Abstract
Tocopherol cyclase (VTE1) plays a key role in promoting the production of γ-tocopherol and improving total tocopherol content in photosynthetic organisms. Walnut is an important source of tocopherols in the human diet, and γ-tocopherol is the major tocopherol compound in walnut kernels. In this study, a full-length cDNA of the VTE1 gene was isolated from walnut using RT-PCR and RACE, and designated as JrVTE1. The full-length cDNA of the JrVTE1 gene contained a 1353-bp open-reading frame encoding a 451-amino-acid protein with a calculated molecular weight of 49.5 kDa. The deduced JrVTE1 protein had a considerable homology with other plant VTE1s and belonged to the tocopherol cyclase family. Functional characterization of JrVTE1 by heterologous expression was carried out in E. coli BL21 (DE3) and microshoot lines of the fruit trees jujube (Zizyphus jujuba var. spinosa) and pear (Pyrus communis) cultivar ‘Old Home’. JrVTE1 in E. coli expressed as a 50 kDa protein, as expected. One or two copies of the transferred JrVTE1 gene were detected in the genomes of representative transgenic lines (from the initial transgenic plants) of jujube and pear by gel blots analysis. Over-expression of JrVTE1 in jujube and pear resulted in an accumulation of tocopherol and a shift in tocopherol composition in leaf, root and stem tissues. In the transgenic jujube, the total tocopherol content increased by 29.8 μg/g in the stems of line J3, 43.7 and 22.5 μg/g in the roots and leaves of line J1, respectively, whereas in the transgenic pear it increased by 47.3 μg/g in the leaf of line P3, and 16.7 and 10.4 μg/g in roots and stems of line P9, respectively. In the examined tissues of transgenic plants, the highest accumulation rate was the γ-tocopherol. These results indicate that JrVTE1 is one of the rate-limiting enzymes for tocopherol production and could be used to improve the tocopherol content of tree crops through genetic engineering.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vitamin E, a group of minor but ubiquitous lipid-soluble compounds, consists of four forms of tocopherol (α-, β-, γ-, δ-tocopherol) and tocotrienol (α-, β-, γ-, δ-tocotrienol). Among these, α-tocopherol (αT) was considered the most biologically active form and had captured much attention (Jiang et al. 2004; Amaral et al. 2005). However, several studies have shown that γ-tocopherol (γT) may induce cell death in human prostate cancer cells and induce human breast cancer cells to undergo apoptosis (Jiang et al. 2004; Yu et al. 2008; Hassim et al. 2014). Lee et al. (2009) found that γ- and δ-tocopherols, but not α-tocopherol, activated peroxisome proliferator-activated receptor-γ and antagonized estrogen action in breast cancer.
Vitamin E, synthesized only by photosynthetic organisms, is obtained mainly from vegetable oils, nuts, cereals, green vegetables, fruits and oil seeds, and γ-tocopherol is the major form in seeds (Amaral et al. 2005; Gilliland et al. 2006). Previously, five enzymes (VTE1, VTE2, VTE3, VTE4 and VTE5) functioning in the tocopherol biosynthetic pathway have been elucidated in Arabidopsis (Addlesee et al. 1996; Collakova and DellaPenna 2001; Porfirova et al. 2002; Bergmüller and Dörmann 2003; Sattler et al. 2003; Valentin et al. 2006). Among them, the tocopherol cyclase (TC, VTE1) has been reported to be the key enzyme that catalyzes conversion of 2,3-dimethyl-6-phytyl-1,4-benzoquinone (DMPBQ) to γ-tocopherol and promotes the production of γ-tocopherol and the total vitamin E content (Porfirova et al. 2002; Cheng et al. 2003; Kanwischer et al. 2005; Vidi et al. 2006). Hence, much effort has been expended in overexpressing VTE1 to increase γ-tocopherol production and vitamin E content in plants such as Arabidopsis (Kanwischer et al. 2005), transgenic rapeseed (Kumar et al. 2005), transgenic lettuce (Lee et al. 2007) and transgenic tobacco (Yabuta et al. 2013).
Walnut (Juglans regia), an excellent source of many nutrients (Isabel et al. 2013), has high oil, protein, vitamin and mineral content (Ma et al. 2010) and has been proposed as a promising natural food. The beneficial effects of walnut consumption, based on their vitamin E activity, have been well documented (Sze-Tao and Sathe 2000; Mène-Saffrané and DellaPenna 2010; Isabel et al. 2013). γ-Tocopherol has been found to be the major vitamin in walnuts (Lavedrine et al. 1997; Amaral et al. 2005), but only a few studies have been conducted on identifying the enzymes which limit the synthesis of γ-tocopherol (Kanwischer et al. 2005; Kumar et al. 2005; Lee et al. 2007; Yabuta et al. 2013). In this study, we aimed to isolate the JrVTE1 gene from the developing embryo of walnut cultivar ‘Xiangling’ and to accomplish functional characterization of JrVTE1 by heterologous expression analysis in E. coli BL21 (DE3), and in microshoot lines of the woody plants, jujube (Zizyphus jujuba var. spinosa) and pear (Pyrus communis) cultivar ‘Old Home’. We also expect to improve the tocopherols content of tree crops through genetic engineering.
Materials and methods
Plant materials
The JrVTE1 gene was isolated from a developing walnut embryo at 90 days after flowering (DAF). The nut was taken from a 10-year-old tree of walnut cultivar ‘Xiangling’ grown at the forestry experimental station of Shandong Agricultural University, Taian, Shandong Province, China. Microshoot lines of jujube (Z. jujuba var. spinosa) and pear (P. communis) cultivar ‘Old Home’ were obtained from the Shandong Institute of Pomology, Taian, Shandong Province, China, and used for genetic transformation to characterize the function of JrVTE1.
Isolation of JrVTE1 gene from walnut
Total RNA was extracted from the developing walnut embryo following a modified CTAB method (Xu et al. 2012). The partial VTE1 cDNA sequence was amplified by nested polymerase chain reaction (PCR) with a one-step RT-PCR kit (TaKaRa, Dalian, China) using degenerate oligonucleotide primers NGSPF and NGSPR (Table S1). The PCR product was purified using a TaKaRa MiniBEST Gel Extraction Kit (TaKaRa) and ligated into the pMD18-T vector (TaKaRa, Dalian, China) for sequencing at Sangon Biotech (Shanghai, China). Based on the partial VTE1 cDNA sequence, the specific primers GSR3 and GSR5 (Table S1) were designed to perform 5′ rapid amplification of cDNA ends (RACE), and 3′ RACE reactions using the SMARTTM RACE cDNA Amplification kit (Takara, Clontech, China). The 5′ end and 3′ end cDNA sequences were assembled to obtain the full-length cDNA sequence by inserting into a pMD18-T vector for sequencing. Primers JrVTE1-FLF and JrVTE1-FLR (Table S1) were designed to generate the full-length cDNA of JrVTE1 using 5′-RACE-Ready cDNA as a template. The PCR product was inserted into the pMD18-T vector for sequencing, and the vector was named as pMD18-T-JrVTE1.
The identity of walnut JrVTE1 gene was confirmed via the nucleotide-nucleotide basic local alignment search tool (BLASTn) in the NCBI database. The sequence of JrVTE1 gene was aligned with thirteen of known homologous genes from other plant species by ClustalW (Thompson et al. 1997; Larkin et al. 2007). The neighbor-joining tree (NJ) was constructed based on the p-distance in software MEGA5 (Tamura et al. 2007, 2011). DNAMAN version 4.0 software (Lynnon Biosoft, USA) was used for the deduced amino acid sequences analysis and assembly. pI/Mw Tool software at ExPaSy (http://web.expasy.org/compute_pi/) was used to predict the calculated molecular weight of the deduced JrVTE1.
Quantification of JrVTE1 transcripts in walnut embryos
The quantification of JrVTE1 transcription in walnut embryos at 60, 90 and 120 DAF was determined by real-time quantitative PCR (RT-qPCR). Primers JrVTE1-YF and JrVTE1-YR, ACTINF and ACTINR (Table S1) were designed to amplify a common fragment shared by the cDNA sequence of JrVTE1 and the sequence of the β-actin gene (a house-keeping gene). Total RNA was isolated from each walnut embryo and treated with RNase-free DNase I at 37 °C for 30 min using the DNase I kit (TaKaRa, Dalian, China). For RT-qPCR, samples were analyzed using a Bio-Rad iQ5 real-time PCR detection system with iQ5 optical system software (Carlson et al. 2008). Each reaction volume was 50 μL: 4 µL DNA (~400 ng), 2 µM of each primer, 25µL SYBR Premix Ex Taq™II (TaKaRa, Dalian, China) and sterile water added to the final volume. The PCR conditions were at 95 °C for 3 min, 40 cycles at 94 °C for 20 s, 53 °C for 20 s and 72 °C for 2 min. Each sample was replicated three times and reactions with no template were used as negative control.
Expression of JrVTE1 in E. coli
The open-reading frame (ORF) of JrVTE1 was amplified from pMD18-T-JrVTE1 using the primers JrVTE1-OF and JrVTE1-OR (Table S1) with BamHI and HindIII restriction sites at the 5′ and 3′ ends, respectively. The amplified DNAs were cloned into a pMD18-Simple vector (TaKaRa, Dalian, China) for sequencing, and this vector was named pMD18-Simple-JrVTE1. BamHI and HindIII were used to digest both pMD18-Simple-JrVTE1 and the prokaryotic expression vector pET-28a (preserved in our laboratory). Then, vector pET-28a-JrVTE1 was created by cloning the digested product JrVTE1 into corresponding region of the pET-28a expression vector and confirmed by restriction enzyme digestion and sequencing. Finally, pET-28a-JrVTE1 was transformed into E. coli BL21 (DE3) competent cells (TransGen Biotech, Beijing, China), in which the expression of the JrVTE1 protein was induced by 1 mM isopropyl-beta-d thiogalactopyranoside (IPTG) at different induction times (0, 2, 4, 6, 8 and 10 h). The protein was analyzed on a 12.5 % SDS-PAGE with the empty vector serving as a control.
Genetic transformation of JrVTE1 in jujube and pear
Plant expression vector construction: The ORF of JrVTE1 was amplified from pMD18-T-JrVTE1 using the primers JrVTE1-ZOF and JrVTE1-ZOR (Table S1) with Nde I and Sal I restriction sites at the 5′ and 3′ ends, respectively. The amplified DNAs were cloned into a pMD18-Simple vector (TaKaRa, Dalian, China) and sequenced. The resulting vector was named pMD18-Simple-ZJrVTE1. Nde I and Sal I were used to digest both pMD18-Simple-ZJrVTE1 and the plant expression vector pRI101 (TaKaRa, Dalian, China). Finally, JrVTE1 was cloned into vector pRI101 to create vector pRI101-JrVTE1 and confirmed by sequencing.
Genetic transformation: pRI101-JrVTE1 was transformed into Agrobacterium tumefaciens (strain AGL1) and used to transform microshoot lines of jujube (Z. jujuba var. spinosa) and pear (P. communis) cultivar ‘Old Home’ by the leaf transformation method described by Sun et al. (2011). The calli of jujube and pear produced excisable shoots (2–3 mm) after a period of 7–8 months on selection medium. The marker gene neomycin phosphotransferase-II (NptII) was used to select the transgenic plants. Later roots were induced from regenerated explants on rooting medium. The transgenic jujube and pear were grown in a culture room at 25 °C with a photoperiod of 16-h light and 8-h darkness. They were transferred to pots and acclimated for 2 weeks in the culture room and then moved to the greenhouse.
Identification of transgenic lines
PCR analysis: Transgenic plants were confirmed by PCR using the primers JrVTE1-ZOF and JrVTE1-ZOR (Table S1). The PCR conditions were at 95 °C for 3 min, 30 cycles at 94 °C for 20 s, 54 °C for 20 s and 72 °C for 2 min, and the last step was at 72 °C for 5 min. The wild types were used as the control.
Southern blot analysis: Ten μg of genomic DNA from each transgenic plant was digested at 37 °C overnight using HindIII restriction enzyme (TaKaRa, Japan), subjected to electrophoresis on 0.7 % agarose gel and transferred to positively charged nylon membrane (Amersham, USA). Probe labeling, pre-hybridization, hybridization and subsequent luminescent detection were performed using the DIG High Prime DNA Labeling and Detection Starter Kit II following the protocol (Roche, USA). Twenty µL of denatured DIG-labeled probe was added to the blots and was incubated at 37 °C overnight. Blots were washed and the hybridization signals were detected by chemiluminescent detection following the manufacturer’s suggested protocol for DIG/CDP-Star (Roche, USA). Blots were exposed to X-ray films for 1–3 h at room temperature, and autoradiograms were developed (Dandekar et al. 1998; Kumar et al. 2015).
Determination of tocopherols by HPLC
Standard and sample preparation: Tocopherol (α-, β-, γ- and δ) (Sigma-Aldrich Co., China) was used as internal standard (IS). A stock solution of the IS (10 mg/mL) was prepared in n-hexane, kept at −20 °C and protected from light, and the final concentration of standard α-, β-, γ- and δ-tocopherols was 1 mg/mL.
One gram of finely ground walnut was accurately weighed and introduced into glass test tubes with 100 μL of butylated hydroxytoluene (BHT) solution (10 mg/mL) as antioxidant (Amaral et al. 2005), and the extraction was extracted with 20 mL n-hexane by ultrasound for 15 min. Extracts were evaporated until dried in a vacuum rotoevaporator (Ya Rong, Shanghai) at 37 °C. The residue was dissolved in 20 mL n-hexane, filtered (0.22 μm membrane) and into 1.5 mL autosampler tubes.
One gram of each tissue (leaf, root and stem) from transgenic and non-transgenic jujube and pear was extracted with 10 mL n-hexane by ultrasound for 15 min. One gram of anhydrous sodium was added and then vortexed immediately and vigorously for 1.5 min. After centrifugation for 5 min at 5000 revolutions per minute (rpm), the supernatant (5 mL) was concentrated to 1 mL and filtered (0.22 μm membrane) into 1.5-mL autosampler tubes.
Determination of tocopherols: Tocopherols were separated by normal-phase HPLC (90 % [v/v] n-hexane and 10 % methyl tertbutyl ether) using an injection volume of 20 μL, a flow rate of 1.0 mL/min, and a run time of 20 min. HPLC analysis was performed on a LiChrospher Si-60 column (5 μm) (Merck, Germany) (Pinheiro-Sant’ana et al. 2011), using a UV detector (Shimadzu, 10AVP, Japan). The detection wavelength was set at 285 nm. Peaks of α-, β-, γ- and δ-tocopherol were identified by comparing their retention times with commercially available authentic standards. Tocopherol concentration and composition was calculated from standard curves.
Results
Tocopherol content in the developing walnut embryo
HPLC measurements showed that α-, β-, γ- and δ-tocopherols were all present in the developing walnut embryo (Supplement 2). Total tocopherol content was approximately 32, 96 and 108 µg/g in walnut embryos at 60, 90, and 120 DAF, respectively, and γ-tocopherol was the main form at each time period. The total tocopherol content at 90 DAF was twofold higher than at 60 DAF, and γ-tocopherol accumulated rapidly from 60 to 90 DAF but more slowly during the 90–120 DAF intervals (Supplement 2).
Isolation and phylogenetic analysis of the JrVTE1 gene
The full-length cDNA of the JrVTE1 gene was isolated and designated as JrVTE1 (GenBank accession no. KC751543) which contained a 1353 bp ORF encoding a 451-amino-acid protein with a calculated molecular weight of 49.5 kDa.
Alignment of the deduced amino acid sequences of VTE1 homologs from Glycine max (XP_003522704) and other plant species showed that JrVTE1 shared 72 % identity with Eucalyptus gunnii and Glycine max, 71 % with Hevea brasiliensis and Vitis vinifera, 69 % with Sesamum indicum, and 67 % with Brassica napus (Supplement 3).
The neighbor-joining tree (NJ) showed that the VTE1s formed three clades with Clade 1 consisting of VTE1s from nine genera, including Arabidopsis, Eucalyptus and Solanum. Clade 2, a sister to clade 1, was comprised of VTE1s from species of the legume family such as Glycine max, Medicago truncatula and Cicer arietinum. VTE1 from Triticum aestivum, a monocot, was more distantly related. JrVTE1 was a sister to Vitis vinifera, being nested into Clade 1, indicating a common ancestor with other VTE1s (Fig. 1). These all indicated that JrVTE1 protein had a considerable homology with other plant VTE1s which belong to the tocopherol cyclase family.
JrVTE1 gene expression in the developing walnut embryo
To investigate JrVTE1 gene expression at different periods during development of the walnut embryo, total RNA was isolated from embryos at 60, 90 and 120 DAF. Real-time PCR analysis revealed that JrVTE1 transcripts were present in all tested samples with the greatest amount of transcript detected at 90 DAF (Supplement 6).
JrVTE1 expression in prokaryotic cells
When recombinant expression plasmid pET28a-JrVTE1 was transferred into E. coli and expression of the target protein was induced by addition of IPTG to cells cultured for 0–10 h, a protein of the expected 49.5 kDa molecular weight was expressed and increased in concentration with increasing induction time. This protein was absent in non-induced cells transformed with the same vector. The target protein did not appear in cells transformed with the negative control plasmid pET-28a during any of the culture periods (Supplement 7). Expression of the target protein maximized at ~6 to 10 h (Supplement 7).
Genetic transformation
Approximately 35 and 65 % of the jujube and pear calli produced excisable shoots (2–3 mm) on selection medium. Fifteen transgenic lines of jujube and forty-one transgenic lines of pear were obtained. The plant regeneration from transformed pear and jujube lines are presented in Fig. 2.
Plant regeneration from jujube and pear transformed with JrVTE1. a, e Putative transgenic adventitious calli from leaves of pear and jujube, respectively. b, f Putative transgenic adventitious bud regeneration from leaf calli of pear and jujube, respectively. c, g Transgenic shoot proliferation of pear and jujube, respectively. d, h Transgenic plants of pear and jujube, respectively
Confirmation of transgenic lines
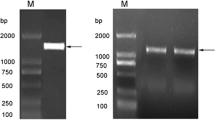
PCR analysis: Genome DNA extracted from each putative transgenic line of jujube and pear was used as the template for PCR analysis. Transgenic lines J1, J3, J4, J6, J8 and J9 of jujube and transgenic lines P3, P7, P9, P10, P12, P15, P16 and P18 of pear were all detected the DNA fragment of 1353 bp; No amplification product was detected from DNA samples of transgenic lines P4, P6 and the wild types JW, PW (Supplement 8).
Southern hybridization analysis: Integration of the transgene-bearing T-DNA (Fig. 3a) into the recipient jujube and pear genomes was further confirmed by Southern hybridization analysis. Gel blots of HindIII-digested genomic DNA of three representative transformants of jujube and pear gave different hybridization bands, and no band was found in the wild type (Fig. 3b). The transgenic lines J1, J3, J4 of jujube and lines P3, P7, P9 of pear all arose from independent transformation events, and the genomic loci into which the T-DNA was inserted were likely different. Two copies of the transferred JrVTE1 gene were detected in the genomes of lines J1, J3, P3, P7, P9 and one copy in line J4 of jujube.
Southern blot analysis of six representative transgenic lines. a Schematic representation of T-DNA region of binary vectors pRI101. RB: right border, CaMV35S: cauliflower mosaic virus 35S promoter, AtAD5′-UTR: the enhancer of JrVTE1 expression, JrVTE1: tocopherol cyclase gene, NOST: nopaline synthase gene terminator, NPTII: neomycin phosphotransferase II, LB: left border. b Southern blot analysis of transgenic jujube and pear lines. lane 1 is the positive control: PCR product of JrVTE1 (1.3 kb) gene, lanes 2, 3, and 4 show the presence of JrVTE1 integration in transgenic jujube lines 4, 3, and 1, lane 5 is wild jujube DNA not showing any band, lanes 6, 7, and 8 show the presence of JrVTE1 integration in transgenic pear lines 3, 7 and 9, lane 9 is wild pear DNA not showing any band
Functional analysis of JrVTE1
HPLC analysis showed that overexpression of JrVTE1 results in accumulation of tocopherol and a shift in tocopherol composition in each tissue of transgenic jujube and pear examined. Total tocopherol and α-tocopherol were found mainly in the leaves of both of jujube and pear, not in roots and stems, and δ-tocopherol was not detected in any tissue (Figs. 4, 5). Overexpression of JrVTE1 using a 35S promoter resulted in an increase of 29.8 μg/g in total tocopherol in the stems of the jujube transformant line J3, and an increase of 43.7 and 22.5 μg/g in the roots and the leaves of line J1, respectively (Fig. 4d). The γ-tocopherol content in the roots of the transformants was 53 % ~2.3-fold higher than wild type (JW) (Fig. 4c). In the pear transformant line P9, total tocopherol content was increased by 16.7 and 10.4 μg/g in roots and stems, respectively, and 47.3 μg/g increased in the leaf of line P3 (Fig. 5d). The γ-tocopherol content in the leaves of the transformants was almost 4.8–16.2-fold higher than wild type (PW) (Fig. 5c).
Analysis of tocopherol content in transgenic jujube roots, stems and leaves. a The α-tocopherol content (μg/g) in roots, stems and leaves of wild-type (JW) and transgenic 35S::JrVTE1 jujube lines (J1, J3, J4, J6, J8 and J9). b The β-tocopherol contents (μg/g) in roots, stems and leaves of wild-type (JW) and transgenic 35S::JrVTE1 jujube lines (J1, J3, J4, J6, J8 and J9). c The γ-tocopherol content (μg/g) in roots, stems and leaves of wild-type (JW) and transgenic 35S::JrVTE1 jujube lines (J1, J3, J4, J6, J8 and J9). d The total tocopherol content (μg/g) in roots, stems and leaves of wild-type (JW) and transgenic 35S::JrVTE1 jujube lines (J1, J3, J4, J6, J8 and J9). The experiments were independently replicated three times
Analysis of tocopherol content in transgenic pear roots, stems and leaves. a The α-tocopherol content (μg/g) in roots, stems and leaves of wild-type (PW) and transgenic 35S::JrVTE1 pear lines (P3, P7, P9, P10, P12, P15, P16 and P18). b The β-tocopherol contents (μg/g) in roots, stems and leaves of wild-type (PW) and transgenic 35S::JrVTE1 pear lines (P3, P7, P9, P10, P12, P15, P16 and P18). c The γ-tocopherol contents (μg/g) in roots, stems and leaves of wild-type (PW) and transgenic 35S::JrVTE1 pear lines (P3, P7, P9, P10, P12, P15, P16 and P18). d The total tocopherol contents (μg/g) in roots, stems and leaves of wild-type (PW) and transgenic 35S::JrVTE1 pear lines (P3, P7, P9, P10, P12, P15, P16 and P18). The experiments were independently replicated three times
Discussion
JrVTE1 gene was isolated from walnut and belonged to the tocopherol cyclase family
The aim of forest tree genomics is to support genetic improvement programs via understanding the association between gene function and morphological traits (Neale and Kremer 2011). The draft genome sequence of the cultivar ‘Chandler’ has already been produced (http://www.ncbi.nlm.nih.gov/bioproject/285351). And many functional genes have been isolated from walnut, such as a gene encoding a 2S albumin seed storage protein precursor (Teuber et al. 1998), two full orthologous cDNAs of chalcone synthase (CHS) which is the key enzyme of flavonoid biosynthesis (Claudot et al. 1999), the orthologous cDNAs of the jrAG and jrAP3 genes which have shown a preferential expression in flowers (Breton et al. 2004), the walnut SDH (shikimate dehydrogenase) gene that is essential for gallic acid (GA) synthesis (Muir et al. 2011), the JrPAL (Phenylalanine ammonia lyase) gene which is the first key enzyme gene of the phenylpropanoid pathway (Xu et al. 2012), and Araji et al. (2014) suggested that the jrPPO1 gene (enzyme polyphenol oxidase) plays a novel and a fundamental role in secondary metabolism and acts as an indirect regulator of cell death in walnut. However, walnut is a healthy dietary component with high micronutrient content and there is significant potential benefit in isolating the relevant genes in order to understand the molecular and biochemical basis of their impact on the micronutrient level of walnut (Gilliland et al. 2006). Hence, the full-length cDNA of the VTE1 gene, which plays a key role in promoting the production of γ-tocopherol and improving total tocopherol content in photosynthetic organisms, was successfully isolated from the developing embryo of the walnut cultivar ‘Xiangling’. Sequence multi-alignment demonstrated that the deduced JrVTE1 protein had a considerable homology to other plant VTE1s and belonged to the tocopherol cyclase family (Supplement 3). Phylogenetic analysis showed that JrVTE1 has a common ancestor with other VTE1s and is a sister to the VTE1 from Vitis vinifera (Fig. 1).
The dynamics of γ-tocopherol accumulation are consistent with JrVTE1 expression in walnut developing embryo
The ‘Xiangling’ walnut cultivar is popular in China. During its embryonic developmental process, fat and soluble sugar content peaks and protein metabolism is most active at 60–100 DAF. Fat content increases from 5.1 % at 60 DAF to 50.11 % at 100 DAF (Li et al. 2012). The total tocopherol content of the developing ‘Xiangling’ embryo at 60, 90 and 120 DAF is shown in Supplement 2, and γ-tocopherol is the main type of tocopherol present. γ-Tocopherol accumulated rapidly from 60 to 90 DAF and then slowly during the remaining 90–120 DAF. Similar trends have been observed in developing rapeseed seeds: γ-Tocopherol accumulated slowly over 12–41 DAF, and the greatest γ-tocopherol accumulation occurred from 41 to 53 DAF. For the rapeseed, both total tocopherol content and α- to γ-tocopherol ratios remained constant from 53 DAF to seed maturity (Goffman et al. 1999). Real-time PCR showed that JrVTE1 gene transcripts could be detected in all walnut samples at 60, 90 and 120 DAF, with the highest transcript occurrence at 90 DAF (Supplement 6). Similarities between γ-tocopherol accumulation and JrVTE1 expression in the walnut developing embryo suggest that γ-tocopherol accumulation is correlated with JrVTE1 expression.
JrVTE1 expressed a 49.5 kDa expected protein in E. coli
To validate the reliability and accuracy of the gene cloned from walnut, plasmid pET-28a-JrVTE1 was created and expressed in E. coli BL21 (DE3) competent cells. After the gene expression in E. coli., a protein of 50 kDa was detected as theoretically expected (49.5 kDa) (Supplement 7). This result was similar to that of the VTE1 from Arabidopsis in which the cyclase activity eluted with a molecular mass of about 50 kDa, corresponding to the calculated 47 kDa molecular mass (Kumar et al. 2005; Kanwischer et al. 2005). SDS-PAGE analysis showed that the content of the target protein increased as the induction time prolonged while the target protein was absent in non-induced cells (Supplement 7). The successful heterologous expression of JrVTE1 gene in E. coli could lay a basis for further study of JrVTE1 protein function.
Overexpression of JrVTE1 results in an accumulation of tocopherol in jujube and pear mainly due to the increase in γ-tocopherol content
VTE1 catalyzes the cyclization of DMPQ resulting in the formation of γ-tocopherol that is subsequently converted to α-tocopherol (Cheng et al. 2003; Kanwischer et al. 2005; Vidi et al. 2006). Jujube and pear are important fruit crops with high commercial values (Nakajima et al. 2013; Liu et al. 2014). In this study, we successfully obtained the transgenic lines of jujube and pear using leaf as explants and employed the protocol of Sun et al. (2011), and detected one or two copies of the transferred JrVTE1 gene in the representative transgenic lines of jujube and pear by gel blot (Fig. 3). We also found that in the transgenic jujube the total tocopherol content increased by 29.8 μg/g in the stems of the transgenic jujube line J3, 43.7 and 22.5 μg/g in the roots and leaves of line J1, respectively (Fig. 4), whereas in the transgenic pear the total tocopherol content increased by 47.3 μg/g in the leaf of line P3, and 16.7 and 10.4 μg/g in roots and stems of line P9, respectively (Fig. 5). And in the examined tissues of transgenic plants, the highest accumulation rate was the γ-tocopherol relative to wild type (Figs. 4, 5). Total tocopherol levels in leaf, root and stem of transgenic plants (jujube and pear) increased significantly due to an increase in γ-tocopherol, indicating that JrVTE1 is one of the rate-limiting enzymes for tocopherol production and γ-tocopherol accumulation in these tissues of transgenic plants. The commercial values of pear and jujube are in the fruits, so tocopherols content in the fruits of transgenic pear and jujube would be determined in our further work.
Previous studies reported that overexpression of the Arabidopsis VTE1 gene in seeds of transgenic rapeseed plants, and in leaves of Arabidopsis, transgenic tobacco and transgenic lettuce, all resulted in an increase in total tocopherol, due mainly to an increase in γ-tocopherol (Kumar et al. 2005; Kanwischer et al. 2005; Lee et al. 2007; Yabuta et al. 2013; Harish et al. 2013). In addition, overexpressing VTE1 can enhance tolerance to the environmental stresses. Liu et al. (2008) reported that overexpressing VTE1 from Arabidopsis in tobacco could enhance tolerance to drought. Transgenic plants overexpressing OsVTE1 from Oryza sativa showed higher tolerance to salt stress and increased the antioxidant capacity of rice seedlings (Ouyang et al. 2011). Therefore, abiotic stress tests of transgenic jujube and pear plants overexpressing JrVTE1 could be used to identify superior lines which have wide adaptability to the environment as well as high content of tocopherols.
References
Addlesee HA, Gibson LCD, Jensen PE, Hunter CN (1996) Cloning, sequencing and functional assignment of the chlorophyll biosynthesis gene, chlP, of Synechocystis sp. PCC 6803. FEBS Lett 389:126–130
Amaral JS, Alves MR, Seabra RM, Oliveira BPP (2005) Vitamin E composition of walnuts (Juglans regia L.): a 3-year comparative study of different cultivars. J Agric Food Chem 53:5467–5472
Araji S, Grammer TA, Gertzen R, Anderson SD, Mikulic-Petkovsek M, Veberic R, Phu ML, Solar A, Leslie CA, Dandekar AM, Escobar MA (2014) Novel roles for the polyphenol oxidase enzyme in secondary metabolism and the regulation of cell death in walnut. Plant Physiol 164:1191–1203
Bergmüller ES, Dörmann Porfirova P (2003) Characterization of an Arabidopsis mutant deficient in γ-tocopherol methyltransferase. Plant Mol Biol 52:1181–1190
Breton C, Cornu D, Chriqui D, Sauvanet A, Capelli P, Germain E, Jay-Allemand C (2004) Somatic embryogenesis, micropropagation and plant regeneration of “early mature” walnut trees (Juglans regia) that flower in vitro. Tree Physiol 24:425–435
Carlson ME, Hsu M, Conboy IM (2008) Imbalance between pSmad3 and Notch induces CDK inhibitors in old muscle stem cells. Nature 454:528–532
Cheng Z, Sattler S, Maeda H, Sakuragi Y, Bryant DA, DellaPenna D (2003) Highly divergent methyltransferases catalyze a conserved reaction in tocopherol and plastoquinone synthesis in cyanobacteria and photosynthetic eukaryotes. Plant Cell 15:2343–2356
Claudot AC, Ernst D, Jra HS, Drouet A (1999) Cloning and characterization of two members of the chalcone synthase gene family from walnut. Plant Physiol Biochem 37:721–730
Collakova E, DellaPenna D (2001) Isolation and functional analysis of homogentisate phytyltransferase from Synechocystis sp. PCC 6803 and Arabidopsis. Plant Physiology 127:1113–1124
Dandekar AM, McGranahan GH, Vail PV, Uratsu SL, Leslie CA, Tebbets JS (1998) High levels of expression of full-length cryIA(c) gene from Bacillus thuringiensis in transgenic somatic walnut embryos. Plant Sci 131:181–193
Gilliland LU, Magallanes-Lundback M, Hemming C, Supplee A, Koornneef M, Leo B, Della Penna D (2006) Genetic basis for natural variation in seed vitamin E levels in Arabidopsis thaliana. Proc Natl Acad Sci USA 49:18834–18841
Goffman FD, Velasco L, Becker HC (1999) Tocopherols accumulation in developing seeds and pods of rapeseed (Brassica napus L.). Lipid/Fett 101:400–403
Harish MC, Dachinamoorthy P, Balamurugan S, Sathishkumar R (2013) Overexpression of homogentisate phytyltransferase (HPT) and tocopherol cyclase (TC) enhances α-tocopherol content in transgenic tobacco. Biol Plant 57:395–400
Hassim F, Zulkafli N, Rahman T, Kornain NK, Nawawi H (2014) Pure tocotrienols reduce endothelial expression of interleukin-6, matrix metalloproteinases in experimentally induced mild and severe atherosclerotic rabbits. Atherosclerosis 2:e122
Isabel TM, Ramón SMJ, Jesús GP, Rosario R, Teresa H, David GG (2013) Comparative study of the nutritional and bioactive compounds content of four walnut (Juglans regia L.) cultivars. J Food Compos Anal 31:232–237
Jiang Q, Wong J, Fyrst H, Saba JD, Ames BN (2004) γ-Tocopherol or combinations of vitamin E forms induce cell death in human prostate cancer cells by interrupting sphingolipid synthesis. Proc Natl Acad Sci USA 51:17825–17830
Kanwischer M, Porfirova S, Bergmüller E, Dörmann P (2005) Alterations in tocopherol cyclase activity in transgenic and mutant plants of Arabidopsis affect tocopherol content, tocopherol composition, and oxidative stress. Plant Physiol 137:713–723
Kumar R, Raclaru M, Schüßeler T, Gruber J, Sadre R, Lühs W, Zarhloul KM, Friedt W, Enders D, Frentzen M, Weier D (2005) Characterisation of plant tocopherol cyclases and their overexpression in transgenic Brassica napus seeds. FEBS Lett 579:1357–1364
Kumar S, AlAbed D, Worden A, Novak S, Wu HX, Ausmus C, Beck M, Robinson H, Minnicks T, Hemingway D, Lee R, Skaggs N, Wang LZ, Marri P, Gupta M (2015) A modular gene targeting system for sequential transgene stacking in plants. J Biotechnol 207:12–20
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Lavedrine F, Ravel A, Poupard A, Alary J (1997) Effect of geographic origin, variety and storage on tocopherol concentrations in walnuts by HPLC. Food Chem 58:135–140
Lee K, Lee SM, Park SR, Jung J, Moon JK, Cheong JJ, Kim M (2007) Overexpression of Arabidopsis homogentisate phytyltransferase or tocopherol cyclase elevates vitamin E content by increasing γ-tocopherol level in lettuce (Lactuca sativa L.). Mol Cells 24:301–306
Lee HJ, Ju J, Paul S, So JY, DeCastro A, Smolarek A, Lee MJ, Yang CS, Newmark HL, Suh N (2009) Mixed tocopherols prevent mammary tumorigenesis by inhibiting estrogen action and activating PPAR-γ. Clin Cancer Res 15:4242–4249
Li YT, Ma SM, Wang YF, Xuan XL, Hou LQ, Sun QR, Yang KQ (2012) The dynamics of fat, protein and sugar metabolism during walnut (Juglans regia L.) fruit development. Afr J Biotechnol 5:1267–1276
Liu XL, Hua XJ, Guo J, Qi DM, Wang LJ, Liu ZP, Jin ZP, Chen SY, Liu GS (2008) Enhanced tolerance to drought stress in transgenic tobacco plants overexpressing VTE1 for increased tocopherol production from Arabidopsis thaliana. Biotechnol Lett 30:1275–1280
Liu J, Liu HB, Ma LY, Wang SQ, Gao J, Li YY, Wu RL, Pang XM (2014) A Chinese jujube (Ziziphus jujuba Mill.) fruit-expressed sequence tag (EST) library: annotation and EST-SSR characterization. Sci Hortic 165:99–105
Ma Y, Njike VY, Millet J, Dutta S, Doughty K, Treu JA, Katz DL (2010) Effects of walnut consumption on endothelial function in type 2 diabetic subjects: a randomized controlled cross-over trial. Diabetes Care 33:227–232
Mène-Saffrané L, DellaPenna D (2010) Biosynthesis, regulation and functions of tocochromanols in plants. Plant Physiol Biochem 48:301–309
Muir RM, Ibáñez AM, Uratsu SL, Ingham ES, Leslie CA, McGranahan GH, Batra N, Goyal S, Joseph J, Jemmis ED, Dandekar AM (2011) Mechanism of gallic acid biosynthesis in bacteria (Escherichia coli) and walnut (Juglans regia). Plant Mol Biol 75:555–565
Nakajima I, Sato Y, Saito T, Moriguchi T, Yamamoto T (2013) Agrobacterium-mediated genetic transformation using cotyledons in Japanese pear (Pyrus pyrifolia). Breed Sci 63:275–283
Neale DB, Kremer A (2011) Forest tree genomics: growing resources and applications. Nat Rev Genet 12:111–122
Ouyang SQ, He SJ, Liu P, Zhang WK, Zhang JS, Chen SY (2011) The role of tocopherol cyclase in salt stress tolerance of rice (Oryza sativa). Life Sci 2:181–188
Pinheiro-Sant’ana HM, Guinazi M, Oliveira Dda S, Della Lucia CM, Reis Bde L, Brandão SC (2011) Method for simultaneous analysis of eight vitamin E isomers in various foods by high performance liquid chromatography and fluorescence detection. J Chromatogr A 1218:8496–8502
Porfirova S, Bergmuller E, Tropf S, Lemke R, Dormann P (2002) Isolation of an Arabidopsis mutant lacking vitamin E and identification of a cyclase essential for all tocopherol biosynthesis. Proc Natl Acad Sci USA 19:12495–12500
Sattler SE, Cahoon EB, Coughlan SJ, DellaPenna D (2003) Characterization of tocopherol cyclases from higher plants and Cyanobacteria. Evolutionary implications for tocopherol synthesis and function. Plant Physiol 132:2184–2195
Sun QR, Zhao Y, Sun HY, Hammond RW, Davis RE, Li X (2011) High-efficiency and stable genetic transformation of pear (Pyrus communis L.) leaf segments and regeneration of transgenic plants. Acta Physiol Plant 33:383–390
Sze-Tao KWC, Sathe SK (2000) Walnuts (Juglans regia L): proximate composition, protein solubility, protein amino acid composition and protein in vitro digestibility. J Sci Food Agric 80:1393–1401
Tamura K, Dudley J, Nei M, Kurmar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Teuber SS, Dandekar AM, Peterson WR, Sellers CL (1998) Cloning and sequencing of a gene encoding a 2S albumin seed storage protein precursor from English walnut (Juglans regia), a major food allergen. J Allergy Clin Immunol 101:807–814
Thompson DJ, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Valentin HE, Lincoln K, Moshiri F, Jensen PK, Qi Q, Venkatesh TV, Karunanandaa B, Baszis SR, Norris SR, Savidge B (2006) The Arabidopsis vitamin E pathway gene5-1 mutant reveals a critical role for phytol kinase in seed tocopherol biosynthesis. Plant Cell 18:212–224
Vidi PA, Kanwischer M, Baginsky S, Austin JR, Csucs G, Dörmann P, Kessler F, Bréhélin C (2006) Tocopherol cyclase (VTE1) localization and vitamin e accumulation in chloroplast plastoglobuli lipoprotein particles. J Biol Chem 16:11225–11234
Xu F, Deng G, Cheng S, Zhang W, Huang X, Li L, Cheng H, Rong X, Li J (2012) Molecular cloning, characterization and expression of the phenylalanine ammonialyase gene from Juglans regia. Molecules 17:7810–7823
Yabuta Y, Tanaka H, Yoshimura S, Suzuki A, Tamoi M, Maruta T, Shigeoka S (2013) Improvement of vitamin E quality and quantity in tobacco and lettuce by chloroplast genetic engineering. Transgenic Res 22:391–402
Yu WP, Park SK, Jia L, Tiwary R, Scott WW, Li J, Wang P, Simmons-Menchaca M, Sanders BG, Kline K (2008) RRR-γ-tocopherol induces human breast cancer cells to undergo apoptosis via death receptor 5 (DR5)-mediated apoptotic signaling. Cancer Lett 259:165–176
Acknowledgments
This study was supported by a Grant from the National ‘Twelfth Five-Year’ Plan for Science & Technology Support (No. 2013BAD14B01), the National Natural Science Foundation of China (No. 31470680), and the Improved Variety Program of Shandong Province of China (No. 2014LZ14-07).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wang, C., Li, C., Leslie, C.A. et al. Molecular cloning and heterologous expression analysis of JrVTE1 gene from walnut (Juglans regia). Mol Breeding 35, 222 (2015). https://doi.org/10.1007/s11032-015-0414-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0414-2