) Contractile forces in tumor cell migration. Eur J Cell Biol 87(8–9):669–676
Pharoah PDP, Guilford P, Caldas C (2001) Incidence of gastric cancer and breast cancer in CDH1 (E-cadherin) mutation carriers from hereditary diffuse gastric cancer families. Gastroenterology 121(6):1348–1353
Landin Malt A, Cagliero J, Legent K, Silber J, Zider A, Flagiello D (2012) Alteration of TEAD1 expression levels confers apoptotic resistance through the transcriptional up-regulation of livin. PLoS ONE 7(9):e45498
Zhang N et al (2014) The RNA-seq approach to discriminate gene expression profiles in response to melatonin on cucumber lateral root formation. J Pineal Res 56(1):39–50
Kaufhold S, Bonavida B (2014) Central role of Snail1 in the regulation of EMT and resistance in cancer: a target for therapeutic intervention. J Exp Clin Cancer Res 33(1):62
Jung H-Y, Yang J (2015) Unraveling the TWIST between EMT and cancer stemness. Cell Stem Cell 16(1):1–2
Mehta GA, Khanna P, Gatza ML (2019) Emerging role of SOX proteins in breast cancer development and maintenance. J Mammary Gland Biol Neoplasia 24(3):213–230
Singh S, Singh S, Lillard JW Jr, Singh R (2017) Drug delivery approaches for breast cancer. IJN 12:6205–6218
Brabletz T (2012) EMT and MET in Metastasis: where are the cancer stem cells? Cancer Cell 22(6):699–701
Guo J, Fu Z, Wei J, Lu W, Feng J, Zhang S (2015) PRRX1 promotes epithelial–mesenchymal transition through the Wnt/β-catenin pathway in gastric cancer. Med Oncol 32(1):393
Craene BD, Berx G (2013) Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer 13(2):97–110
Onder TT, Gupta PB, Mani SA, Yang J, Lander ES, Weinberg RA (2008) Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Can Res 68(10):3645–3654
Ren C et al (2014) Double-negative feedback loop between ZEB2 and miR-145 regulates epithelial–mesenchymal transition and stem cell properties in prostate cancer cells. Cell Tissue Res 358(3):763–778
Cong N et al (2013) Downregulated microRNA-200a promotes EMT and tumor growth through the Wnt/β-catenin pathway by targeting the E-cadherin repressors ZEB1/ZEB2 in gastric adenocarcinoma. Oncol Rep 29(4):1579–1587
Valastyan S, Weinberg RA (2011) Tumor metastasis: molecular insights and evolving paradigms. Cell 147(2):275–292
Huber MA, Kraut N, Beug H (2005) Molecular requirements for epithelial–mesenchymal transition during tumor progression. Curr Opin Cell Biol 17(5):548–558
Choi Y et al (2013) Epithelial–mesenchymal transition increases during the progression of in situ to invasive basal-like breast cancer. Hum Pathol 44(11):2581–2589
Oliveras-Ferraros C et al (2012) Epithelial-to-mesenchymal transition (EMT) confers primary resistance to trastuzumab (Herceptin). Cell Cycle 11(21):4020–4032
Cheng Q et al (2014) A signature of epithelial–mesenchymal plasticity and stromal activation in primary tumor modulates late recurrence in breast cancer independent of disease subtype. Breast Cancer Res 16(4):407
Lui Y et al (2015) Mammalian models of chemically induced primary malignancies exploitable for imaging-based preclinical theragnostic research. Quant Imaging Med Surg 5(5):708–729
Acknowledgements
Funding was provided through the generous support of Clemson University Creative Inquiry and Undergraduate Research Program.
Funding
This work was supported by internal funds secured through Clemson University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. The research reported in this article was performed with approval of the Clemson University Institutional Animal Care and Use Committee (IACUC) Animal Use Protocols (AUP) (AUP# 2015-064, AUP# 2019-004). This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moss, M.A., Williams, B., Ferdous, F. et al. Methodology to analyze gene expression patterns of early mammary development in pig models. Mol Biol Rep 47, 3241–3248 (2020). https://doi.org/10.1007/s11033-020-05362-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05362-1