Abstract
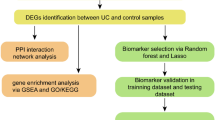
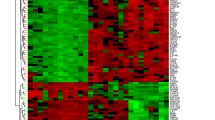
In this study we aimed to screen effective biomarkers for differential diagnosis of ulcerative colitis (UC) and Crohn’s disease (CD). By using the gene expression profile dataset GSE24287 including 47 ileal CD, 27 UC and 25 non-inflammatory bowel diseases control downloaded from Gene Expression Omnibus database, we identified the differentially expressed genes (DEGs) between UC patients and controls as well as between CD patients and controls (|log2FC(fold change)| > 1 and p < 0.05). Then Gene Ontology (GO) functional enrichment analyses were performed for these DEGs in two groups, followed by the construction of weight PPI (protein–protein interaction) networks. Subnets enriched for the PPIs and differentially expressed genes were constructed based on the weight PPI networks. The overlapping genes between the genes in the top 10 subnets with smallest p value and the DEGs were selected as the candidate genes of disease. A total of 75 DEGs were identified in UC group and 87 ones in CD group. There were 69 and 57 specific DEGs in CD group and UC group, respectively. The DEGs in CD group were mainly enriched in “inflammatory response” and “defense response”, while the most significantly enriched GO terms in UC group were “anion transport” and “chemotaxis”. FOS and SOCS3 were identified as candidate genes for CD and other three genes HELB, ZBTB16 and FAM107A were candidate genes for UC. In conclusion, there were distinct genetic alterations between UC and CD. The candidate genes identified in current study may be used as biomarkers for differential diagnosis of CD and UC.



Similar content being viewed by others
References
Xavier R, Podolsky D (2007) Unravelling the pathogenesis of inflammatory bowel disease. Nature 448(7152):427–434
Wu F, Dassopoulos T, Cope L, Maitra A, Brant SR, Harris ML, Bayless TM, Parmigiani G, Chakravarti S (2007) Genome-wide gene expression differences in Crohn’s disease and ulcerative colitis from endoscopic pinch biopsies: insights into distinctive pathogenesis. Inflamm Bowel Dis 13(7):807–821
Gophna U, Sommerfeld K, Gophna S, Doolittle WF, Van Zanten SJV (2006) Differences between tissue-associated intestinal microfloras of patients with Crohn’s disease and ulcerative colitis. J Clin Microbiol 44(11):4136–4141
Zhang T, DeSimone RA, Jiao X, Rohlf FJ, Zhu W, Gong QQ, Hunt SR, Dassopoulos T, Newberry RD, Sodergren E (2012) Host genes related to Paneth cells and xenobiotic metabolism are associated with shifts in human ileum-associated microbial composition. PLoS ONE 7(6):e30044
Podolsky DK (1991) Inflammatory bowel disease. N Engl J Med 325(13):928–937
Loftus EV Jr (2004) Clinical epidemiology of inflammatory bowel disease: incidence, prevalence, and environmental influences. Gastroenterology 126(6):1504–1517
Legnani P, Kornbluth A (2001) Difficult differential diagnoses in IBD: ileitis and indeterminate colitis. In: seminars in gastrointestinal disease. pp 211–222
Kaser A, Zeissig S, Blumberg RS (2010) Inflammatory bowel disease. Annu Rev Immunol 28:573–621. doi:10.1146/annurev-immunol-030409-101225
Bouma G, Strober W (2003) The immunological and genetic basis of inflammatory bowel disease. Nat Rev Immunol 3(7):521–533
Parronchi P, Romagnani P, Annunziato F, Sampognaro S, Becchio A, Giannarini L, Maggi E, Pupilli C, Tonelli F, Romagnani S (1997) Type 1 T-helper cell predominance and interleukin-12 expression in the gut of patients with Crohn’s disease. Am J Pathol 150(3):823
Heller F, Florian P, Bojarski C, Richter J, Christ M, Hillenbrand B, Mankertz J, Gitter AH, Bürgel N, Fromm M (2005) Interleukin-13 is the key effector Th2 cytokine in ulcerative colitis that affects epithelial tight junctions, apoptosis, and cell restitution. Gastroenterology 129(2):550–564
Vasiliauskas E (2003) Recent advances in the diagnosis and classification of inflammatory bowel disease. Curr Gastroenterol Rep 5(6):493–500
Davis S, Meltzer PS (2007) GEOquery: a bridge between the gene expression omnibus (GEO) and bio conductor. Bioinformatics 23(14):1846–1847
Diboun I, Wernisch L, Orengo C, Koltzenburg M (2006) Microarray analysis after RNA amplification can detect pronounced differences in gene expression using limma. BMC Genom 7(1):252
Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA (2003) DAVID: database for annotation, visualization, and integrated discovery. Genome Biol 4(5):P3
Licata L, Briganti L, Peluso D, Perfetto L, Iannuccelli M, Galeota E, Sacco F, Palma A, Nardozza AP, Santonico E (2012) MINT, the molecular interaction database: 2012 update. Nucleic Acids Res 40(D1):D857–D861
Goel R, Muthusamy B, Pandey A, Prasad TK (2011) Human protein reference database and human proteinpedia as discovery resources for molecular biotechnology. Mol Biotechnol 48(1):87–95
Chatr-aryamontri A, Breitkreutz B-J, Heinicke S, Boucher L, Winter A, Stark C, Nixon J, Ramage L, Kolas N, O’Donnell L (2013) The BioGRID interaction database: 2013 update. Nucleic Acids Res 41(D1):D816–D823
Wu C, Zhu J, Zhang X (2012) Integrating gene expression and protein–protein interaction network to prioritize cancer-associated genes. BMC Bioinformatics 13(1):182
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci 102(43):15545–15550
Liu Z-P, Wang Y, Zhang X-S, Chen L (2010) Identifying dysfunctional crosstalk of pathways in various regions of Alzheimer’s disease brains. BMC Syst Biol 4(Suppl 2):S11
Fiocchi C (1998) Inflammatory bowel disease: etiology and pathogenesis. Gastroenterology 115(1):182–205
Sartor RB (2006) Mechanisms of disease: pathogenesis of Crohn’s disease and ulcerative colitis. Nat Clin Pract Gastroenterol Hepatol 3(7):390–407
Samten B, Townsend JC, Weis SE, Bhoumik A, Klucar P, Shams H, Barnes PF (2008) CREB, ATF, and AP-1 transcription factors regulate IFN-γ secretion by human T cells in response to mycobacterial antigen. J Immunol 181(3):2056–2064
Roy S, Charboneau R, Melnyk D, Barke RA (2000) Interleukin-4 regulates macrophage interleukin-12 protein synthesis through a c-fos mediated mechanism. Surgery 128(2):219–224
Miao T, Wu D, Zhang Y, Bo X, Xiao F, Zhang X, Magoulas C, Subang MC, Wang P, Richardson PM (2008) SOCS3 suppresses AP-1 transcriptional activity in neuroblastoma cells through inhibition of c-Jun N-terminal kinase. Mol Cell Neurosci 37(2):367–375
Lovato P, Brender C, Agnholt J, Kelsen J, Kaltoft K, Svejgaard A, Eriksen KW, Woetmann A, Ødum N (2003) Constitutive STAT3 activation in intestinal T cells from patients with Crohn’s disease. J Biol Chem 278(19):16777–16781
Boirivant M, Fuss IJ, Chu A, Strober W (1998) Oxazolone colitis: a murine model of T helper cell type 2 colitis treatable with antibodies to interleukin 4. J Exp Med 188(10):1929–1939
Heller F, Fuss IJ, Nieuwenhuis EE, Blumberg RS, Strober W (2002) Oxazolone colitis, a Th2 colitis model resembling ulcerative colitis, is mediated by IL-13-producing NK-T cells. Immunity 17(5):629–638
Engel I, Kronenberg M (2012) Making memory at birth: understanding the differentiation of natural killer T cells. Curr Opin Immunol 24(2):184–190
Kholodnyuk ID, Kozireva S, Kost-Alimova M, Kashuba V, Klein G, Imreh S (2006) Down regulation of 3p genes, LTF, SLC38A3 and DRR1, upon growth of human chromosome 3–mouse fibrosarcoma hybrids in severe combined immunodeficiency mice. Int J Cancer 119(1):99–107
Mari F, Hermanns P, Giovannucci-Uzielli ML, Galluzzi F, Scott D, Lee B, Renieri A, Unger S, Zabel B, Superti-Furga A (2009) Refinement of the 12q14 microdeletion syndrome: primordial dwarfism and developmental delay with or without osteopoikilosis. Eur J Hum Genet 17(9):1141–1147
Koepsell H, Lips K, Volk C (2007) Polyspecific organic cation transporters: structure, function, physiological roles, and biopharmaceutical implications. Pharm Res 24(7):1227–1251
Wojtal KA, Eloranta JJ, Hruz P, Gutmann H, Drewe J, Staumann A, Beglinger C, Fried M, Kullak-Ublick GA, Vavricka SR (2009) Changes in mRNA expression levels of solute carrier transporters in inflammatory bowel disease patients. Drug Metab Dispos 37(9):1871–1877
Acknowledgments
This work was supported partly by the Science and Technology Program of Liaoning Province (2014021083 & 2013225303); and the Science and Technology Program of Shenyang (F14-158-9-49).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lin, LJ., Zhang, Y., Lin, Y. et al. Identifying candidate genes for discrimination of ulcerative colitis and Crohn’s disease. Mol Biol Rep 41, 6349–6355 (2014). https://doi.org/10.1007/s11033-014-3469-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3469-y