Abstract
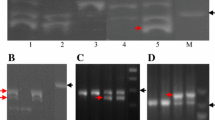
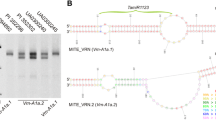
A dominant allele of the vernalization gene Vrn-2 is the wild type conferring winter growth habit, whereas a recessive vrn-2 allele confers spring growth habit. The recessive vrn-2 allele is mutated due to the deletion of the complete gene (a null form) or alternation of a key amino acid in the VRN-2 protein (a nonfunctional form) in diploid wheat or tetraploid wheat. VRN-2 is also denoted ZCCT due to the presence of a zinc finger and a CCT domain in its protein. There are two paralogous ZCCT genes at the VRN-2 locus in diploid Triticum monococcum and three paralogous ZCCT genes on each of the A and B genomes in tetraploid wheat, but little is known about the allelic variation in VRN-2 in hexaploid wheat. In the study reported here, we performed a one-shot PCR to simultaneously amplify the promoter regions of the three ZCCT-1 genes from hexaploid wheat, including the 302-bp fragment from ZCCT-A1, the 294-bp fragment from ZCCT-B1, and the 320-bp fragment from ZCCT-D1. Each amplicon could be differentiated by electrophoresis in an acrylamide/bisacrylamide gel. This PCR marker for different lengths of the three ZCCT-1 genes was used to search for null alleles in hexaploid wheat. A null allele was found in each of ZCCT-A1, ZCCT-B1, and ZCCT-D1 among 74 cultivars and genetic stocks of U.S. hexaploid wheat. Among 54 Chinese wheat cultivars, breeding lines, and landraces, we identified three accessions carrying a single null allele at ZCCT-A1, three accessions carrying a null allele at ZCCT-B1, and one accession carrying a double null allele at both ZCCT-A1 and ZCCT-D1. The potential application of these natural ZCCT-1 mutant materials in wheat breeding programs and studies on the genetics of wheat is discussed.


Similar content being viewed by others
References
Akhunov ED, Akhunova AR, Dvorak J (2005) BAC libraries of Triticum urartu, Aegilops speltoides and Ae. tauschii, the diploid ancestors of polyploid wheat. Theor Appl Genet 111:1617–1622
Bai G, Guo P, Kolb FL (2003) Genetic relationships among head blight resistant cultivars of wheat assessed on the basis of molecular markers. Crop Sci 43:498–507
Baloch DM, Karow RS, Marx E, Kling JG, Witt MD (2003) Vernalization studies with Pacific Northwest Wheat. Agron J 95:1201–1208
Berry GJ, Salisbury PA, Halloran GM (1980) Expression of vernalization genes in near-lsogenic wheat lines: duration of vernalization period. Ann Bot 46:235–241
Cenci A, Chantret N, Kong X, Gu Y, Anderson OD, Fahima T, Distelfeld A, Dubcovsky J (2003) Construction and characterization of a half million clone BAC library of durum wheat (Triticum turgidum ssp. durum). Theor Appl Genet 107:931–939
Crofts HJ (1989) On defining a winter wheat. Euphytica 44:225–234
Distelfeld A, Tranquilli G, Li C, Yan L, Dubcovsky J (2009) Genetic and molecular characterization of the VRN2 loci in tetraploid wheat. Plant Physiol 149:245–257
Dubcovsky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316:1862–1866
Dubcovsky J, Galvez AF, Dvorak J (1994) Comparison of the genetic organization of the early salt stress response gene system in salt-tolerant Lophopyrum elongatum and salt-sensitive wheat. Theor Appl Genet 87:957–964
Dubcovsky J, Lijavetzky D, Appendino L, Tranquilli G (1998) Comparative RFLP mapping of Triticum monococcum genes controlling vernalization requirement. Theor Appl Genet 97:968–975
Dubcovsky J, Chen C, Yan L (2005) Molecular characterization of the allelic variation at the VRN-H2 vernalization locus in barley. Mol Breed 15:395–407
Dubcovsky J, Loukoianov A, Fu D, Valarik M, Sanchez A, Yan L (2006) Effect of photoperiod on the regulation of wheat vernalization genes VRN1 and VRN2. Plant Mol Biol 60:469–480
Fayt VI, Symonenko LK, Mokanu NV, Popova NV (2007) Chromosomal location of genes for vernalization requirement duration (Vrd) in winter bread wheat. Russ J Genet 43:143–148
Fu D, Szucs P, Yan L, Helguera M, Skinner J, Hayes P, Dubcovsky J (2005) Large deletions in the first intron of the VRN-1 vernalization gene are associated with spring growth habit in barley and polyploid wheat. Mol Genet Genomics 273:54–65
Li C, Dubcovsky J (2008) Wheat FT protein regulates VRN1 transcription through interactions with FDL2. Plant J 55:543–554
Loukoianov A, Yan L, Blechl A, Sanchez A, Dubcovsky J (2005) Regulation of VRN-1 vernalization genes in normal and transgenic polyploid wheat. Plant Physiol 138:2364–2373
Shimada S, Ogawa T, Kitagawa S, Suzuki T, Ikari C, Shitsukawa N, Abe T, Kawahigashi H, Kikuchi R, Handa H, Murai K (2009) A genetic network of flowering-time genes in wheat leaves, in which an APETALA1/FRUITFULL-like gene, VRN1, is upstream of FLOWERING LOCUS T. Plant J 58:668–681
Shitsukawa N, Ikari C, Shimada S, Kitagawa S, Sakamoto K, Saito H, Ryuto H, Fukunishi N, Abe T, Takumi S, Nasuda S, Murai K (2007) The einkorn wheat (Triticum monococcum) mutant, maintained vegetative phase, is caused by a deletion in the VRN1 gene. Genes Genet Syst 82:167–170
Takahashi R, Yasuda S (1971) Genetics of earliness and growth habit in barley. In: Nilan RA (ed) Proc 2nd Int Barley Genet Symp. Washington State University Press, Pullman, pp 388–408
Tranquilli G, Dubcovsky J (2000) Epistatic interactions between vernalization genes Vrn-A m 1 and Vrn-A m 2 in diploid wheat. J Hered 91:304–306
Trevaskis B, Hemming MN, Peacock WJ, Dennis ES (2006) HvVRN2 responds to daylength, whereas HvVRN1 is regulated by vernalization and developmental status. Plant Physiol 140:1397–1405
Trevaskis B, Hemming MN, Dennis ES, Peacock WJ (2007) The molecular basis of vernalization-induced flowering in cereals. Trends Plant Sci 12:352–357
Wang SY, Ward RW, Ritchie JT, Fischer RA, Schulthess U (1995) Vernalization in wheat II. Genetic variability for the interchangeability of plant age and vernalization duration. Field Crops Res 44:67–72
Wenkel S, Turck F, Singer K, Gissot L, Le Gourrierec J, Samach A, Coupland G (2006) CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering of Arabidopsis. Plant Cell 18:2971–2984
Xue WY, Xing YZ, Weng XY, Zhao Y, Tang WJ, Wang L, Zhou HJ, Yu SB, Xu CG, Li XH, Zhang QF (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767
Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of wheat vernalization gene VRN1. Proc Natl Acad Sci U S A 100:6263–6268
Yan L, Helguera M, Kato K, Fukuyama S, Sherman J, Dubcovsky J (2004a) Allelic variation at the VRN-1 promoter region in polyploid wheat. Theor Appl Genet 109:1677–1686
Yan L, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, SanMiguel P, Bennetzen JL, Echenique V, Dubcovsky J (2004b) The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science 303:1640–1644
Yan L, Fu D, Li C, Blechl A, Tranquilli G, Bonafede M, Sanchez A, Valarik M, Dubcovsky J (2006) The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proc Natl Acad Sci U S A 103:19581–19586
Zhang XK, Xia XC, Xiao YG, Dubcovsky J, He ZH (2008) Allelic variation at the vernalization genes Vrn-A1, Vrn-B1, Vrn-D1 and Vrn-B3 in Chinese common wheat cultivars and their association with growth habit. Crop Sci 48:458–470
Acknowledgments
This study was supported by the Oklahoma Center of Advanced Science and Technology (OCAST) and partially funded by the Oklahoma Agricultural Experiment Station and the Oklahoma Wheat Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Xinkai Zhu and ChorTee Tan contributed equally to the work.
Rights and permissions
About this article
Cite this article
Zhu, X., Tan, C., Cao, S. et al. Molecular differentiation of null alleles at ZCCT-1 genes on the A, B, and D genomes of hexaploid wheat. Mol Breeding 27, 501–510 (2011). https://doi.org/10.1007/s11032-010-9447-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-010-9447-8