Abstract
Genome-wide association studies were conducted using a globally diverse safflower (Carthamus tinctorius L.) Genebank collection for grain yield (YP), days to flowering (DF), plant height (PH), 500 seed weight (SW), seed oil content (OL), and crude protein content (PR) in four environments (sites) that differed in water availability. Phenotypic variation was observed for all traits. YP exhibited low overall genetic correlations (rGoverall) across sites, while SW and OL had high rGoverall and high pairwise genetic correlations (rGij) across all pairwise sites. In total, 92 marker-trait associations (MTAs) were identified using three methods, single locus genome-wide association studies (GWAS) using a mixed linear model (MLM), the Bayesian multi-locus method (BayesR), and meta-GWAS. MTAs with large effects across all sites were detected for OL, SW, and PR, and MTAs specific for the different water stress sites were identified for all traits. Five MTAs were associated with multiple traits; 4 of 5 MTAs were variously associated with the three traits of SW, OL, and PR. This study provided insights into the phenotypic variability and genetic architecture of important safflower agronomic traits under different environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Safflower (Carthamus tinctorius L.) is a member of the Compositae family, grown as a vegetable, cut flower, herbal medicine, animal feed, birdseed, and oilseed, etc. in over 60 geographical regions covering the Middle East, Africa, America, Europe, and Asia (Knowles and Ashri 1995). In recent years, with a growing demand for healthy cooking oil and clean biofuel and bio-lubricants, safflower has emerged as a modern industrial oilseed crop due to its higher oleic and linoleic acid content compared to other oilseed crops (Fernández-Martinez et al. 1993; Khalid et al. 2017). In 2019, FAO data showed safflower seed production world-wide was approximately 0.6 million tonnes, and the top 4 largest growers (Kazakhstan, United States, Russian Federation, and Mexico) produce over 75% of total production (FAO 2019). The Australian safflower growing area is currently about 40,000 ha, down from its peak of 74,688 ha in 1979 (Jochinke et al. 2008). As a potential crop that could grow in a drier environment, safflower is gaining more research attention (Li and Mündel 1996).
To date, genetic analyses for agronomic traits in safflower have largely been undertaken using conventional family-based methods (Kotecha 1979; Ramachandram and Goud 1981). This has allowed the identification of genes and quantitative trait loci (QTL) for traits such as plant height, seed oil content, and days to flowering (Hamdan et al. 2012; Pearl et al. 2014). Association mapping approaches have also been used to identify QTL in safflower. Study using AFLP markers detected four marker-trait associations (MTAs) for PH under drought conditions in safflower (Ebrahimi et al. 2017a). Six MTAs for PH, five MTAs for DF, and several MTAs for oil content, oleic acid content, and linoleic acid content were identified in an association study using microsatellite markers (Ambreen et al. 2018). The Fad2 gene family (Fatty acid desaturases, FAD) in safflower has been sequenced with genes being isolated and cloned (Cao et al. 2013; Wood et al. 2018). However, no genome-wide association studies (GWAS) based on single nucleotide polymorphisms (SNPs) markers have been reported in safflower.
Statistical methods used in GWAS analysis are important for identifying MTAs for complex traits (Wang et al. 2019; Zhang et al. 2010). Single-locus GWAS with mixed linear models (MLM-GWAS) has been widely used to detect the MTAs for agronomic traits in a variety of plants, including wheat (Ledesma-Ramírez et al. 2019), rapeseed (Qu et al. 2017), soybean (Leamy et al. 2017a), etc.. To increase power to discover SNP with small effects and reduce false-positive associations, summary statistic-based methods (meta-GWAS) have been adopted in some studies (Joukhadar et al. 2021; Pasaniuc and Price 2017). In canola, a meta-GWAS analysis identified 79 genomic regions conferring potential candidate resistance to canola blackleg disease, more significant SNPs than single-locus GWAS (Fikere et al. 2020). Differing from single-locus MLM-GWAS testing one marker at a time, multi-locus GWAS have been applied by fitting all loci simultaneously to improve fine-mapping (Kaler et al. 2020; Tamba et al. 2017). As a multi-locus Bayesian method, BayesR simultaneously accommodates all SNPs in the model, and SNPs effects were a mixture of four normal distributions, which include SNPs with 0, small, and moderate effects. In each distribution, fewer SNPs explain the gradually more genetic variance (Daetwyler et al. 2014; Erbe et al. 2012). BayesR has been used to identify QTL or associations in dairy cattle and wheat (Pasam et al. 2017; Xiang et al. 2021).
The variation in phenotypes among genotypes in different environments is evaluated as the extent of the genotype-by-environment interaction (G × E), which is also referred to as the traits phenotypic plasticity (Bradshaw 1965). Identifying G × E interaction patterns and their genetic basis under multi-environment trials can deepen the knowledge of the genetic architecture of traits (Das et al. 2019; Kusmec et al. 2018). In a canola study, 12 environment-stable QTL and 43 environment-specific QTL were detected for flowering time in three different ecological conditions, which provided new insights into the genetic regulatory network underlying the control of flowering time (Li et al. 2018a). Few studies investigating G × E interaction patterns have been reported in safflower, which were carried out to evaluate genotypes and yield stability (Alizadeh et al. 2017; Jamshidmoghaddam and Pourdad 2013).
In Australia, crop production is challenged by spatial drought patterns due to seasonal rainfall and high temperatures (Chenu et al. 2013). Therefore, understanding the G × E interaction and genetic basis underlying grain yield and related agronomic traits are important for safflower breeding. In this study, a globally diverse Genebank collection of 406 accessions was grown in 4 different field environments (2 trials in one location but with different field management in 2017 and 2 locations in 2018). The aims were to (1) assess genetic variability in the different environments and the level of G × E interaction for grain yield and related agronomic traits and (2) identify MTAs via GWAS at each environment to study the genetic basis of the G × E interaction for grain yield and related agronomic traits.
Materials and methods
Plant material and phenotyping
A total of 406 globally diverse safflower accessions were sourced from the Australian Grain Genebank (AGG), including elite cultivars, breeding lines, and landraces. Accessions information and the field trial experiment design are previously described (Zhao et al. 2021). In brief, with a randomized complete block design, all accessions were sown at two field sites in two consecutive years (2017 and 2018, a total of four sites) with plot size of 1 m x 5 m, 5 rows in each plot, and 220 seeds sowed per plot. Sites 1 and 2 were sown in 2017 at the same location (Horsham, Victoria) in a low rainfall zone. Site 1 was flood irrigated before sowing and considered an optimal site with a full soil water profile. Site 2 was rainfed, with soil water stress starting in late spring (during the flowering stage). Sites 3 and 4 were sown in 2018. Site 3 was at the same location as sites 1 and 2 but was rainfed and experienced soil water stress during the entire growing season, with minimal rain in the early spring and high temperature towards the end of the season. Site 4 was in a higher rainfall zone (Wonwondah, Victoria) and received more rain overall than site 3, but also experienced soil water stress.
Days to flowering (DF) was recorded as the number of days from sowing to 25% of the plot flowering. Plant height (PH) was measured at the late flowering stage from the ground surface to the top of the plot canopy in cm. Seed weight (SW) was measured as random 500 achenes from the whole plot in grams. Grain yield was measured as yield per plot (YP) in kilograms harvested by machine. Seed protein (PR) and seed oil content (OL) were determined by near-infrared reflectance spectroscopy (NIR, Foss Pacific Pty Ltd, Denmark) with calibration by the Dumas nitrogen combustion method for protein (TruMac, Leco Corporation St Joseph USA), and the Soxhlet extraction for oil (Soxtec 25,050, FOSS, Hilleroed, Denmark). The NIR prediction models R-squared (R2) and standard error of prediction (SEP) were 0.93 and 0.7% for seed protein content and 0.95 and 1.2% for seed total oil content.
Statistical analysis of phenotypic data
Summary statistics were calculated for each trait at each site. The best linear unbiased estimates (BLUE) for each trait at each site were calculated by a single site linear mixed model with safflower accessions fitted as fixed effects. The model was illustrated as:
where Ymijk is the phenotypes of accessions m in rep j at row i, column k; µ is the overall mean, gm is the fixed accession genetic effect, and Rj is the replicate effect; ri is the row effect, ck is the column effect, and ɛmijk is the residual, including the AR1 × AR1 covariance structure to adjust spatial variation.
Pearson’s correlation at each site was calculated based on the BLUEs of each trait. BLUEs were used as the “phenotypes” for the GWAS.
To assess the G × E level for each trait, the four sites were combined, and the genetic effect associated with accessions was decomposed into two components, the genetic effect of accessions and the interaction effect between accessions and sites (G × E effect), which were assumed to be homogenous for all sites. The linear mixed model was:
where Yijk is the phenotype of accession k in rep j at site i, µ is the overall mean, Si is the fixed i-th site effect, Rj is the fixed replicate effect, Gk is the random accession genetic effect, SGik is the random G × E effect, and ɛijk is the residual. Two models, including and excluding the G × E effect, were compared, and a log-likelihood ratio test was used to test the significance of the G × E effect for each trait (Kendall and Stuart 1979). The genetic correlation among the four sites (rGoverall) was estimated as the ratio of the genetic effect of accessions to total genetic variance, calculated as rGoverall = \({\sigma }_{G}^{2}/({\sigma }_{G}^{2}+{\sigma }_{GE}^{2})\), where \({\sigma }_{G}^{2}\) is the genetic variance of accessions and \({\sigma }_{GE}^{2}\) is the variance for G × E interaction. High genetic correlation among sites indicated low G × E interaction, while low genetic correlation indicated high G × E interaction (Li et al. 2016).
A heterogeneous variance structure was also fitted in the linear mixed model, which assumes that accessions genetic effect is independent at each site. It can be illustrated as:
where the terms are the same as above, with the site as a fixed effect and the accession and trial replicate effects both nested within sites as a random effect with different variance for each site. The residual variance was also nested within the site, with the AR1 × AR1 covariance structure used to adjust spatial variation across columns and rows. The genetic correlation of the accession effect between two sites was calculated as rGij = \({\sigma }_{GiGj}/\sqrt{{\sigma }_{Gi}^{2}{*\sigma }_{Gj}^{2}}\), where the \({\sigma }_{Gi }^{2}\) and \({\sigma }_{Gj }^{2}\) are the variance of the accession genetic effect at sites i and j, respectively. The \({\sigma }_{GiGj}\) is the covariance of the accessions genetic effect at sites i and j. Similar to the above, high genetic correlations between two sites indicated a low G × E interaction. The significance of the genetic correlation between two sites was tested for deviation from 1 using likelihood ratio tests. If rGij significantly differed from 1, it suggested the ranking of accessions at the two sites was different. Akaike information criterion (AIC) was used to compare the fitness of models 2 and 3.
SNP genotyping and population structure
A total of 349 accessions were genotyped using a genotyping-by-sequencing assay as described in (Zhao et al. 2021). In brief, genomic DNA was extracted from six crushed seeds per accession, digested with restriction endonucleases PstI (6-bp cutter) and MseI (4-bp cutter), followed by the amplification, purification, and sequencing by Illumina Hiseq 3000 sequencer. SNP discovery and genotype calling were conducted with custom scripts, and SNPs were filtered for a missing data rate < 30% and minor allele frequency (MAF) > 0.01 and imputed with LinkImpute (Money et al. 2015). A total of 318 samples were passed the filtering, and population structure was evaluated from the genomic relationship matrix (GRM) according to VanRaden (VanRaden 2008). SNPs were further filtered with MAF > 0.05 and heterozygosity < 0.3 for 318 samples for the genome-wide association study. The physical position of the filtered SNP was determined by mapping their flanking sequences to a draft safflower genome assembly (unpublished data) with 12 main scaffolds (pseudochromosomes). Linkage disequilibrium (LD) was calculated for all pairwise SNP using PLINK (Purcell et al. 2007).
Genome-wide association study
Single site GWAS was conducted for each trait using the BLUEs of each trait as the “phenotypes” (Supplementary Table s1). First, a single SNP regression model, referred to as MLM-GWAS, implemented in the GCTA software (Yang et al. 2011), was performed with the GRM fitted to account for population structure. Second, the Bayesian multi-locus approach-BayesR was performed using the Markov chain Monte Carlo (MCMC) method with 50,000 iterations and 25,000 burn-in. SNPs with large effects were declared if they had a nonzero effect with at least a 0.7 posterior probability, averaged over 5 runs (Erbe et al. 2012). And third, meta-GWAS implemented in the software Metal (Willer et al. 2010) was performed for each trait, with each single site MLM-GWAS treated as an independent study. Manhattan and quantile–quantile (Q-Q) plot generated with an R script (Yu et al. 2006) were used to visualize associations for each trait. SNP identified by all three methods were considered candidate MTAs for each trait.
Results
Phenotypic variation and correlations
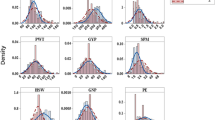
In total, 406 globally diverse safflower accessions were evaluated in four field trials. The phenotypic distributions and means for grain yield (YP), plant height (PH), days to flowering (DF), 500 seed weight (SW), seed protein (PR), and oil content (OL) are shown in Fig. 1. The mean YP was the highest at site 1 (1.89 kg/plot), a third less at site 2 (1.21 kg/plot), and halved at sites 3 and 4 (0.66 and 0.72 kg/plot) (Supplementary Table s2). The distribution for YP was much narrower at sites 3 and 4 compared with sites 1 and 2 for YP (Fig. 1). PH showed a similar distribution pattern to YP, with lower means at sites 3 and 4 (~ 60 cm) compared to sites 1 and 2 (~ 115 cm). DF had the highest mean value and narrowest distribution at site 2 (~ 160 days, Fig. 1). There were differences in trait means between sites for the three seed traits (SW, PR, and OL), but they were more subtle than those observed for YP and PH. The SW mean was higher at sites 3 and 4 (~ 20.6 g) than at sites 1 and 2 (19.56 g and 18.77 g, respectively). The mean of PR ranged from 15.14 to 15.92% across four sites, and OL decreased about 1% with different water stress environments, from 31.83% at site 1 down to 29.69% at site 3. Similarly, the distributions for the three seed traits did not change dramatically across the four sites (Fig. 1, Supplementary Table s2).
Pearson correlations between traits at each site showed that YP was positively correlated with PH (0.34–0.479) and negatively correlated with PR (− 0.208– − 0.405). SW was negatively correlated with OL (− 0.518– − 0.505), PR (− 0.55– − 0.383), and PR was positively correlated with OL (0.19–0.639) over all sites. However, DF is positively correlated with PH at sites 1, 3, and 4 (0.464–0.62) and negatively correlated with PH at site 2 (Table 1).
G × E interaction
G × E interactions for each trait were determined through combined site analysis, and the overall genetic correlations ranged from 0 to 1 among sites for all traits (model 2). The model including G × E interaction effects had a higher log-likelihood than the model excluding G × E interactions, and the G × E interaction effects were all significant (Supplementary Table s3, s4). High overall genetic correlations (rGoverall) among sites were observed for SW and OL (0.95 and 0.94, respectively), indicating low G × E interaction for those traits, while low rGoverall for YP (0.48) indicated a strong G × E interaction. G × E levels were moderate for PH, DF, and PR, with rGoverall value ranging from 0.67 to ~ 0.79 (Supplementary Table s4).
To account for differences in G × E interactions between sites, a linear mixed model assuming heterogeneous genetic and residual variances (model 3) was adopted. The AIC was lower for all traits for model 3 than for model 2, suggesting that model 3 fitted the data better (Supplementary Table s5). The genetic correlation (rGij) between pairwise site combinations varied; however, traits with high rGoverall also had high rGij between pairwise sites (Table 2). According to Robertson (Robertson 1959), a correlation of performance between environments ≤ 0.8 indicates a considerable re-ranking of individuals. SW and OL had high pairwise genetic correlations between all sites with rGij values > 0.9, especially between sites 3 and 4, in which a rGij (0.99) was not significantly different from 1. As the rest traits all had rGij ≥ 0.80 between sites 3 and 4, suggesting that those two sites could be treated as a single site with limited re-ranking. Genetic correlations for PH were uniformly high between pairwise sites (rGij ≥ 0.80), while the genetic correlations for DF and YP varied, with those for YP being the most variable.
Genome-wide associations
The heatmap of the genomic relationship matrix (G) revealed a strong population structure among 318 accessions (Supplementary Figure s1), which was consistent with the previous observation (Zhao et al. 2021). After further filtering, a total of 1806 SNPs were used for GWAS studies, with 1780 positioned on the 12 pseudochromosomes of the draft safflower genome assembly, about 100–200 SNPs per pseudochromosome. LD decayed rapidly over a short physical distance, followed by a slower decline over longer pairwise distances (Supplementary Figure s2).
Combined QQ plots for the four sites showed that the inclusion of G matrix in the GWAS effectively accounted for the observed population structure (Fig. 2, Supplementary Figure s3). A relaxed significance threshold of -log10(p) ≥ 2 was used to denote MTAs in the MLM-GWAS, which resulted in the identification of between 41 and 71 putative MTAs for each trait across the four sites. For the meta-GWAS, the number of significant SNPs was more than twice the number found in the MLM-GWAS for OL, SW, and PR. Fewer MTAs were detected using the BayesR method, especially for PR (Table 3). SNPs with large effects in the BayesR analysis typically overlapped with SNPs above the significance threshold in the MLM-GWAS (Fig. 2, Supplementary Figure s3).
a QQ plot for seed oil content (OL) at the four trial sites; b negative log10 p values are plotted against for all SNPs across four sites for OL, and the blue line indicates the threshold of significance with the MLM-GWAS method; and c SNP effects are plotted against for all SNPs across four sites for OL with the BayesR GWAS method. SNPs are sorted according to physical position in the 12 pseudochromosomes and are in the same order for each of the four trial sites
A total of 92 significant MTAs were detected by all three GWAS approaches (Table 3, Supplementary Tables s6-s11). Heatmap of pairwise LD between significant MTAs for each trait was plotted to show that MTAs were not tightly linked (Supplementary Figure s4). By comparing MLM-GWAS and BayesR results, the significant MTAs were classified as site-specific or shared among sites, where sites 3 and 4 were treated as a single site. Traits with low overall G × E (high genetic correlation) had a higher percentage of shared MTAs across sites (SW and OL), while traits with high to moderate overall G × E (PH, DF, and YP) detected limited shared MTA between pairwise sites. Five and six MTAs shared across all sites were observed for SW and OL, respectively, with Locus 6195 and 28,935 for OL and Locus 3057 and 27,064 for SW showing large effects (> 0.15). PR had three shared MTAs, and locus 3057 with large effects were detected across all sites. Seven MTAs for PH were shared between two sites, and five of seven were between sites 1 and 2. Two and one MTAs were shared between two sites for DF and YP, respectively. All site-specific MTAs were consistent in effect direction but varied in magnitude across sites, except the MTAs for locus 25,179 and 6025 associated with DF and PH, respectively, which had opposite directions with near zero effect at nonsignificant sites. The number of site-specific MTAs observed for each trait differed among sites. 12 out of 18 site-specific SNPs for OL were observed at site 3 (or site 4), six of nine site-specific MTAs for SW were observed at site 1, and eight of 14 site-specific MTAs for DF were observed at site 1 (Table 3, Supplementary Tables s6–11).
Five MTAs had significant associations with more than one trait. Locus 5628 was associated with PH at sites 2 and 3 and DF at site 1. Locus 9819 was associated with DF at site 1, OL at site 3, and PR at sites 2 and 3. Locus 17,302 and 3057 increased SW but decreased PR and OL at site 2 and site 3 (or site 4), respectively. Locus 28,935 was associated with low SW and high OL at sites 2 and 3 (or site 4) (Table 4).
Discussion
Understanding G × E is an important initial step to developing strategies for a breeding program in the target environment(s). Our results showed that G × E patterns differed between safflower traits. The identification of site-shared and site-specific MTAs in GWAS provides knowledge to broaden our understanding of the genetic basis of G × E interactions for important safflower traits.
Different G × E interaction patterns were observed for safflower traits
The seasonal rainfall in the Victoria Wimmera region (Horsham and Wonwondah) had an impact on safflower agronomic traits. Safflower is normally sown in Winter in Australia to maximize the usage of the available water from Winter and early Spring rain (Wachsmann et al. 2008). In our study, we observed that water stress during flowering decreased safflower grain yield substantially. Further, insufficient Spring rain heavily reduced safflower production via poor biomass accumulation. Similar grain yield instability induced by rainfall patterns has also been reported in other crops (Sadras and Dreccer 2015). Besides grain yield, a 1% decrease of OL was observed under differing water stress, consistent with previous studies that oil content decreased under drought stresses (Ebrahimi et al. 2017b; Joshan et al. 2019). The positive correlation between OL and PR (0.14 to ~ 0.46 across sites), which was also reported in a previous study (r = 0.476) (Oz 2016), indicated that artificial selection for OL in safflower has a limited impact on seed protein compared with soybean (Leamy et al. 2017b). The negative correlation between SW with PR and OL across all four sites suggests a negative relationship between carbohydrate accumulation and protein and oil accumulation, which could be similar to the competition in cereal crops (Bjarnason and Vasal 1992; Pasam et al. 2012).
The overall G × E interactions were significant for all the traits studied. However, there were different levels of G × E for the different traits. The heterogeneous model further revealed detailed G × E patterns for each trait, which indicated the rank changes of accessions between sites. The high G × E observed for YP across sites was consistent with studies in other crops (He et al. 2019; Tolessa et al. 2013). Low to moderate pairwise genetic correlation indicated re-ranking for YP was high among sites. Only 4 accessions showed yield stability through presence in the top 50 high yield accessions across all sites, which could be used for the future breeding program. The cultivars and breeding lines performed well at site 1 (19 accessions out of the 50 top YP accessions) but not at the other three sites, suggesting that introgression of water stress tolerance from landraces could improve safflower yield stability. The low level of G × E for OL with limited re-ranking across sites observed in our study was also indicated in a soybean study (Sudarić et al. 2006). According to the BLUEs for each site, about half of the top 30 accessions with high OL were cultivars and breeding lines, reflecting breeding efforts to improve OL in safflower cultivars. Although a moderate rGoverall was observed for DF and PH, the pairwise genetic correlation showed that lines were reranked strongly for DF at site 2 compared with other sites. The genetic divergence of DF among the accessions in response to water stress at flowering implied DF is important in developing drought tolerant varieties (Bhandari et al. 2020).
GWAS identified MTAs for safflower traits
GWAS has been widely used to study the genetic basis of the important agronomy traits with diverse germplasm in crops (Liu and Yan 2019). Multi-environment trials normally were combined to present the overall phenotypic variation for GWAS to detecting the associations between markers and traits (Landers and Stapleton 2014; Leamy et al. 2017c). However, with diverse germplasm, the phenotypic variation displayed under differed environments can be used to measure the plasticity of the traits or trait G × E level with proper statistical models (Des Marais et al. 2013; Malosetti et al. 2013). Environmental stable and environmental-specific MTAs can help our understanding of the genetic basis of trait G × E, and it also will enrich our knowledge of the genetic architecture of the important agronomy traits (Li et al. 2018b; Timpson et al. 2018). In our study, GWAS was carried out with a globally diverse safflower collection for six agronomic traits in four field trials that differed with water availability. MTAs shared across sites were identified for traits with low G × E, and site-specific MTAs were discovered for all traits with more site-specific MTAs than shared MTAs identified for moderate overall G × E traits.
A high number of significant MTAs were identified for seed oil content (OL) by all three GWAS approaches, of which 18 were shared across sites, and 12 were site-specific, indicating the complex genetic control of this trait. Studies with canola showed that 24 candidate genes were involved in fatty acid biosynthesis (Qu et al. 2017). In safflower, a transcriptome study showed that a significant number of transcription factors were involved in oil accumulation in safflower seeds (Li et al. 2021). The six MTAs shared across four sites will be of interest to safflower breeders and geneticists as sources of genetic variation to improve the seed oil content in safflower under different growing conditions. Similarly, numerous MTA (total 20) were identified for seed weight (SW), of which 11 were shared across sites. Three MTAs explaining more than 10–20% phenotypic variance across sites will provide useful information for breeders to modify SW in safflower (Supplementary Table s9).
The molecular basis for G × E interactions could be due to site-specific QTLs, gene expression, or differences in the magnitude of expression across environments (Des Marais et al. 2013; Li et al. 2018b). In our study, all site-specific MTAs showed differing allelic effects across sites for each trait (Supplementary Table s6–11); however, the effects were significant in some environments but not in other environments. We observed moderate overall G × E for PH and DF with a higher number of site-specific MTAs. Markers associated with PH and DF under drought conditions in safflower have been reported (Ebrahimi et al. 2017a). Only one MTA was identified for DF at site 2, which could be related to the narrow phenotypic variation observed at site 2 (Stich and Melchinger 2010). Few MTAs for PR and YP were detected by all three GWAS methodologies. However, those that were identified explained a high proportion of the phenotypic variation for each trait, indicating their potential importance for genetic improvement.
Correlations between traits can be caused by pleiotropy or a close linkage of loci associated with the traits (Chen and Lübberstedt 2010). Shared major genes or QTL for flowering time and plant height have been reported in soybean (Cober and Morrison 2010; Fang et al. 2017). In our study, locus 5628 was associated with DF at site 1 and PH at site 2 and 3, suggesting the MTA is likely tightly linked to different QTL affecting both traits, rather than being a single QTL with pleiotropic effects. In canola, a QTL affecting both OL and PR in repulsion was reported, suggesting the PR and OL biosynthesis pathways interfere and/or compete with one another (Chao et al. 2017). In maize, high OL and high PR were achieved using the opaque2 modifier genes. However, a yield reduction was noted (Vanous et al. 2019). In our study, we identified four MTAs affecting three traits, one MTA influencing both SW and OL, one MTA associated with PR and OL, and two MTAs interfering SW, OL, and PR. The allelic effects of those MTAs were consistent with the correlation observed in the field among the three traits that PR and OL are positively correlated, and both traits are negatively correlated with SW. This suggested that safflower breeding for PR and OL may differ from canola and maize. However, balancing seed weight and seed quality (OL and PR) would be a challenge. There were other strong phenotypic correlations, such as YP with PH and YP with PR, but associated markers were not identified. The reason could be the low number of significant SNPs that were observed for grain yield.
The interplay of GWAS results and genetic architectures
SW and OL are known as highly heritable traits in many crops, while yield is more quantitative in nature (Ward et al. 2019; Xiao et al. 2019). The number of MTAs identified by the three GWAS methods did not fully reflect the complexity of the trait genetic architecture. One reason for this could be the thresholds used by the three methods. The p value used in our single locus MLM-GWAS was relaxed, and a significant number of candidate MTAs were observed for all traits. With meta-GWAS, we reported significant SNPs number with -log10P value at 3 instead of 2, which detected more significant SNPs for each trait indicating the increased power (Supplementary Table s12). However, multi-locus BayesR, which can improve association mapping resolution by removing multiple SNPs being in LD with the same QTL, could detect SNPs with larger effects (Kemper et al. 2015; Pasam et al. 2017). We observed fewer MTAs for all traits with the BayesR methods with the arbitrary threshold of 0.7 posterior probability of a SNP having an effect. This threshold may have been too stringent for polygenic traits such as YP and PR. Only 10 MTAs associated with YP, and 5 MTAs associated with PR were detected with BayesR, which explained 5–28% of the phenotypic variance.
Heterogenous model fit the data better
Mixed linear models are widely used for G × E analysis in crop research (Smith et al. 2005). Falconer and Mackay (1996) suggested that the same trait measured in different environments should be considered as different (but correlated) traits. In our study, the homogenous model 2 combined the four sites together and estimated the overall GXE pattern with only three parameters. However, the heterogeneous model 3 treated each site as an independent environment, and a total of 26 parameters were estimated. The increased number of parameters allowed dissection of G × E among individual environments to reveal hidden patterns of genetic correlation between sites. Furthermore, the AIC, BIC, and Logl were improved for all traits, indicating model 3 was better able to fit the data (Hirotugu. 1998). These findings agreed with Malosetti et al. (2013), who compared different models to study G × E interactions and concluded that sophisticated mixed models are necessary to allow for heterogeneity of genetic variances and correlations across environments.
In conclusion, two mixed linear models were applied to analyse the G × E pattern for grain yield (YP), days to flowering (DF), plant height (PH), 500 seed weight (SW), seed oil content (OL), and seed protein content (PR) in a globally diverse safflower collection grown in four field trials. The heterogenous mixed linear model (MLM) fitted data better and provided a detailed estimation of the G × E pattern. We observed that different water stress conditions impacted the performance of each of these traits differently, with low overall G × E observed for OL and SW and high overall G × E for YP. In total, 92 MTAs were identified with large effects MTAs detected for OL, SW, and PR across all sites. Site-specific MTAs were detected for all traits with differed allelic effects, suggesting these MTAs could be associated with trait G × E. Five MTAs were associated with multiple traits. The uniform GWAS thresholds used in the study could have impacted the number of significant SNP identified for complex traits. This study has provided new insights into the genetic architecture of the traits studied, and it presents opportunities to exploit the MTA identified in breeding programs to increase yield stability and local adaptation in safflower.
Data Availability
The phenotypic datasets supporting the conclusions of this article are included within the article and the attached additional files. And the genotype dataset, please see the previously published paper: Genomic prediction and genomic heritability of yield-related traits in Safflower https://doi.org/10.1002/tpg2.20064.
References
Alizadeh K, Mohammadi R, Shariati A, Eskandari M (2017) Comparative analysis of statistical models for evaluating of genotype x environment interaction in rainfed safflower. Agricultural Research 6
Ambreen H, Kumar S, Kumar A, Agarwal M, Jagannath A, Goel S (2018) Association Mapping for Important Agronomic Traits in Safflower (Carthamus tinctorius L.) Core collection using microsatellite markers. Frontiers in Plant Science 9
Bhandari A, Sandhu N, Bartholome J, Cao-Hamadoun T-V, Ahmadi N, Kumari N, Kumar A (2020) Genome-wide association study for yield and yield related traits under reproductive stage drought in a diverse indica-aus rice panel. Rice 13:53
Bjarnason M, Vasal SK (1992) Breeding of Quality Protein Maize (QPM). Plant Breeding Reviews, pp 181–216
Bradshaw AD (1965) Evolutionary significance of phenotypic plasticity in plants. Adv Genet 13:115–155
Cao S, Zhou XR, Wood CC, Green AG, Singh SP, Liu L, Liu Q (2013) A large and functionally diverse family of Fad2 genes in safflower (Carthamus tinctorius L.). BMC Plant Biol 13:5
Chao H, Wang H, Wang X, Guo L, Gu J, Zhao W, Li B, Chen D, Raboanatahiry N, Li M (2017) Genetic dissection of seed oil and protein content and identification of networks associated with oil content in Brassica napus. Sci Rep 7:46295
Chen Y, Lübberstedt T (2010) Molecular basis of trait correlations. Trends Plant Sci 15:454–461
Chenu K, Deihimfard R, Chapman SC (2013) Large-scale characterization of drought pattern: a continent-wide modelling approach applied to the Australian wheatbelt–spatial and temporal trends. New Phytol 198:801–820
Cober ER, Morrison MJ (2010) Regulation of seed yield and agronomic characters by photoperiod sensitivity and growth habit genes in soybean. Theor Appl Genet 120:1005–1012
Daetwyler HD, Bansal U, Bariana H, Hayden M, Hayes B (2014) Genomic prediction for rust resistance in diverse wheat landraces. TAG Theoretical and applied genetics Theoretische und angewandte Genetik 127
Das A, Parihar AK, Saxena D, Singh D, Singha KD, Kushwaha KPS, Chand R, Bal RS, Chandra S, Gupta S (2019) Deciphering genotype-by- environment interaction for targeting test environments and rust resistant genotypes in field pea (Pisum sativum L.). Frontiers in Plant Science 10
Des Marais DL, Hernandez KM, Juenger TE (2013) Genotype-by-environment interaction and plasticity: exploring genomic responses of plants to the abiotic environment. Annu Rev Ecol Evol Syst 44:5–29
Ebrahimi F, Majidi MM, Arzani A, Mohammadi-Nejad G (2017a) Association analysis of molecular markers with traits under drought stress in safflower. Crop Pasture Sci 68:167–175
Ebrahimi F, Majidi MM, Arzani A, Mohammadi-Nejad G (2017b) Association analysis of molecular markers with traits under drought stress in safflower. Crop Pasture Sci 68(167–175):169
Erbe M, Hayes BJ, Matukumalli LK, Goswami S, Bowman PJ, Reich CM, Mason BA, Goddard ME (2012) Improving accuracy of genomic predictions within and between dairy cattle breeds with imputed high-density single nucleotide polymorphism panels. J Dairy Sci 95:4114–4129
Falconer DS, Mackay TFC (1996) Introduction to Quantitative Genetics, 4th Edition, 4th edition edn. Longman Group Ltd
Fang C, Ma Y, Wu S, Liu Z, Wang Z, Yang R, Hu G, Zhou Z, Yu H, Zhang M, Pan Y, Zhou G, Ren H, Du W, Yan H, Wang Y, Han D, Shen Y, Liu S, Liu T, Zhang J, Qin H, Yuan J, Yuan X, Kong F, Liu B, Li J, Zhang Z, Wang G, Zhu B, Tian Z (2017) Genome-wide association studies dissect the genetic networks underlying agronomical traits in soybean. Genome Biol 18:161
FAO (2019)
Fernández-Martinez J, del Rio M, de Haro A (1993) Survey of safflower (Carthamus tinctorius L.) germplasm for variants in fatty acid composition and other seed characters. Euphytica 69:115–122
Fikere M, Barbulescu DM, Malmberg MM, Spangenberg GC, Cogan NOI, Daetwyler HD (2020) Meta-analysis of GWAS in canola blackleg (Leptosphaeria maculans) disease traits demonstrates increased power from imputed whole-genome sequence. Sci Rep 10:14300
Hamdan Y, García-Moreno M, Fernández-Martínez J, Velasco L, Pérez-Vich B (2012) Mapping of major and modifying genes for high oleic acid content in safflower. Mol Breeding 30:1279–1293
He S, Thistlethwaite R, Forrest K, Shi F, Hayden MJ, Trethowan R, Daetwyler HD (2019) Extension of a haplotype-based genomic prediction model to manage multi-environment wheat data using environmental covariates. Theor Appl Genet 132:3143–3154
Hirotugu A (1998) Information theory and an extension of the maximum likelihood principle. In: Parzen E. TK, Kitagawa G. (ed) Selected Papers of Hirotugu Akaike Springer Series in Statistics (Perspectives in Statistics). Springer, New York, NY
Jamshidmoghaddam M, Pourdad SS (2013) Genotype × environment interactions for seed yield in rainfed winter safflower (Carthamus tinctorius L.) multi-environment trials in Iran. Euphytica 190:357–369
Jochinke D, Wachsmann N, Potter T, Norton R (2008) Growing safflower in Australia: Part 1 - History, experiences and current constraints on production. The 7th international safflower conference Waga Wagga, Australia
Joshan YSB, Jabbari H, Mozafari H, Moaveni P (2019) Effect of drought stress on oil content and fatty acids composition of some safflower genotypes. Plant Soil Environ 65:4
Joukhadar R, Thistlethwaite R, Trethowan R, Keeble-Gagnère G, Hayden MJ, Ullah S, Daetwyler HD (2021) Meta-analysis of genome-wide association studies reveal common loci controlling agronomic and quality traits in a wide range of normal and heat stressed environments. Theor Appl Genet 134:2113–2127
Kaler AS, Gillman JD, Beissinger T, Purcell LC (2020) Comparing different statistical models and multiple testing corrections for association mapping in soybean and maize. Frontiers in Plant Science 10
Kemper KE, Reich CM, Bowman PJ, vander Jagt CJ, Chamberlain AJ, Mason BA, Hayes BJ, Goddard ME, (2015) Improved precision of QTL mapping using a nonlinear Bayesian method in a multi-breed population leads to greater accuracy of across-breed genomic predictions. Genet Sel Evol 47:29
Kendall MG, Stuart A (1979) The advantage theory of statistics Griffin and Co., London
Khalid N, Khan RS, Hussain MI, Farooq M, Ahmad A, Ahmed I (2017) A comprehensive characterisation of safflower oil for its potential applications as a bioactive food ingredient - a review. Trends Food Sci Technol 66:176–186
Knowles P, Ashri A (1995) Safflower -- Carthamus Tinctorius (Compositae). In: Smartt J, Simmonds NW (eds) Evolution of Crop Plants. Longman, Harlow, UK
Kotecha A (1979) Inheritance and Association of Six Traits in Safflower1. Crop Science 19:cropsci1979.0011183X001900040022x
Kusmec A, de Leon N, Schnable PS (2018) Harnessing phenotypic plasticity to improve maize yields. Frontiers in Plant Science 9
Landers DA, Stapleton AE (2014) Genetic interactions matter more in less-optimal environments: a focused review of “phenotype uniformity in combined-stress environments has a different genetic architecture than in single-stress treatments” (Makumburage and Stapleton, 2011). Frontiers in Plant Science 5
Leamy LJ, Zhang H, Li C, Chen CY, Song B-H (2017a) A genome-wide association study of seed composition traits in wild soybean (Glycine soja). BMC Genomics 18:18
Ledesma-Ramírez L, Solís-Moya E, Iturriaga G, Sehgal D, Reyes-Valdes MH, Montero-Tavera V, Sansaloni CP, Burgueño J, Ortiz C, Aguirre-Mancilla CL, Ramírez-Pimentel JG, Vikram P, Singh S (2019) GWAS to identify genetic loci for resistance to yellow rust in wheat pre-breeding lines derived from diverse exotic crosses. Frontiers in Plant Science 10
Li Y, Wilcox P, Telfer E, Graham N, Stanbra L (2016) Association of single nucleotide polymorphisms with form traits in three New Zealand populations of radiata pine in the presence of genotype by environment interactions. Tree Genet Genomes 12:63
Li B, Zhao W, Li D, Chao H, Zhao X, Ta N, Li Y, Guan Z, Guo L, Zhang L, Li S, Wang H, Li M (2018a) Genetic dissection of the mechanism of flowering time based on an environmentally stable and specific QTL in Brassica napus. Plant Sci 277:296–310
Li X, Guo T, Mu Q, Li X, Yu J (2018b) Genomic and environmental determinants and their interplay underlying phenotypic plasticity. Proc Natl Acad Sci 115:6679
Li D, Mündel HH (1996) Safflower, Carthamus tinctorius L. promoting the conservation and use of underutilized and neglected crops 7. Institute of Plant Genetics and Crop Plant Research, Gatersleben/International Plant Genetic Resources Institute, Rome, Italy
Li D, Wang Q, Xu X, Yu J, Chen Z, Wei B, Wu W (2021) Temporal transcriptome profiling of developing seeds reveals candidate genes involved in oil accumulation in safflower (Carthamus tinctorius L.). BMC Plant Biol 21:181
Liu HJ, Yan J (2019) Crop genome-wide association study: a harvest of biological relevance. Plant J 97:8–18
Malosetti M, Ribaut J-M, van Eeuwijk FA (2013) The statistical analysis of multi-environment data: modeling genotype-by-environment interaction and its genetic basis. Frontiers in Physiology 4
Money D, Gardner K, Migicovsky Z, Schwaninger H, Zhong GY, Myles S (2015) LinkImpute: fast and accurate genotype imputation for nonmodel organisms. G3 5:2383–2390
Oz M (2016) Relationship between Sowing Time, Variety, and Quality in Safflower. J Chem 2016:9835641
Pasam RK, Sharma R, Malosetti M, van Eeuwijk FA, Haseneyer G, Kilian B, Graner A (2012) Genome-wide association studies for agronomical traits in a world wide spring barley collection. BMC Plant Biol 12:16
Pasam RK, Bansal U, Daetwyler HD, Forrest KL, Wong D, Petkowski J, Willey N, Randhawa M, Chhetri M, Miah H, Tibbits J, Bariana H, Hayden MJ (2017) Detection and validation of genomic regions associated with resistance to rust diseases in a worldwide hexaploid wheat landrace collection using BayesR and mixed linear model approaches. Theor Appl Genet 130:777–793
Pasaniuc B, Price AL (2017) Dissecting the genetics of complex traits using summary association statistics. Nat Rev Genet 18:117–127
Pearl SA, Bowers JE, Reyes-Chin-Wo S, Michelmore RW, Burke JM (2014) Genetic analysis of safflower domestication. BMC Plant Biol 14:43
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Qu C, Jia L, Fu F, Zhao H, Lu K, Wei L, Xu X, Liang Y, Li S, Wang R, Li J (2017) Genome-wide association mapping and Identification of candidate genes for fatty acid composition in Brassica napus L. using SNP markers. BMC Genomics 18:232
Ramachandram M, Goud JV (1981) Genetic analysis of seed yield, oil content and their components in safflower (Carthamus tinctorius L.). Theor Appl Genet 60:191–195
Robertson A (1959) The sampling variance of the genetic correlation coefficient. Biometrics 15:469–485
Sadras V, Dreccer MF (2015) Adaptation of wheat, barley, canola, field pea and chickpea to the thermal environments of Australia. Crop Pasture Sci 66:1137–1150
Smith AB, Cullis BR, Thompson R (2005) The analysis of crop cultivar breeding and evaluation trials: an overview of current mixed model approaches. J Agric Sci 143:449–462
Stich B, Melchinger A (2010) An introduction to association mapping in plants. Cab Reviews: Perspectives in Agriculture, Veterinary Science, Nutrition and Natural Resources, v5, 1-9 (2010) 5
Sudarić A, ŠImić D, Vratarić M, (2006) Characterization of genotype by environment interactions in soybean breeding programmes of southeast Europe. Plant Breeding 125:191–194
Tamba CL, Ni Y-L, Zhang Y-M (2017) Iterative sure independence screening EM-Bayesian LASSO algorithm for multi-locus genome-wide association studies. PLOS Comput Biol 13:e1005357
Timpson NJ, Greenwood CMT, Soranzo N, Lawson DJ, Richards JB (2018) Genetic architecture: the shape of the genetic contribution to human traits and disease. Nat Rev Genet 19:110–124
Tolessa T, Keneni G, Gela TS, Jarso M, Bekele Y (2013) Genotype × Environment Interaction and Performance Stability for Grain Yield in Field Pea (Pisum sativum L.) Genotypes. International Journal of Plant Breeding 7:116–123
Vanous A, Gardner C, Blanco M, Martin-Schwarze A, Wang J, Li X, Lipka AE, Flint-Garcia S, Bohn M, Edwards J, Lübberstedt T (2019) Stability analysis of kernel quality traits in exotic-derived doubled haploid maize lines. Plant Genome 12:170114
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423
Wachsmann NJD, Potter T, Norton R (2008) Growing safflower in Australia: part 2 - agronomic research and suggestions to increase yields and production. In: Knights SEP, T.D. (ed) Safflower: unesploited potential and world adaptability 7th International Safflower Conference, . Agri-MC marketing and Communication, Wagga Wagga, New South Wales, Australia., pp 1–8
Wang MH, Cordell HJ, Van Steen K (2019) Statistical methods for genome-wide association studies. Semin Cancer Biol 55:53–60
Ward BP, Brown-Guedira G, Kolb FL, Van Sanford DA, Tyagi P, Sneller CH, Griffey CA (2019) Genome-wide association studies for yield-related traits in soft red winter wheat grown in Virginia. PLoS ONE 14:e0208217–e0208217
Willer CJ, Li Y, Abecasis GR (2010) METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26:2190–2191
Wood CC, Okada S, Taylor MC, Menon A, Mathew A, Cullerne D, Stephen SJ, Allen RS, Zhou XR, Liu Q, Oakeshott JG, Singh SP, Green AG (2018) Seed-specific RNAi in safflower generates a superhigh oleic oil with extended oxidative stability. Plant Biotechnol J 16:1788–1796
Xiang R, Breen EJ, Prowse-Wilkins CP, Chamberlain AJ, Goddard ME (2021) Bayesian genome-wide analysis of cattle traits using variants with functional and evolutionary significance. bioRxiv:2021.2005.2005.442705
Xiao Z, Zhang C, Tang F, Yang B, Zhang L, Liu J, Huo Q, Wang S, Li S, Wei L, Du H, Qu C, Lu K, Li J, Li N (2019) Identification of candidate genes controlling oil content by combination of genome-wide association and transcriptome analysis in the oilseed crop Brassica napus. Biotechnol Biofuels 12:216
Yang J, Lee SH, Goddard ME, Visscher PM (2011) GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 88:76–82
Yu J, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhang Z, Ersoz E, Lai CQ, Todhunter RJ, Tiwari HK, Gore MA, Bradbury PJ, Yu J, Arnett DK, Ordovas JM, Buckler ES (2010) Mixed linear model approach adapted for genome-wide association studies. Nat Genet 42:355–360
Zhao H, Li Y, Petkowski J, Kant S, Hayden MJ, Daetwyler HD (2021) Genomic prediction and genomic heritability of grain yield and its related traits in a safflower genebank collection. The Plant Genome n/a:e20064
Acknowledgements
The authors thank the Australian Grains Genebank for providing seed for the safflower
accessions. The collective efforts of the Agriculture Victoria field and lab technical staff are most
appreciated.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions This study was funded by Agriculture Victoria Research, Victoria state government, Australia.
Author information
Authors and Affiliations
Contributions
HZ, MH, and HD conceived and designed the experiment. PM, JT, SK, and MH performed and supervised the phenotyping, genotyping, and genome assembly; KS performed SNP annotation and previewed the manuscript; YL assisted with the statistical analysis; EB assisted with BayesR analysis; HZ performed the data analysis and wrote the manuscript; MH and HD supervised the study and data interpretation. All authors revised the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, H., Savin, K.W., Li, Y. et al. Genome-wide association studies dissect the G × E interaction for agronomic traits in a worldwide collection of safflowers (Carthamus tinctorius L.). Mol Breeding 42, 24 (2022). https://doi.org/10.1007/s11032-022-01295-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-022-01295-8