Abstract
Extracts of Anthemis hyalina (Ah), Nigella sativa (Ns) and peels of Citrus sinensis (Cs) have been used as folk medicine to fight antimicrobial diseases. To evaluate the effect of extracts of Ah, Ns and Cs on the replication of coronavirus (CoV) and on the expression of TRP genes during coronavirus infection, HeLa-CEACAM1a (HeLa-epithelial carcinoembryonic antigen-related cell adhesion molecule 1a) cells were inoculated with MHV-A59 (mouse hepatitis virus–A59) at moi of 30. 1/50 dilution of the extracts was found to be the safe active dose. ELISA kits were used to detect the human IL-8 levels. Total RNA was isolated from the infected cells and cDNA was synthesized. Fluidigm Dynamic Array nanofluidic chip 96.96 was used to analyze the mRNA expression of 21 TRP genes and two control genes. Data was analyzed using the BioMark digital array software. Determinations of relative gene expression values were carried out by using the 2−∆∆Ct method (normalized threshold cycle (Ct) value of sample minus normalized Ct value of control). TCID50/ml (tissue culture infectious dose that will produce cytopathic effect in 50 % of the inoculated tissue culture cells) was found for treatments to determine the viral loads. The inflammatory cytokine IL-8 level was found to increase for both 24 and 48 h time points following Ns extract treatment. TRPA1, TRPC4, TRPM6, TRPM7, TRPM8 and TRPV4 were the genes which expression levels changed significantly after Ah, Ns or Cs extract treatments. The virus load decreased when any of the Ah, Ns or Cs extracts was added to the CoV infected cells with Ah extract treatment leading to undetectable virus load for both 6 and 8hpi. Although all the extract treatments had an effect on IL-8 secretion, TRP gene expression and virus load after CoV infection, it was the Ah extract treatment that showed the biggest difference in virus load. Therefore Ah extract is the best candidate in our hands that contains potential treatment molecule(s).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronaviruses (CoV), members of the Coronaviridae family, are enveloped viruses that contain non-segmented, positive-stranded genomic RNA [1–6]. Ultrastructural analysis of CoV have revealed that they form pleomorphic particles that are roughly spherical but show variations in size (80–120 nm in diameter) and shape [7–10]. The entire CoV replication cycle take places in the cytoplasm. In general, enveloped viruses are able to use a variety of cellular membranes at different steps of their life cycles. In fact, almost any membranous subcellular compartment appears to be used by particular viruses to support their replication complex and assembly process [10, 11]. Coronaviridae family causes various diseases including bronchitis, gastroenteritis, hepatitis systemic diseases and etc. on humans, animal and birds [12, 13]. Both host and viral factors affect the coronavirus virulence and the disease severity in animals. The disease is generally most severe in newborns [14].
In spring 2003, a new human CoV has been infamously notorious due to an epidemic outbreak in South East Asia and Canada [15]. At the time, the accused virus was rapidly identified as the SARS-CoV but it did not look like the human CoVs. The SARS-CoV alarmed the world because it sickened more than 8,000 people and killed nearly 800 of them [16]. Recently, a novel coronavirus caused respiratory infectious disease with high mortality. The virus is called Middle East respiratory syndrome coronavirus (MERS-CoV). This novel MERS-CoV was first reported in Saudi Arabia and eight other countries in 2012 [17]. Currently, there are no approved drugs against CoVs but some potential therapies have been proposed.
It has been revealed that nucleocapsid (N) and non-structural protein 3 (nsp3) interactions is important for CoV replication [18]. The nsp3 interacts with N protein through its EF motif site that contains a calcium binding domain, which can mean that this interaction could depend on calcium. It has been shown that intracellular calcium signaling triggers the elevation of reactive oxygen species in mitochondria that effect replication of influenza virus [19]. Also poliovirus infection increases intracellular calcium concentration [20]. These findings suggest that intracellular calcium concentration might be important for viral replication; therefore viruses may be targeting cellular mechanisms that regulate this concentration.
One of the possible targets of viruses to change the intracellular calcium concentration are the ion channels. The transient receptor potential proteins (TRPs) is group of ion channels family that are responsible for a wide range of cellular functions including adjusting intracellular Ca2+ concentration [21]. The TRP superfamily can be divided into seven families: TRPC (canonical), TRPV (vanilloid), TRPM (melastatin), TRPN and TRPA (ankyrin), TRPML (mucolipin) and TRPP (polycystin) [22]. TRP channels are triggered by a many different stimuli such as; growth factors, chemicals, temperature, depletion of intracellular calcium level and etc. Triggering mechanism of channels and selectivity can be different between the TRPs [23]. The expression levels of TRPM and TRPV families have been associated with tumor growth and progression [23–25].
Herbal remedies have been used as folk medicine to treat many diseases. Some herb extracts were shown to inhibit virus replication [26, 27]. One of these herbs is a chamomile plant called Anthemis hyalina (Ah). A. hyalina belongs to plant family of Asreraceae genus; Anthemis, species; A. hyalina DC. Chamomile is regularly used in tea making and to relieve stomachache and digestive system problems. Chamomile was shown to have anti-inflammatory effects and to decrease wound healing time [28]. Antimicrobial activity of different species in the genus of Anthemis have been documented [29]. Twenty-four different components were characterized in Ah extract. The major components were carvacrol (38.4 %) and α-pinene (30.9 %) [30].
Citrus sinensis (Cs) belongs to plant family of Rutaceae, genus citrus, species Citrus sinensis (L.) Osbeck. Another herb used for various treatments is Cs, known as sweet orange. In Turkey, people consume Cs by making jam and eating especially during flu seasons in order to get more vitamin C, which is believed to help recover from flu. Cs peels extract contain high amount of flavonoids, limonene and linalool [31]. Cs peel extract has antioxidant and antimicrobial activities [31, 32]. Limonene is an insecticidal compound with low oral or contact toxicity to mammals [33]. Flavonoids have also been shown to have antiviral activity [34] [35].
Nigella Sativa belongs to the plant family of Ranunculaceae, genus; Nigella, species; Nigella sativa L. Ns seeds that are known as black cumin is another plant that is used for various treatments. In Turkey, Ns seeds are commonly used on breads and cookies. In addition, some people mix Ns seeds and honey to use for upper respiratory tract infections and some stomach diseases. Ns extract has antiviral, antitumor and antimicrobial activities [36–38]. Ns extract oil contains thymoquinone (27.8–57.0 %), ρ-simen (7.1–15.5 %), karvakrol (5.8–11.6 %), t-anetol (0.25–2.3 %), 4-terpineol (2.0–6.6 %) and longifoline (1.0–8.0 %) [39, 40]. It has been revealed that thymoquinone ve dithymoquinone are active components in Ns extract [41].
In this study we set to find the effects of Ah, Cs and Ns extracts on the replication of CoV and the possible involvement of TRP genes in the replication of CoV.
Materials and methods
Cells, virus and time-course analysis of MHV infection
HeLa-CEACAM1a (the epithelial carcinoembryonic antigen-related cell adhesion molecule 1) and murine fibroblast LR7 cells [42] that were used to propagate and titrate MHV-A59 (mouse hepatitis virus–A59) were maintained in Dulbecco’s Modified Eagle Medium (DMEM; Sigma, St. Louis, MO) containing 10 % fetal calf serum (Thermo, Waltham, MA), 100 IU of penicillin/ml and 100 μg/ml of streptomycin (both from Life Technologies, Rochester, NY). HeLa-CEACAM1a cells were inoculated with MHV-A59 at a moi of 30 [43, 44]. After 30 min, the infected cells were washed and maintained in complete medium. Subsequently, the infected cells and culture supernatants were collected for analysis at 0, 6, 8 h, p.i. When plant extracts were added to the HeLa-CEACAM1a cells, they were added after viral infection and were washed away 1 h later.
Preparation of extracts
Air-dried plant powdered; seeds of Ns, flowers and buds of Ah and peels of Cs were used for extraction (100 g of material of each separately). All extraction experiments were performed by 200 ml ethanol addition onto powders and incubation at room temperature for a 24 h. The extracts were filtered using Whatman filter paper and ethanol was eliminated by rotary vacuum evaporator at 55 °C. The plant extracts were dissolved in 10 ml distilled water.
Quantitative analysis of IL-8 by enzyme-linked immunosorbent assay (ELISA)
ELISA kits were used to detect the human IL-8 according to the user’s manual (Quantikine, ELISA, Human CXCL8/IL-8 Immunoassay, DY208, R&D Systems, Minneapolis, MN). HeLa CEACAM1 cells were treated with 1/50 and 1/100 diluted concentrations of Ns, Ah and Cs extracts. Additionally HeLa CEACAM1 cells were infected with MHV-A59. Infected cells were treated with 1/50 and 1/100 diluted concentrations of Ns, Ah and Cs extracts. Subsequently, culture supernatants were collected at 0, 6, and 8 h post infection to measure IL-8 levels by using enzyme-linked immunosorbent assay (ELISA).
Monitoring the extracellular release of the virus
Infected HeLa CEACAM1a supernatants were collected and end point dilutions were made for TCID50 analysis at 6 and 8 h post infection. The amount of virus present in the culture supernatants were evaluated by using LR7 cells and TCID50 values were calculated.
Isolation of total RNA and TaqMan analysis
Total RNA was isolated from the infected cells using RNeasy mini-kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions with subsequent DNaseI treatment on the column. The amount of RNA concentration was determined by spectrometry using a Nanodrop 100 (Thermo Scientific, DE, USA).
cDNA synthesis
Complementary DNA (cDNA) was synthesized by using Qiagen miScript Reverse Transcription Kit (Qiagen Sample and Assay Technologies, Hilden, Germany) according to manufacturer’s instructions. Mixture of 5× miScript RT buffer and miScript Reverse Transcription mix was prepared at given ratios and 1.25 μl of it was distributed for each reaction. 0.2 μg of RNA samples was added on reaction mixtures and they were incubated at 37 °C for 60 min then at 95 °C for 5 min. Synthesized cDNAs were then diluted 1:5 with low EDTA TE buffer and subsequently transferred on ice block and kept at −20 °C till use.
Fluidigm real-time PCR quantifications
Fluidigm Dynamic Array nanofluidic chip 96.96 was used for mRNA expression assay (Fluidigm, South San Francisco, CA USA). mRNA expression levels of 23 genes were analyzed by BioMark™ HD System real-time PCR (Fluidigm) (Table 1). Scrutinizing of the data and counting of the generated signals in reaction chambers was achieved by fluidigm real-time PCR Analysis software (Fluidigm).
Intracellular calcium level determination using atomic absorption spectrometer
HeLa CEACAM1a cells were infected with MHV-A59 cells. After infection, HeLa CEACAM1 cells were incubated with Ns, Ah and Cs extracts for 6 and 8 h. Infected cells were collected for quantification of total calcium ions. Concentration of calcium ions (μg/l) was determined by using an AAS-400 Perkin-Elmer (USA) atomic absorption spectrometer (FAAS) in an air-acetylene flame, according to the user’s manual provided by the manufacturer. A calcium hallow cathode lamp was used as the radiation source at 423 nm.
Statistical analysis
Data was analyzed using the BioMark digital array software and the numbers of positive chambers were corrected to estimate the true number of copies. This number was used to determine the number of copies in the original sample. Determinations of relative gene expression were carried out by using the 2−∆∆Ct method (comparative Ct method) where Ct values of the samples of interest are compared with a control (uninfected cells) after the Ct values of the control and the samples are normalized to an endogenous housekeeping gene. Expression levels were shown relative to the uninfected cells. GAPDH was used as the housekeeping gene for normalization of the expressions.
Results and discussion
Determination of maximum non-toxic doses
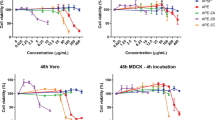
In order to assess any possible toxic effects of Ns, Ah and Cs extracts on HeLa CEACAM1a cells, serial dilutions (1/10, 1/50, 1/100, 1/1000, 1/10.000) of Ns, Ah and Cs extracts were incubated with HeLa CEACAM1a cells for 24 and 48 h. The cell viability was assessed by the tetrazolium salt 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. These three plant extracts showed toxic effect at 1/10 dilution, however toxicity of the extracts clearly decreased at 1/50 dilution for 24 and 48 h time points (Figs. 1, 2). These results suggest that the cytotoxic effect of these plants extracts were concentration dependent but not time-dependent in that less cells were viable at 1/10 dilution compared to 1/50 dilution. Based on these results, the plant extract dilution of 1/50 was accepted as an active dose.
Effects of Ns, Ah and Cs extracts on the survival of HeLa CEACAM1a cells. HeLa CEACAM1a cells were incubated with the diluted extracts for 24 h and cell toxicity was measured by MTT assay. The bars show the averages from 2 independent experiments with 4 repeats for each treatment. Error bars show standard error. C negative control
Effects of Ns, Ah and Cs extracts on the survival of HeLa CEACAM1a cells. HeLa CEACAM1a cells were incubated with the extracts for 48 h and cell toxicity was measured by MTT assay. The bars show the averages from two independent experiments with four repeats for each treatment. Error bars show standard error. C negative control
IL-8 levels following extract treatments of CoV infected cells
The levels of the inflammatory cytokine IL-8 in the supernatants of cells that were treated with the plant extracts were determined by ELISA (Fig. 3). IL-8 was chosen for its possible role in preventing DC cells from priming T-cells during CoV infection and inducing calcium signaling that might be related to TRP genes [45]. IL-8 levels increased following treatment by Ns, Ah and Cs extracts for both 1/50 and 1/100 dilutions at 24 h. However IL-8 levels had decreased at 48 h for Ah and Cs treated cells. This suggests that Ns extract was better at stimulating IL-8 secretion from infected cells to lead to a bigger inflammatory response. When IL-8 levels were measured following plant extract treatment of infected cells 6 and 8 h post-infection it was found that IL-8 levels were below the lowest standard and very low (Supplementary figure). Therefore we cannot comment on the effect of plant extracts on IL-8 secretion from CoV infected cells.
Expressions of TRP genes following extract treatment of CoV infected cells
To show effects of the Ns, Ah and Cs extracts on expression of TRP genes during CoV infection, active doses of the Ns, Ah and Cs extracts were added to HeLa CEACAM1a cells one hour before CoV infection. Subsequently, the cells were infected with CoV for 6 h post-infection and 8 h. mRNA expression levels for TRP channel families were investigated by using fluidigm RT PCR. GAPDH expression was used for normalization of gene expression. Uninfected cells were used as a negative control. The infected but not plant extract exposed cells were used as positive control.
The expressions of TRP genes were evaluated 6 and 8 h post-infection. These time points were chosen because CoV shows its cytopathic effects by 8 h post-infection [43]. The changes in the expression levels were determined by calculating fold-changes (2−ΔΔCt) for the TRP genes and normalizing them by dividing with the fold-change for the control gene GAPDH.
Fold-change values greater than 1 indicate an up-regulation and fold-change values less than 1 indicate down-regulation. However fold-change values more than 2 and less than 0.5 are considered significant.
Based on the TRP genes expression analysis results, the expression levels were mainly down (Fig. 4). The TRP channels affected by extracts were TRPA1, TRPC4, TRPM6, TRPM7, TRPM8 and TRPV4. Treatment with Ns, Ah and Cs extracts during CoV infection resulted in the down-regulation of TRPM6 and TRPA1 for both 6 and 8 h time points. TRPC4 was down regulated following Ns and Ah treatments for both 6 and 8 h time points. TRPM7 was down regulated only after Ns treatment for the two time points. Ah treatment resulted in down-regulation of TRPV4 for the two time points. Cs treatment resulted in down-regulation of TRPM8 for the two time points.
Monitoring the extracellular release of the virus
In order to determine the number of viruses following extracts treatment and viral infection, TCID50/ml was determined for the conditions and it was found that compared to the control group, extract treatments lowered the virus loads (Fig. 5). In the case of Ah treatments there was no detectable virus. Following Ns treatment the number of viruses was very low at 6 h post infections and it was 1/10th of the control amount by 8 h post infections. Cs treatment resulted in the same number of viruses at 6 and 8 h post infections, which were 1/10th of the positive control numbers by 8 h.
Virus loads following Ns, Ah and Cs extract treatment of infected HeLa CEACAM1a cells after 6 and 8 h post infections. The amount of virus present in the culture supernatants were evaluated by using LR7 cells and TCID50 values were calculated. The bars show the averages from two independent experiments with two repeats
Intracellular calcium levels
Following the determination of the expression of TRP genes, we sought to find the intracellular calcium levels (Fig. 6). Intracellular calcium levels did not change after addition of plant extracts to uninfected cells. However addition of plant extracts to CoV infected cells increased intracellular calcium levels.
Conclusion
The results presented here suggest that treatment of cells with Ns, Ah or Cs extracts prior to infection with CoV decreases the replication of the virus. IL-8 secretion was increased at 24 h for Ns and Ah extract treatments but Cs treatment did not show a significant increase in IL-8 secretion at 24 h. However, IL-8 secretion was very low at early time points after infection (Fig. 3). Therefore it is not known if IL-8 levels are related to a decreased virus load.
Following gene expression analysis the TRP genes with changed expression levels were identified as follows: TRPM6 and TRPA1 were the two genes that were down regulated for both 6 and 8 h time points in CoV infected and extract treated cells, TRPC4 gene was down regulated following Ns and Ah treatments for both 6 and 8 h time points, TRPM7 was down regulated only after Ns treatment for the two time points, TRPV4 gene was down regulated following Ah treatment for the two time points and TRPM8 was down regulated following Cs treatment for the two time points.
A general down regulation trend for TRP genes was observed in the CoV infected and extract treated cells. Together with the data showing that virus loads were decreased upon extract treatments we predicted that TRP genes might be involved in CoV survival inside cells. Considering that replication of other viruses were related to intracellular calcium concentration and TRPs modulate ion concentrations, including calcium we measured intracellular calcium concentrations and found that they increased following plant extract treatments of infected cells. The controversial TRP gene expression and calcium concentration results suggest that they may not be related to decreased viral loads in plant extract treated infected cells or decreased TRP gene expression and/or increased calcium concentration decrease viral loads through unknown mechanisms. Undetectable levels of the virus after Ah treatment can mean a specific factor being involved following that treatment and this factor can be the TRPV4 gene, whose expression went down only in the case of Ah treatment for both 6 and 8 h time points. Currently there is no literature tying TRP genes to viral survival.
The reason for decreased virus loads following extract treatments is not known at this point. Further research is required in order to identify the specific ingredient(s) in the extracts that led to this phenotype and also to identify the mechanism by which it prevents virus replication or leads to more efficient virus killing. Undetectable virus loads after Ah extract treatment can mean future research can identify certain molecule(s) in Ah extract that can be used as highly effective drugs against not only CoV but also other viruses as well.
References
Perlman S, Netland J (2009) Coronaviruses post-SARS: update on replication and pathogenesis. Nat Rev Microbiol 7:439–450
Sawicki SG, Sawicki DL, Siddell SG (2007) A contemporary view of coronavirus transcription. J Virol 81:20–29
Wijegoonawardane PK, Cowley JA, Phan T, Hodgson RA, Nielsen L, Kiatpathomchai W, Walker PJ (2008) Genetic diversity in the yellow head nidovirus complex. Virology 380:213–225
Brian DA, Baric RS (2005) Coronavirus genome structure and replication. Curr Top Microbiol Immunol 287:1–30
Ziebuhr J, Snijder EJ, Gorbalenya AE (2000) Virus-encoded proteinases and proteolytic processing in the Nidovirales. J Gen Virol 81:853–879
Lai MM, Cavanagh D (1997) The molecular biology of coronaviruses. Adv Virus Res 48:1–100
Barcena M, Oostergetel GT, Bartelink W, Faas FG, Verkleij A, Rottier PJ, Koster AJ, Bosch BJ (2009) Cryo-electron tomography of mouse hepatitis virus: insights into the structure of the coronavirion. Proc Natl Acad Sci USA 106:582–587
Davies HA, Macnaughton MR (1979) Comparison of the morphology of three coronaviruses. Arch Virol 59:25–33
de Haan CA, Rottier PJ (2005) Molecular interactions in the assembly of coronaviruses. Adv Virus Res 64:165–230
Masters PS (2006) The molecular biology of coronaviruses. Adv Virus Res 66:193–292
de Haan CA, van Genne L, Stoop JN, Volders H, Rottier PJ (2003) Coronaviruses as vectors: position dependence of foreign gene expression. J Virol 77:11312–11323
Chang CK, Sue SC, Yu TH, Hsieh CM, Tsai CK, Chiang YC, Lee SJ, Hsiao HH, Wu WJ, Chang WL, Lin CH, Huang TH (2006) Modular organization of SARS coronavirus nucleocapsid protein. J Biomed Sci 13:59–72
Weiss SR, Navas-Martin S (2005) Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol Mol Biol Rev 69:635–664
Garbino J, Crespo S, Aubert JD, Rochat T, Ninet B, Deffernez C, Wunderli W, Pache JC, Soccal PM, Kaiser L (2006) A prospective hospital-based study of the clinical impact of non-severe acute respiratory syndrome (Non-SARS)-related human coronavirus infection. Clin Infect Dis 43:1009–1015
Drosten C, Gunther S, Preiser W, van der Werf S, Brodt HR, Becker S, Rabenau H, Panning M, Kolesnikova L, Fouchier RA, Berger A, Burguiere AM, Cinatl J, Eickmann M, Escriou N, Grywna K, Kramme S, Manuguerra JC, Muller S, Rickerts V, Sturmer M, Vieth S, Klenk HD, Osterhaus AD, Schmitz H, Doerr HW (2003) Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med 348:1967–1976
Liang G, Chen Q, Xu J, Liu Y, Lim W, Peiris JS, Anderson LJ, Ruan L, Li H, Kan B, Di B, Cheng P, Chan KH, Erdman DD, Gu S, Yan X, Liang W, Zhou D, Haynes L, Duan S, Zhang X, Zheng H, Gao Y, Tong S, Li D, Fang L, Qin P, Xu W (2004) Laboratory diagnosis of four recent sporadic cases of community-acquired SARS, Guangdong Province. China. Emerg Infect Dis 10:1774–1781
Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA (2012) Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med 367:1814–1820
Hurst KR, Ye R, Goebel SJ, Jayaraman P, Masters PS (2010) An interaction between the nucleocapsid protein and a component of the replicase-transcriptase complex is crucial for the infectivity of coronavirus genomic RNA. J Virol 84:10276–10288
Gong G, Waris G, Tanveer R, Siddiqui A (2001) Human hepatitis C virus NS5A protein alters intracellular calcium levels, induces oxidative stress, and activates STAT-3 and NF-kappa B. Proc Natl Acad Sci USA 98:9599–9604
Irurzun A, Arroyo J, Alvarez A, Carrasco L (1995) Enhanced intracellular calcium concentration during poliovirus infection. J Virol 69:5142–5146
Gkika D, Prevarskaya N (2009) Molecular mechanisms of TRP regulation in tumor growth and metastasis. Biochim Biophys Acta 1793:953–958
Santoni G, Farfariello V (2011) TRP channels and cancer: new targets for diagnosis and chemotherapy. Endocr Metab Immune Disord Drug Targets 11:54–67
Clapham DE (2003) TRP channels as cellular sensors. Nature 426:517–524
Amantini C, Mosca M, Nabissi M, Lucciarini R, Caprodossi S, Arcella A, Giangaspero F, Santoni G (2007) Capsaicin-induced apoptosis of glioma cells is mediated by TRPV1 vanilloid receptor and requires p38 MAPK activation. J Neurochem 102:977–990
Duncan LM, Deeds J, Hunter J, Shao J, Holmgren LM, Woolf EA, Tepper RI, Shyjan AW (1998) Down-regulation of the novel gene melastatin correlates with potential for melanoma metastasis. Cancer Res 58:1515–1520
Pleschka S, Stein M, Schoop R, Hudson JB (2009) Anti-viral properties and mode of action of standardized Echinacea purpurea extract against highly pathogenic avian influenza virus (H5N1, H7N7) and swine-origin H1N1 (S-OIV). Virol J. 6:197
Zhu H, Zhang Y, Ye G, Li Z, Zhou P, Huang C (2009) In vivo and in vitro antiviral activities of calycosin-7-O-beta-D-glucopyranoside against coxsackie virus B3. Biol Pharm Bull 32:68–73
Reddy KK, Grossman L, Rogers GS (2011) Common complementary and alternative therapies with potential use in dermatologic surgery: risks and benefits. J Am Acad Dermatol 68:e127–135
Uzel A, Guvensen A, Cetin E (2004) Chemical composition and antimicrobial activity of the essential oils of Anthemis xylopoda O. Schwarz from Turkey. J Ethnopharmacol 95:151–154
Rustaiyana A, Masoudib S, Ezatpourb L, Danaiib E, Taherkhanib M, Aghajanic Z (2011) Composition of the Essential Oils of Anthemis Hyalina DC., Achillea Nobilis L. and Cichorium intybus L. Three Asteraceae Herbs Growing Wild in Iran. Journal of Essential Oil Bearing Plants. 14:472–480
Ademosun AO, Oboh G (2013) Anticholinesterase and antioxidative properties of water-extractable phytochemicals from some citrus peels. J Basic Clin Physiol Pharmacol 2:1–6
Dhiman A, Nanda A, Ahmad S, Narasimhan B (2011) In vitro antimicrobial activity of methanolic leaf extract of Psidium guajava L. J Pharm Bioallied Sci. 3:226–229
Kumar P, Mishra S, Malik A, Satya S (2011) Insecticidal evaluation of essential oils of Citrus sinensis L. (Myrtales: Myrtaceae) against housefly, Musca domestica L. (Diptera: Muscidae). Parasitol Res 110:1929–1936
Carvalho OV, Botelho CV, Ferreira CG, Ferreira HC, Santos MR, Diaz MA, Oliveira TT, Soares-Martins JA, Almeida MR, Silva Junior A (2013) In vitro inhibition of canine distemper virus by flavonoids and phenolic acids: implications of structural differences for antiviral design. Res Vet Sci 95:717–724
Su X, D’Souza DH (2013) Naturally occurring flavonoids against human norovirus surrogates. Food Environ Virol. 5:97–102
Ait Mbarek L, Ait Mouse H, Elabbadi N, Bensalah M, Gamouh A, Aboufatima R, Benharref A, Chait A, Kamal M, Dalal A, Zyad A (2007) Anti-tumor properties of blackseed (Nigella sativa L.) extracts. Braz J Med Biol Res 40:839–847
Bakathir HA, Abbas NA (2012) Detection of the antibacterial effect of Nigella sativa ground seeds with water. Afr J Tradit Complement Altern Med 8:159–164
Salem ML, Hossain MS (2000) Protective effect of black seed oil from Nigella sativa against murine cytomegalovirus infection. Int J Immunopharmacol 22:729–740
Ghosheh OA, Houdi AA, Crooks PA (1999) High performance liquid chromatographic analysis of the pharmacologically active quinones and related compounds in the oil of the black seed (Nigella sativa L.). J Pharm Biomed Anal 19:757–762
Hosseinzadeh H, Parvardeh S (2004) Anticonvulsant effects of thymoquinone, the major constituent of Nigella sativa seeds, in mice. Phytomedicine 11:56–64
Khan N, Sharma S, Sultana S (2003) Nigella sativa (black cumin) ameliorates potassium bromate-induced early events of carcinogenesis: diminution of oxidative stress. Hum Exp Toxicol 22:193–203
Verheije MH, Raaben M, Mari M, Te Lintelo EG, Reggiori F, van Kuppeveld FJ, Rottier PJ, de Haan CA (2008) Mouse hepatitis coronavirus RNA replication depends on GBF1-mediated ARF1 activation. PLoS Pathog 4:e1000088
Ulasli M, Verheije MH, de Haan CA, Reggiori F (2010) Qualitative and quantitative ultrastructural analysis of the membrane rearrangements induced by coronavirus. Cell Microbiol 12:844–861
Verheije MH, Wurdinger T, van Beusechem VW, de Haan CA, Gerritsen WR, Rottier PJ (2006) Redirecting coronavirus to a nonnative receptor through a virus-encoded targeting adapter. J Virol 80:1250–1260
Yoshikawa T, Hill T, Li K, Peters CJ, Tseng CT (2009) Severe acute respiratory syndrome (SARS) coronavirus-induced lung epithelial cytokines exacerbate SARS pathogenesis by modulating intrinsic functions of monocyte-derived macrophages and dendritic cells. J Virol 83:3039–3048
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Ulasli, M., Gurses, S.A., Bayraktar, R. et al. The effects of Nigella sativa (Ns), Anthemis hyalina (Ah) and Citrus sinensis (Cs) extracts on the replication of coronavirus and the expression of TRP genes family. Mol Biol Rep 41, 1703–1711 (2014). https://doi.org/10.1007/s11033-014-3019-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3019-7