Abstract
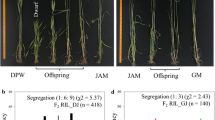
Gibberellin-sensitive dwarfing gene Rht18 was mapped in two durum wheat recombinant inbred lines (RIL) populations developed from crosses, Bijaga Yellow/Icaro and HI 8498/Icaro. Rht18 was mapped within genetic interval of 1.8 cM on chromosome 6A. Simple sequence repeat (SSR) markers S470865SSR4, barc37 and TdGA2ox-A9 specific marker showed co-segregation with Rht18 in Bijaga Yellow/Icaro population consisting 256 RILs. Effect of Rht18 on plant height was validated in HI 8498/Icaro RIL population which segregated for Rht18 and Rht-B1b. Rht-B1b from HI 8498 showed pleiotropic effect on plant height and coleoptile length, on the other hand, Rht18 did not show effect on coleoptile length. The SSR and SNP markers linked to Rht18 were also validated by assessing their allelic frequency in 89 diverse durum and bread wheat accessions. It was observed that 204 bp allele of S470865SSR4 could differentiate Icaro from rest of the wheat accessions except HI 8498, suggesting its utility for selection of Rht18 in wheat improvement programs. Rht18 associated alleles of TdGA2ox-A9, IAW4371 and IAW7940 were absent in most of the tall Indian local durum wheat and bread wheat, hence could be used to transfer Rht18 to bread wheat and local durum wheat. SSR marker barc3 showed high recombination frequency with Rht18, though it showed allele unique to Icaro. Since semidwarf wheat with GA-sensitive dwarfing genes are useful in dry environments owing to their longer coleoptile, better emergence and seedling vigor, Rht18 may provide a useful alternative to widely used GA-insensitive dwarfing genes under dry environments.



Similar content being viewed by others
References
Allan RE, Vogel OA, Peterson CJ (1962) Seedling emergence rate of fall sown wheat and its association with plant height and coleoptile length. Agron J 54:347–350
Appelford NEJ, Lenton JR (1991) Gibberellins and leaf expansion in near-isogenic wheat lines containing Rht1 and Rht3 dwarfing alleles. Planta 183:229–236
Bai C, Liang Y, Hawkesford MJ (2013) Identification of QTLs associated with seedling root traits and their correlation with plant height in wheat. J Exp Bot 64:1745–1753
Bhagwat SG, Bhatia CR (1994) Quantitative differences in the gibberellin induced alpha amylase activity from aleurone layers of tall and semidwarf wheat cultivars. Cereal Res Commun 22:129–134
Botwright TL, Rebetzke GJ, Condon AG, Richards RA (2001) The effect of rht genotype and temperature on coleoptile growth and dry matter partitioning in young wheat seedlings. Aust J Plant Physiol 28:417–423
Chen L, Phillips AL, Condon AG, Parry MAJ, Hu Y-G (2013) GA-responsive dwarfing gene Rht12 affects the developmental and agronomic traits in common bread wheat. PLoS One 8:e62285. doi:10.1371/journal.pone.0062285
Ellis MH, Rebetzke GJ, Azanza F, Richards RA, Spielmeyer W (2005) Molecular mapping of gibberellin-responsive dwarfing genes in bread wheat. Theor Appl Genet 111:423–430
Ellis MH, Spielmeyer W, Gale KR, Rebetzke GJ, Richards RA (2002) Perfect markers for the Rht-B1b and Rht-D1b dwarfing genes in wheat. Theor Appl Genet 105:1038–1042
Griffiths S, Simmonds J, Leverington M, Wang Y, Fish L et al (2012) Meta-QTL analysis of the genetic control of crop height in elite European winter wheat germplasm. Mol Breeding 29:159–171
Haque MA, Martinek P, Watanabe N, Kuboyama T (2011) Genetic mapping of gibberellic acid-sensitive genes for semidwarfism in durum wheat. Cereal Res Commun 39:171–178
Hedden P (2003) The genes of the Green Revolution. Trends Genet 19:5–9
Hedden P, Phillips AL (2000) Gibberellin metabolism: new insights revealed by the genes. Trends Plant Sci 5:523–530
Konzak CF (1987) Mutations and mutation breeding. In: Heyne EC (ed) Wheat and wheat improvement—Agronomy Monograph no. 13, 2nd edn. American Society of Agronomy, Madison, pp 428–443
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MapMaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Li P, Chen J, Wu P, Zhang J, Chu C, See D, Brown-Guedira G, Zemetra R, Souza E (2011) Quantitative trait loci analysis for the effect of Rht-B1 dwarfing gene on coleoptile length and seedling root length and number of bread wheat. Crop Sci 51:2561–2568
Lo SF, Yang SY, Chen KT, Hsing YI, Zeevaart JA, Chen LJ, Yu SM (2008) A novel class of gibberellin 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 20:2603–2618
Maccaferri M, Ricci A, Salvi S, Milner SG, Noli E, Martelli P, Casadio R, Akhunov E, Scalabrin S, Vendramin V, Ammar K, Blanco A, Desiderio F, Distelfeld A, Dubcovsky J, Fahima T, Faris J, Korol A, Massi A, Mastrangelo AM, Morgante M, Pozniak C, N’Diaye A, Xu S, Tuberosa R (2015) A high-density, SNP-based consensus map of tetraploid wheat as a bridge to integrate durum and bread wheat genomics and breeding. Plant Biotechnol J 13:648–663
Maluszynski M, Szarejko I (2003) Induced mutations in the Green and Gene Revolutions. In: Tuberosa R, Phillips RL, Gale M (eds) Proceedings of the international congress bologna, Italy, ‘in the wake of the double helix: from the Green Revolution to the gene revolution’. ©2005 Avenue media, Bologna, pp 403–425
Pearce S, Huttly AK, Prosser IM, Li Y, Vaughan SP, Gallova B, Patil A, Coghill JA, Dubcovsky J, Hedden P, Phillips AL (2015) Heterologous expression and transcript analysis of gibberellin biosynthetic genes of grasses reveals novel functionality in the GA3ox family. BMC Plant Biol 15:130. doi:10.1186/s12870-015-0520-7
Ramkumar G, Prahalada GD, Hechanova SL, Vinarao R, Jena KK (2015) Development and validation of SNP-based functional codominant markers for two major disease resistance genes in rice (O. sativa L.) Mol Breeding 35:129. doi:10.1007/s11032-015-0323-4
Rebetzke GJ, Richards RA (2000) Gibberellic acid-sensitive dwarfing genes reduce plant height to increase kernel number and grain yield of wheat. Aust J Agric Res 51:235–245
Rebetzke GJ, Appels R, Morrison AD, Richards RA, McDonald G, Ellis MH, Spielmeyer W, Bonnett DG (2001) Quantitative trait loci on chromosome 4B for coleoptile length and early vigour in wheat. Aust J Agric Res 52:1221–1234
Rebetzke GJ, Bruce S, Kirkegaard JA (2005) Genotypic increases in coleoptile length improves emergence and early vigour with crop residues. Plant Soil 270:87–100
Rebetzke GJ, Ellis MH, Bonnett DG, Condon AG, Falk D, Richards RA (2011) The Rht13 dwarfing gene reduces peduncle length and plant height to increase grain number and yield of wheat. Field Crop Res 124:323–331
Rebetzke GJ, Richards RA, Fettel NA, Long M, Condon AG, Forrester RI, Botwright TL (2007) Genotypic increases in coleoptile length improves stand establishment, vigour and grain yield of deep-sown wheat. Field Crops Res 100:10–12
Rebetzke GJ, Ellis MH, Bonnett DG, Mickelson B, Condon AG, Richards RA (2012) Height reduction and agronomic performance for selected gibberellin-responsive dwarfing genes in bread wheat (Triticum aestivum L.) Field Crops Res 126:87–96
Richards RA (1992) The effect of dwarfing genes in spring wheat in dry environments II. Growth, water use and water use efficiency. Aust J Agric Res 43:529–539
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76
Spielmeyer W, Hyles J, Joaquim P, Azanza F, Bonnett D, Ellis ME, Moore C, Richards RA (2007) A QTL on chromosome 6A in bread wheat (Triticum aestivum) is associated with longer coleoptiles, greater seedling vigour and final plant height. Theor Appl Genet 115:59–66
Voorrips RE (2002) Mapchart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wuddineh WA, Mazarei M, Zhang J-Y, Poovaiah CR, Mann DGJ, Ziebell A, Sykes RW, Davis MF, Udvardi MK, Stewart CN Jr (2015) Identification and overexpression of gibberellic acid 2-oxidase (GA2ox) in switchgrass (Panicum virgatum L.) for improved plant architecture and reduced biomass recalcitrance. Plant Biotechnol J 13:636–647
Yang Z, Zheng J, Liu C, Wang Y, Condon AG, Chen Y, Hu Y (2015) Effects of the GA-responsive dwarfing gene Rht18 from tetraploid wheat on agronomic traits of common wheat. Field Crop Res 183:92–101
Acknowledgements
Authors wish to thank Dr. M. D. Bhagwat for initiating development of mapping populations and useful suggestions. Authors are grateful to Dr. Andrew Phillips, Rothamsted Research, Harpenden, UK, for providing scaffolds containing sequences of three homoeologues of TaGA2ox9. Authors wish to thank three anonymous reviewers for constructive suggestions on the manuscript. Financial support by the Science and Engineering Research Board, Department of Science and Technology, New Delhi under Start-Up Research grants for Young Scientists-SB/FT/LS-243/2012 to RP is gratefully acknowledged. The Junior Research Fellowship by Agharkar Research Institute to PV under project GEN15 is also gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The research was partially funded by the Science and Engineering Research Board, Department of Science and Technology, New Delhi (SB/FT/LS-243/2012) and the Agharkar Research Institute. The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Vikhe, P., Patil, R., Chavan, A. et al. Mapping gibberellin-sensitive dwarfing locus Rht18 in durum wheat and development of SSR and SNP markers for selection in breeding. Mol Breeding 37, 28 (2017). https://doi.org/10.1007/s11032-017-0641-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-017-0641-9