Abstract
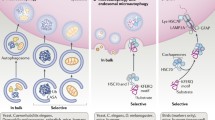
Chaperone-mediated autophagy (CMA), one of the degradation pathways of proteins, is highly selective to substrates that have KFERQ-like motif. In this process, the substrate proteins are first recognized by the chaperone protein, heat shock cognate protein 70 (Hsc70), then delivered to lysosomal membrane surface where the single-span lysosomal receptor, lysosome-associated membrane protein type 2A (LAMP2A) can bind to the substrate proteins to form a 700 kDa protein complex that allows them to translocate into the lysosome lumen to be degraded by the hydrolytic enzymes. This degradation pathway mediated by CMA plays an important role in regulating glucose and lipid metabolism, transcription, DNA reparation, cell cycle, cellular response to stress and consequently, regulating many aging-associated human diseases, such as neurodegeneration, cancer and metabolic disorders. In this review, we provide an overview of current research on the functional roles of CMA primarily from a perspective of understanding and treating human diseases and also discuss its potential applications for diseases.



Similar content being viewed by others
References
Grant BD, Donaldson JG (2009) Pathways and mechanisms of endocytic recycling. Nat Rev Mol Cell Biol 10(9):597–608. https://doi.org/10.1038/nrm2755
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24(1):24–41. https://doi.org/10.1038/cr.2013.168
Santambrogio L, Cuervo AM (2011) Chasing the elusive mammalian microautophagy. Autophagy 7(6):652–654. https://doi.org/10.4161/auto.7.6.15287
Sahu R, Kaushik S, Clement CC, Cannizzo ES, Scharf B, Follenzi A, Potolicchio I, Nieves E, Cuervo AM, Santambrogio L (2011) Microautophagy of cytosolic proteins by late endosomes. Dev Cell 20(1):131–139. https://doi.org/10.1016/j.devcel.2010.12.003
Kaushik S, Cuervo AM (2018) The coming of age of chaperone-mediated autophagy. Nat Rev Mol Cell Biol 19(6):365–381. https://doi.org/10.1038/s41580-018-0001-6
Rodríguez-Muela N, Koga H, García-Ledo L, de la Villa P, de la Rosa EJ, Cuervo AM, Boya P (2013) Balance between autophagic pathways preserves retinal homeostasis. Aging Cell 12(3):478–488. https://doi.org/10.1111/acel.12072
Massey AC, Kaushik S, Sovak G, Kiffin R, Cuervo AM (2006) Consequences of the selective blockage of chaperone-mediated autophagy. Proc Natl Acad Sci U S A 103(15):5805–5810. https://doi.org/10.1073/pnas.0507436103
Kaushik S, Massey AC, Mizushima N, Cuervo AM (2008) Constitutive activation of chaperone-mediated autophagy in cells with impaired macroautophagy. Mol Biol Cell 19(5):2179–2192. https://doi.org/10.1091/mbc.e07-11-1155
Chava S, Lee C, Aydin Y, Chandra PK, Dash A, Chedid M, Thung SN, Moroz K, Wu T, Nayak NC, Dash S (2017) Chaperone-mediated autophagy compensates for impaired macroautophagy in the cirrhotic liver to promote hepatocellular carcinoma. Oncotarget 8(25):40019–40036. https://doi.org/10.18632/oncotarget.16685
Meléndez A, Tallóczy Z, Seaman M, Eskelinen EL, Hall DH, Levine B (2003) Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 301(5638):1387–1391. https://doi.org/10.1126/science.1087782
Cadwell K, Patel KK, Maloney NS, Liu TC, Ng AC, Storer CE, Head RD, Xavier R, Stappenbeck TS, Virgin HW (2010) Virus-plus-susceptibility gene interaction determines Crohn’s disease gene Atg16L1 phenotypes in intestine. Cell 141(7):1135–1145. https://doi.org/10.1016/j.cell.2010.05.009
Bretin A, Carrière J, Dalmasso G, Bergougnoux A, B’Chir W, Maurin AC, Müller S, Seibold F, Barnich N, Bruhat A, Darfeuille-Michaud A, Nguyen HT (2016) Activation of the EIF2AK4-EIF2A/eIF2α-ATF4 pathway triggers autophagy response to Crohn disease-associated adherent-invasive Escherichia coli infection. Autophagy 12(5):770–783. https://doi.org/10.1080/15548627.2016.1156823
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A, Rosen J, Eskelinen EL, Mizushima N, Ohsumi Y, Cattoretti G, Levine B (2003) Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Invest 112(12):1809–1820. https://doi.org/10.1172/jci20039
Zatloukal K, Stumptner C, Fuchsbichler A, Heid H, Schnoelzer M, Kenner L, Kleinert R, Prinz M, Aguzzi A, Denk H (2002) p62 Is a common component of cytoplasmic inclusions in protein aggregation diseases. Am J Pathol 160(1):255–263. https://doi.org/10.1016/s0002-9440(10)64369-6
Hars ES, Qi H, Ryazanov AG, Jin S, Cai L, Hu C, Liu LF (2007) Autophagy regulates ageing in C. elegans. Autophagy 3(2):93–95. https://doi.org/10.4161/auto.3636
Nakai A, Yamaguchi O, Takeda T, Higuchi Y, Hikoso S, Taniike M, Omiya S, Mizote I, Matsumura Y, Asahi M, Nishida K, Hori M, Mizushima N, Otsu K (2007) The role of autophagy in cardiomyocytes in the basal state and in response to hemodynamic stress. Nat Med 13(5):619–624. https://doi.org/10.1038/nm1574
Schneider JL, Suh Y, Cuervo AM (2014) Deficient chaperone-mediated autophagy in liver leads to metabolic dysregulation. Cell Metab 20(3):417–432. https://doi.org/10.1016/j.cmet.2014.06.009
Hubbi ME, Gilkes DM, Hu H, Kshitiz AI, Semenza GL (2014) Cyclin-dependent kinases regulate lysosomal degradation of hypoxia-inducible factor 1α to promote cell-cycle progression. Proc Natl Acad Sci U S A 111(32):E3325-3334. https://doi.org/10.1073/pnas.1412840111
Li L, Fang R, Liu B, Shi H, Wang Y, Zhang W, Zhang X, Ye L (2016) Deacetylation of tumor-suppressor MST1 in Hippo pathway induces its degradation through HBXIP-elevated HDAC6 in promotion of breast cancer growth. Oncogene 35(31):4048–4057. https://doi.org/10.1038/onc.2015.476
Kon M, Kiffin R, Koga H, Chapochnick J, Macian F, Varticovski L, Cuervo AM (2011) Chaperone-mediated autophagy is required for tumor growth. Sci Transl Med 3(109):109ra117. https://doi.org/10.1126/scitranslmed.3003182
Alfaro IE, Albornoz A, Molina A, Moreno J, Cordero K, Criollo A, Budini M (2018) Chaperone mediated autophagy in the crosstalk of neurodegenerative diseases and metabolic disorders. Front Endocrinol (Lausanne) 9:778. https://doi.org/10.3389/fendo.2018.00778
Xilouri M, Brekk OR, Polissidis A, Chrysanthou-Piterou M, Kloukina I, Stefanis L (2016) Impairment of chaperone-mediated autophagy induces dopaminergic neurodegeneration in rats. Autophagy 12(11):2230–2247. https://doi.org/10.1080/15548627.2016.1214777
Lescat L, Véron V, Mourot B, Péron S, Chenais N, Dias K, Riera N, Beaumatin F, Pinel K, Priault M, Panserat S, Salin B, Guiguen Y, Bobe J, Herpin A, Seiliez I (2020) Chaperone-mediated autophagy in the light of evolution: insight from fish. Mol Biol Evol. https://doi.org/10.1093/molbev/msaa127
Dice JF (2007) Chaperone-mediated autophagy. Autophagy 3(4):295–299. https://doi.org/10.4161/auto.4144
Cuervo AM, Knecht E, Terlecky SR, Dice JF (1995) Activation of a selective pathway of lysosomal proteolysis in rat liver by prolonged starvation. Am J Physiol 269(5 Pt 1):C1200-1208. https://doi.org/10.1152/ajpcell.1995.269.5.C1200
Finn PF, Dice JF (2005) Ketone bodies stimulate chaperone-mediated autophagy. J Biol Chem 280(27):25864–25870. https://doi.org/10.1074/jbc.M502456200
Arias E, Cuervo AM (2011) Chaperone-mediated autophagy in protein quality control. Curr Opin Cell Biol 23(2):184–189. https://doi.org/10.1016/j.ceb.2010.10.009
Anguiano J, Garner TP, Mahalingam M, Das BC, Gavathiotis E, Cuervo AM (2013) Chemical modulation of chaperone-mediated autophagy by retinoic acid derivatives. Nat Chem Biol 9(6):374–382. https://doi.org/10.1038/nchembio.1230
Kaushik S, Cuervo AM (2015) Degradation of lipid droplet-associated proteins by chaperone-mediated autophagy facilitates lipolysis. Nat Cell Biol 17(6):759–770. https://doi.org/10.1038/ncb3166
Park C, Suh Y, Cuervo AM (2015) Regulated degradation of Chk1 by chaperone-mediated autophagy in response to DNA damage. Nat Commun 6:6823. https://doi.org/10.1038/ncomms7823
Yang Q, She H, Gearing M, Colla E, Lee M, Shacka JJ, Mao Z (2009) Regulation of neuronal survival factor MEF2D by chaperone-mediated autophagy. Science 323(5910):124–127. https://doi.org/10.1126/science.1166088
Cuervo AM, Hu W, Lim B, Dice JF (1998) IkappaB is a substrate for a selective pathway of lysosomal proteolysis. Mol Biol Cell 9(8):1995–2010. https://doi.org/10.1091/mbc.9.8.1995
Vakifahmetoglu-Norberg H, Kim M, Xia HG, Iwanicki MP, Ofengeim D, Coloff JL, Pan L, Ince TA, Kroemer G, Brugge JS, Yuan J (2013) Chaperone-mediated autophagy degrades mutant p53. Genes Dev 27(15):1718–1730. https://doi.org/10.1101/gad.220897.113
Ali AB, Nin DS, Tam J, Khan M (2011) Role of chaperone mediated autophagy (CMA) in the degradation of misfolded N-CoR protein in non-small cell lung cancer (NSCLC) cells. PLoS One 6(9):e25268. https://doi.org/10.1371/journal.pone.0025268
Welsch T, Younsi A, Disanza A, Rodriguez JA, Cuervo AM, Scita G, Schmidt J (2010) Eps8 is recruited to lysosomes and subjected to chaperone-mediated autophagy in cancer cells. Exp Cell Res 316(12):1914–1924. https://doi.org/10.1016/j.yexcr.2010.02.020
Zhang J, Huang J, Gu Y, Xue M, Qian F, Wang B, Yang W, Yu H, Wang Q, Guo X, Ding X, Wang J, Jin M, Zhang Y (2020) Inflammation-induced inhibition of chaperone-mediated autophagy maintains the immunosuppressive function of murine mesenchymal stromal cells. Cell Mol Immunol. https://doi.org/10.1038/s41423-019-0345-7
Dice JF (1990) Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem Sci 15(8):305–309. https://doi.org/10.1016/0968-0004(90)90019-8
Kirchner P, Bourdenx M, Madrigal-Matute J, Tiano S, Diaz A, Bartholdy BA, Will B, Cuervo AM (2019) Proteome-wide analysis of chaperone-mediated autophagy targeting motifs. PLoS Biol 17(5):e3000301. https://doi.org/10.1371/journal.pbio.3000301
Bandyopadhyay U, Cuervo AM (2008) Entering the lysosome through a transient gate by chaperone-mediated autophagy. Autophagy 4(8):1101–1103. https://doi.org/10.4161/auto.7150
Bandyopadhyay U, Kaushik S, Varticovski L, Cuervo AM (2008) The chaperone-mediated autophagy receptor organizes in dynamic protein complexes at the lysosomal membrane. Mol Cell Biol 28(18):5747–5763. https://doi.org/10.1128/mcb.02070-07
Agarraberes FA, Dice JF (2001) A molecular chaperone complex at the lysosomal membrane is required for protein translocation. J Cell Sci 114(Pt 13):2491–2499
Shin Y, Klucken J, Patterson C, Hyman BT, McLean PJ (2005) The co-chaperone carboxyl terminus of Hsp70-interacting protein (CHIP) mediates alpha-synuclein degradation decisions between proteasomal and lysosomal pathways. J Biol Chem 280(25):23727–23734. https://doi.org/10.1074/jbc.M503326200
Bandyopadhyay U, Sridhar S, Kaushik S, Kiffin R, Cuervo AM (2010) Identification of regulators of chaperone-mediated autophagy. Mol Cell 39(4):535–547. https://doi.org/10.1016/j.molcel.2010.08.004
Dice JF, Chiang HL, Spencer EP, Backer JM (1986) Regulation of catabolism of microinjected ribonuclease A. Identification of residues 7–11 as the essential pentapeptide. J Biol Chem 261(15):6853–6859
Kaushik S, Cuervo AM (2012) Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol 22(8):407–417. https://doi.org/10.1016/j.tcb.2012.05.006
Zhang Y, Xu YY, Yao CB, Li JT, Zhao XN, Yang HB, Zhang M, Yin M, Chen J, Lei QY (2017) Acetylation targets HSD17B4 for degradation via the CMA pathway in response to estrone. Autophagy 13(3):538–553. https://doi.org/10.1080/15548627.2016.1268302
Lv L, Li D, Zhao D, Lin R, Chu Y, Zhang H, Zha Z, Liu Y, Li Z, Xu Y, Wang G, Huang Y, Xiong Y, Guan KL, Lei QY (2011) Acetylation targets the M2 isoform of pyruvate kinase for degradation through chaperone-mediated autophagy and promotes tumor growth. Mol Cell 42(6):719–730. https://doi.org/10.1016/j.molcel.2011.04.025
Thompson LM, Aiken CT, Kaltenbach LS, Agrawal N, Illes K, Khoshnan A, Martinez-Vincente M, Arrasate M, O’Rourke JG, Khashwji H, Lukacsovich T, Zhu YZ, Lau AL, Massey A, Hayden MR, Zeitlin SO, Finkbeiner S, Green KN, LaFerla FM, Bates G, Huang L, Patterson PH, Lo DC, Cuervo AM, Marsh JL, Steffan JS (2009) IKK phosphorylates Huntingtin and targets it for degradation by the proteasome and lysosome. J Cell Biol 187(7):1083–1099. https://doi.org/10.1083/jcb.200909067
Bonhoure A, Vallentin A, Martin M, Senff-Ribeiro A, Amson R, Telerman A, Vidal M (2017) Acetylation of translationally controlled tumor protein promotes its degradation through chaperone-mediated autophagy. Eur J Cell Biol 96(2):83–98. https://doi.org/10.1016/j.ejcb.2016.12.002
Ferreira JV, Soares AR, Ramalho JS, Pereira P, Girao H (2015) K63 linked ubiquitin chain formation is a signal for HIF1A degradation by chaperone-mediated autophagy. Sci Rep 5:10210. https://doi.org/10.1038/srep10210
Cuervo AM, Dice JF (2000) Regulation of lamp2a levels in the lysosomal membrane. Traffic 1(7):570–583. https://doi.org/10.1034/j.1600-0854.2000.010707.x
Cuervo AM, Dice JF (1996) A receptor for the selective uptake and degradation of proteins by lysosomes. Science 273(5274):501–503. https://doi.org/10.1126/science.273.5274.501
Kaushik S, Massey AC, Cuervo AM (2006) Lysosome membrane lipid microdomains: novel regulators of chaperone-mediated autophagy. Embo j 25(17):3921–3933. https://doi.org/10.1038/sj.emboj.7601283
Kiffin R, Christian C, Knecht E, Cuervo AM (2004) Activation of chaperone-mediated autophagy during oxidative stress. Mol Biol Cell 15(11):4829–4840. https://doi.org/10.1091/mbc.e04-06-0477
Ferreira JV, Fôfo H, Bejarano E, Bento CF, Ramalho JS, Girão H, Pereira P (2013) STUB1/CHIP is required for HIF1A degradation by chaperone-mediated autophagy. Autophagy 9(9):1349–1366. https://doi.org/10.4161/auto.25190
Valdor R, Mocholi E, Botbol Y, Guerrero-Ros I, Chandra D, Koga H, Gravekamp C, Cuervo AM, Macian F (2014) Chaperone-mediated autophagy regulates T cell responses through targeted degradation of negative regulators of T cell activation. Nat Immunol 15(11):1046–1054. https://doi.org/10.1038/ni.3003
Pajares M, Rojo AI, Arias E, Díaz-Carretero A, Cuervo AM, Cuadrado A (2018) Transcription factor NFE2L2/NRF2 modulates chaperone-mediated autophagy through the regulation of LAMP2A. Autophagy 14(8):1310–1322. https://doi.org/10.1080/15548627.2018.1474992
Arias E, Koga H, Diaz A, Mocholi E, Patel B, Cuervo AM (2015) Lysosomal mTORC2/PHLPP1/Akt Regulate chaperone-mediated autophagy. Mol Cell 59(2):270–284. https://doi.org/10.1016/j.molcel.2015.05.030
Ormeño F, Hormazabal J, Moreno J, Riquelme F, Rios J, Criollo A, Albornoz A, Alfaro IE, Budini M (2020) Chaperone mediated autophagy degrades TDP-43 protein and is affected by TDP-43 aggregation. Front Mol Neurosci 13:19. https://doi.org/10.3389/fnmol.2020.00019
Li W, Zhu J, Dou J, She H, Tao K, Xu H, Yang Q, Mao Z (2017) Phosphorylation of LAMP2A by p38 MAPK couples ER stress to chaperone-mediated autophagy. Nat Commun 8(1):1763. https://doi.org/10.1038/s41467-017-01609-x
Obayashi H, Nagano Y, Takahashi T, Seki T, Tanaka S, Sakai N, Matsumoto M, Maruyama H (2020) Histone deacetylase 10 knockout activates chaperone-mediated autophagy and accelerates the decomposition of its substrate. Biochem Biophys Res Commun 523(1):246–252. https://doi.org/10.1016/j.bbrc.2019.12.048
Sato M, Ueda E, Konno A, Hirai H, Kurauchi Y, Hisatsune A, Katsuki H, Seki T (2020) Glucocorticoids negatively regulates chaperone mediated autophagy and microautophagy. Biochem Biophys Res Commun 528(1):199–205. https://doi.org/10.1016/j.bbrc.2020.04.132
Arias E, Cuervo AM (2020) Pros and cons of chaperone-mediated autophagy in cancer biology. Trends Endocrinol Metab 31(1):53–66. https://doi.org/10.1016/j.tem.2019.09.007
Cuervo AM, Dice JF (2000) Age-related decline in chaperone-mediated autophagy. J Biol Chem 275(40):31505–31513. https://doi.org/10.1074/jbc.M002102200
Valdor R, García-Bernal D, Riquelme D, Martinez CM, Moraleda JM, Cuervo AM, Macian F, Martinez S (2019) Glioblastoma ablates pericytes antitumor immune function through aberrant up-regulation of chaperone-mediated autophagy. Proc Natl Acad Sci U S A 116(41):20655–20665. https://doi.org/10.1073/pnas.1903542116
Lu W, Zhang Y, McDonald DO, Jing H, Carroll B, Robertson N, Zhang Q, Griffin H, Sanderson S, Lakey JH, Morgan NV, Reynard LN, Zheng L, Murdock HM, Turvey SE, Hackett SJ, Prestidge T, Hall JM, Cant AJ, Matthews HF, Koref MF, Simon AK, Korolchuk VI, Lenardo MJ, Hambleton S, Su HC (2014) Dual proteolytic pathways govern glycolysis and immune competence. Cell 159(7):1578–1590. https://doi.org/10.1016/j.cell.2014.12.001
Orenstein SJ, Kuo SH, Tasset I, Arias E, Koga H, Fernandez-Carasa I, Cortes E, Honig LS, Dauer W, Consiglio A, Raya A, Sulzer D, Cuervo AM (2013) Interplay of LRRK2 with chaperone-mediated autophagy. Nat Neurosci 16(4):394–406. https://doi.org/10.1038/nn.3350
Kabuta T, Furuta A, Aoki S, Furuta K, Wada K (2008) Aberrant interaction between Parkinson disease-associated mutant UCH-L1 and the lysosomal receptor for chaperone-mediated autophagy. J Biol Chem 283(35):23731–23738. https://doi.org/10.1074/jbc.M801918200
Cuervo AM, Stefanis L, Fredenburg R, Lansbury PT, Sulzer D (2004) Impaired degradation of mutant alpha-synuclein by chaperone-mediated autophagy. Science 305(5688):1292–1295. https://doi.org/10.1126/science.1101738
Wang B, Cai Z, Tao K, Zeng W, Lu F, Yang R, Feng D, Gao G, Yang Q (2016) Essential control of mitochondrial morphology and function by chaperone-mediated autophagy through degradation of PARK7. Autophagy 12(8):1215–1228. https://doi.org/10.1080/15548627.2016.1179401
Liu H, Wang P, Song W, Sun X (2009) Degradation of regulator of calcineurin 1 (RCAN1) is mediated by both chaperone-mediated autophagy and ubiquitin proteasome pathways. Faseb j 23(10):3383–3392. https://doi.org/10.1096/fj.09-134296
Wang Y, Martinez-Vicente M, Krüger U, Kaushik S, Wong E, Mandelkow EM, Cuervo AM, Mandelkow E (2010) Synergy and antagonism of macroautophagy and chaperone-mediated autophagy in a cell model of pathological tau aggregation. Autophagy 6(1):182–183. https://doi.org/10.4161/auto.6.1.10815
Huang CC, Bose JK, Majumder P, Lee KH, Huang JT, Huang JK, Shen CK (2014) Metabolism and mis-metabolism of the neuropathological signature protein TDP-43. J Cell Sci 127(Pt 14):3024–3038. https://doi.org/10.1242/jcs.136150
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H, Mann D, Tsuchiya K, Yoshida M, Hashizume Y, Oda T (2006) TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun 351(3):602–611. https://doi.org/10.1016/j.bbrc.2006.10.093
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT, Bruce J, Schuck T, Grossman M, Clark CM, McCluskey LF, Miller BL, Masliah E, Mackenzie IR, Feldman H, Feiden W, Kretzschmar HA, Trojanowski JQ, Lee VM (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314(5796):130–133. https://doi.org/10.1126/science.1134108
Arosio A, Cristofani R, Pansarasa O, Crippa V, Riva C, Sirtori R, Rodriguez-Menendez V, Riva N, Gerardi F, Lunetta C, Cereda C, Poletti A, Ferrarese C, Tremolizzo L, Sala G (2020) HSC70 expression is reduced in lymphomonocytes of sporadic ALS patients and contributes to TDP-43 accumulation. Amyotroph Lateral Scler Frontotemporal Degener 21(1–2):51–62. https://doi.org/10.1080/21678421.2019.1672749
Bauer PO, Goswami A, Wong HK, Okuno M, Kurosawa M, Yamada M, Miyazaki H, Matsumoto G, Kino Y, Nagai Y, Nukina N (2010) Harnessing chaperone-mediated autophagy for the selective degradation of mutant huntingtin protein. Nat Biotechnol 28(3):256–263. https://doi.org/10.1038/nbt.1608
DiFiglia M, Sapp E, Chase KO, Davies SW, Bates GP, Vonsattel JP, Aronin N (1997) Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 277(5334):1990–1993. https://doi.org/10.1126/science.277.5334.1990
Qi L, Zhang XD, Wu JC, Lin F, Wang J, DiFiglia M, Qin ZH (2012) The role of chaperone-mediated autophagy in huntingtin degradation. PLoS One 7(10):e46834. https://doi.org/10.1371/journal.pone.0046834
Vogiatzi T, Xilouri M, Vekrellis K, Stefanis L (2008) Wild type alpha-synuclein is degraded by chaperone-mediated autophagy and macroautophagy in neuronal cells. J Biol Chem 283(35):23542–23556. https://doi.org/10.1074/jbc.M801992200
Mak SK, McCormack AL, Manning-Bog AB, Cuervo AM, Di Monte DA (2010) Lysosomal degradation of alpha-synuclein in vivo. J Biol Chem 285(18):13621–13629. https://doi.org/10.1074/jbc.M109.074617
Malkus KA, Ischiropoulos H (2012) Regional deficiencies in chaperone-mediated autophagy underlie α-synuclein aggregation and neurodegeneration. Neurobiol Dis 46(3):732–744. https://doi.org/10.1016/j.nbd.2012.03.017
Alvarez-Erviti L, Rodriguez-Oroz MC, Cooper JM, Caballero C, Ferrer I, Obeso JA, Schapira AH (2010) Chaperone-mediated autophagy markers in Parkinson disease brains. Arch Neurol 67(12):1464–1472. https://doi.org/10.1001/archneurol.2010.198
Xilouri M, Vogiatzi T, Vekrellis K, Park D, Stefanis L (2009) Abberant alpha-synuclein confers toxicity to neurons in part through inhibition of chaperone-mediated autophagy. PLoS One 4(5):e5515. https://doi.org/10.1371/journal.pone.0005515
Yang Q, Mao Z (2009) The complexity in regulation of MEF2D by chaperone-mediated autophagy. Autophagy 5(7):1073–1074. https://doi.org/10.4161/auto.5.7.9824
Wilson MA, Collins JL, Hod Y, Ringe D, Petsko GA (2003) The 1.1-A resolution crystal structure of DJ-1, the protein mutated in autosomal recessive early onset Parkinson’s disease. Proc Natl Acad Sci U S A 100(16):9256–9261. https://doi.org/10.1073/pnas.1133288100
Kahle PJ, Waak J, Gasser T (2009) DJ-1 and prevention of oxidative stress in Parkinson’s disease and other age-related disorders. Free Radic Biol Med 47(10):1354–1361. https://doi.org/10.1016/j.freeradbiomed.2009.08.003
Andersson FI, Werrell EF, McMorran L, Crone WJ, Das C, Hsu ST, Jackson SE (2011) The effect of Parkinson’s-disease-associated mutations on the deubiquitinating enzyme UCH-L1. J Mol Biol 407(2):261–272. https://doi.org/10.1016/j.jmb.2010.12.029
Xilouri M, Brekk OR, Landeck N, Pitychoutis PM, Papasilekas T, Papadopoulou-Daifoti Z, Kirik D, Stefanis L (2013) Boosting chaperone-mediated autophagy in vivo mitigates α-synuclein-induced neurodegeneration. Brain 136(Pt 7):2130–2146. https://doi.org/10.1093/brain/awt131
Pang S, Chen D, Zhang A, Qin X, Yan B (2012) Genetic analysis of the LAMP-2 gene promoter in patients with sporadic Parkinson’s disease. Neurosci Lett 526(1):63–67. https://doi.org/10.1016/j.neulet.2012.07.044
Wang Y, Martinez-Vicente M, Krüger U, Kaushik S, Wong E, Mandelkow EM, Cuervo AM, Mandelkow E (2009) Tau fragmentation, aggregation and clearance: the dual role of lysosomal processing. Hum Mol Genet 18(21):4153–4170. https://doi.org/10.1093/hmg/ddp367
Chan B, Greenan G, McKeon F, Ellenberger T (2005) Identification of a peptide fragment of DSCR1 that competitively inhibits calcineurin activity in vitro and invivo. Proc Natl Acad Sci U S A 102(37):13075–13080. https://doi.org/10.1073/pnas.0503846102
Park JS, Kim DH, Yoon SY (2016) Regulation of amyloid precursor protein processing by its KFERQ motif. BMB Rep 49(6):337–342. https://doi.org/10.5483/bmbrep.2016.49.6.212
Bates G (2003) Huntingtin aggregation and toxicity in Huntington’s disease. Lancet 361(9369):1642–1644. https://doi.org/10.1016/s0140-6736(03)13304-1
Koga H, Martinez-Vicente M, Arias E, Kaushik S, Sulzer D, Cuervo AM (2011) Constitutive upregulation of chaperone-mediated autophagy in Huntington’s disease. J Neurosci 31(50):18492–18505. https://doi.org/10.1523/jneurosci.3219-11.2011
Martinez-Vicente M, Talloczy Z, Wong E, Tang G, Koga H, Kaushik S, de Vries R, Arias E, Harris S, Sulzer D, Cuervo AM (2010) Cargo recognition failure is responsible for inefficient autophagy in Huntington’s disease. Nat Neurosci 13(5):567–576. https://doi.org/10.1038/nn.2528
Gao FB, Almeida S, Lopez-Gonzalez R (2017) Dysregulated molecular pathways in amyotrophic lateral sclerosis-frontotemporal dementia spectrum disorder. Embo J 36(20):2931–2950. https://doi.org/10.15252/embj.201797568
Tamaki Y, Shodai A, Morimura T, Hikiami R, Minamiyama S, Ayaki T, Tooyama I, Furukawa Y, Takahashi R, Urushitani M (2018) Elimination of TDP-43 inclusions linked to amyotrophic lateral sclerosis by a misfolding-specific intrabody with dual proteolytic signals. Sci Rep 8(1):6030. https://doi.org/10.1038/s41598-018-24463-3
Thorburn A, Debnath J (2011) Targeting chaperone-mediated autophagy in cancer. Sci Transl Med 3(109):109ps145. https://doi.org/10.1126/scitranslmed.3003390
Saha T (2012) LAMP2A overexpression in breast tumors promotes cancer cell survival via chaperone-mediated autophagy. Autophagy 8(11):1643–1656. https://doi.org/10.4161/auto.21654
Lu TL, Huang GJ, Wang HJ, Chen JL, Hsu HP, Lu TJ (2010) Hispolon promotes MDM2 downregulation through chaperone-mediated autophagy. Biochem Biophys Res Commun 398(1):26–31. https://doi.org/10.1016/j.bbrc.2010.06.004
Quintavalle C, Di Costanzo S, Zanca C, Tasset I, Fraldi A, Incoronato M, Mirabelli P, Monti M, Ballabio A, Pucci P, Cuervo AM, Condorelli G (2014) Phosphorylation-regulated degradation of the tumor-suppressor form of PED by chaperone-mediated autophagy in lung cancer cells. J Cell Physiol 229(10):1359–1368. https://doi.org/10.1002/jcp.24569
Gomes LR, Menck CFM, Cuervo AM (2017) Chaperone-mediated autophagy prevents cellular transformation by regulating MYC proteasomal degradation. Autophagy 13(5):928–940. https://doi.org/10.1080/15548627.2017.1293767
Zhou J, Yang J, Fan X, Hu S, Zhou F, Dong J, Zhang S, Shang Y, Jiang X, Guo H, Chen N, Xiao X, Sheng J, Wu K, Nie Y, Fan D (2016) Chaperone-mediated autophagy regulates proliferation by targeting RND3 in gastric cancer. Autophagy 12(3):515–528. https://doi.org/10.1080/15548627.2015.1136770
Han Q, Deng Y, Chen S, Chen R, Yang M, Zhang Z, Sun X, Wang W, He Y, Wang F, Pan X, Li P, Lai W, Luo H, Huang P, Guan X, Deng Y, Yan J, Xu X, Wen Y, Chen A, Hu C, Li X, Li S (2017) Downregulation of ATG5-dependent macroautophagy by chaperone-mediated autophagy promotes breast cancer cell metastasis. Sci Rep 7(1):4759. https://doi.org/10.1038/s41598-017-04994-x
Hubbi ME, Hu H, Kshitiz AI, Levchenko A, Semenza GL (2013) Chaperone-mediated autophagy targets hypoxia-inducible factor-1α (HIF-1α) for lysosomal degradation. J Biol Chem 288(15):10703–10714. https://doi.org/10.1074/jbc.M112.414771
Wu JH, Guo JP, Shi J, Wang H, Li LL, Guo B, Liu DX, Cao Q, Yuan ZY (2017) CMA down-regulates p53 expression through degradation of HMGB1 protein to inhibit irradiation-triggered apoptosis in hepatocellular carcinoma. World J Gastroenterol 23(13):2308–2317. https://doi.org/10.3748/wjg.v23.i13.2308
Suzuki J, Nakajima W, Suzuki H, Asano Y, Tanaka N (2017) Chaperone-mediated autophagy promotes lung cancer cell survival through selective stabilization of the pro-survival protein, MCL1. Biochem Biophys Res Commun 482(4):1334–1340. https://doi.org/10.1016/j.bbrc.2016.12.037
Wang R, Liu Y, Liu L, Chen M, Wang X, Yang J, Gong Y, Ding BS, Wei Y, Wei X (2019) Tumor cells induce LAMP2a expression in tumor-associated macrophage for cancer progression. EBioMedicine 40:118–134. https://doi.org/10.1016/j.ebiom.2019.01.045
Ding ZB, Fu XT, Shi YH, Zhou J, Peng YF, Liu WR, Shi GM, Gao Q, Wang XY, Song K, Jin L, Tian MX, Shen YH, Fan J (2016) Lamp2a is required for tumor growth and promotes tumor recurrence of hepatocellular carcinoma. Int J Oncol 49(6):2367–2376. https://doi.org/10.3892/ijo.2016.3754
Warfel NA, Dolloff NG, Dicker DT, Malysz J, El-Deiry WS (2013) CDK1 stabilizes HIF-1α via direct phosphorylation of Ser668 to promote tumor growth. Cell Cycle 12(23):3689–3701. https://doi.org/10.4161/cc.26930
Aydin Y, Stephens CM, Chava S, Heidari Z, Panigrahi R, Williams DD, Wiltz K, Bell A, Wilson W, Reiss K, Dash S (2018) Chaperone-mediated autophagy promotes beclin1 degradation in persistently infected hepatitis C virus cell culture. Am J Pathol 188(10):2339–2355. https://doi.org/10.1016/j.ajpath.2018.06.022
Liu DX, Li PP, Guo JP, Li LL, Guo B, Jiao HB, Wu JH, Chen JM (2019) Exosomes derived from HBV-associated liver cancer promote chemoresistance by upregulating chaperone-mediated autophagy. Oncol Lett 17(1):323–331. https://doi.org/10.3892/ol.2018.9584
Khan MM, Nomura T, Kim H, Kaul SC, Wadhwa R, Zhong S, Pandolfi PP, Ishii S (2001) PML-RARalpha alleviates the transcriptional repression mediated by tumor suppressor Rb. J Biol Chem 276(47):43491–43494. https://doi.org/10.1074/jbc.C100532200
Zhang H, Guttikonda S, Roberts L, Uziel T, Semizarov D, Elmore SW, Leverson JD, Lam LT (2011) Mcl-1 is critical for survival in a subgroup of non-small-cell lung cancer cell lines. Oncogene 30(16):1963–1968. https://doi.org/10.1038/onc.2010.559
Tang J, Zhan MN, Yin QQ, Zhou CX, Wang CL, Wo LL, He M, Chen GQ, Zhao Q (2017) Impaired p65 degradation by decreased chaperone-mediated autophagy activity facilitates epithelial-to-mesenchymal transition. Oncogenesis 6(10):e387. https://doi.org/10.1038/oncsis.2017.85
Schneider JL, Villarroya J, Diaz-Carretero A, Patel B, Urbanska AM, Thi MM, Villarroya F, Santambrogio L, Cuervo AM (2015) Loss of hepatic chaperone-mediated autophagy accelerates proteostasis failure in aging. Aging Cell 14(2):249–264. https://doi.org/10.1111/acel.12310
Razidlo GL, Wang Y, Chen J, Krueger EW, Billadeau DD, McNiven MA (2013) Dynamin 2 potentiates invasive migration of pancreatic tumor cells through stabilization of the Rac1 GEF Vav1. Dev Cell 24(6):573–585. https://doi.org/10.1016/j.devcel.2013.02.010
Ferguson EC, Rathmell JC (2008) New roles for pyruvate kinase M2: working out the Warburg effect. Trends Biochem Sci 33(8):359–362. https://doi.org/10.1016/j.tibs.2008.05.006
Trencia A, Perfetti A, Cassese A, Vigliotta G, Miele C, Oriente F, Santopietro S, Giacco F, Condorelli G, Formisano P, Beguinot F (2003) Protein kinase B/Akt binds and phosphorylates PED/PEA-15, stabilizing its antiapoptotic action. Mol Cell Biol 23(13):4511–4521. https://doi.org/10.1128/mcb.23.13.4511-4521.2003
Garg AD, Dudek AM, Agostinis P (2013) Calreticulin surface exposure is abrogated in cells lacking, chaperone-mediated autophagy-essential gene, LAMP2A. Cell Death Dis 4(10):e826. https://doi.org/10.1038/cddis.2013.372
Xia HG, Najafov A, Geng J, Galan-Acosta L, Han X, Guo Y, Shan B, Zhang Y, Norberg E, Zhang T, Pan L, Liu J, Coloff JL, Ofengeim D, Zhu H, Wu K, Cai Y, Yates JR, Zhu Z, Yuan J, Vakifahmetoglu-Norberg H (2015) Degradation of HK2 by chaperone-mediated autophagy promotes metabolic catastrophe and cell death. J Cell Biol 210(5):705–716. https://doi.org/10.1083/jcb.201503044
You Y, Li WZ, Zhang S, Hu B, Li YX, Li HD, Tang HH, Li QW, Guan YY, Liu LX, Bao WL, Shen X (2018) SNX10 mediates alcohol-induced liver injury and steatosis by regulating the activation of chaperone-mediated autophagy. J Hepatol 69(1):129–141. https://doi.org/10.1016/j.jhep.2018.01.038
Ma SY, Sun KS, Zhang M, Zhou XM, Zheng XH, Tian SY, Liu YS, Chen L, Gao X, Ye J, Zhou XM, Wang JB, Han Y (2020) Disruption of Plin5 degradation by CMA causes lipid homeostasis imbalance in NAFLD. Liver Int. https://doi.org/10.1111/liv.14492
Sooparb S, Price SR, Shaoguang J, Franch HA (2004) Suppression of chaperone-mediated autophagy in the renal cortex during acute diabetes mellitus. Kidney Int 65(6):2135–2144. https://doi.org/10.1111/j.1523-1755.2004.00639.x
Cuervo AM, Hildebrand H, Bomhard EM, Dice JF (1999) Direct lysosomal uptake of alpha 2-microglobulin contributes to chemically induced nephropathy. Kidney Int 55(2):529–545. https://doi.org/10.1046/j.1523-1755.1999.00268.x
Qin B, He M, Chen X, Pei D (2006) Sorting nexin 10 induces giant vacuoles in mammalian cells. J Biol Chem 281(48):36891–36896. https://doi.org/10.1074/jbc.M608884200
Lock EA, Charbonneau M, Strasser J, Swenberg JA, Bus JS (1987) 2,2,4-Trimethylpentane-induced nephrotoxicity. II. The reversible binding of a TMP metabolite to a renal protein fraction containing alpha 2u-globulin. Toxicol Appl Pharmacol 91(2):182–192. https://doi.org/10.1016/0041-008x(87)90099-8
Venugopal B, Mesires NT, Kennedy JC, Curcio-Morelli C, Laplante JM, Dice JF, Slaugenhaupt SA (2009) Chaperone-mediated autophagy is defective in mucolipidosis type IV. J Cell Physiol 219(2):344–353. https://doi.org/10.1002/jcp.21676
Li Z, Wang C, Wang Z, Zhu C, Li J, Sha T, Ma L, Gao C, Yang Y, Sun Y, Wang J, Sun X, Lu C, Difiglia M, Mei Y, Ding C, Luo S, Dang Y, Ding Y, Fei Y, Lu B (2019) Allele-selective lowering of mutant HTT protein by HTT-LC3 linker compounds. Nature 575(7781):203–209. https://doi.org/10.1038/s41586-019-1722-1
Wang H, Tian C, Sun J, Chen LN, Lv Y, Yang XD, Xiao K, Wang J, Chen C, Shi Q, Shao QX, Dong XP (2017) Overexpression of PLK3 mediates the degradation of abnormal prion proteins dependent on chaperone-mediated autophagy. Mol Neurobiol 54(6):4401–4413. https://doi.org/10.1007/s12035-016-9985-0
Pedrozo Z, Torrealba N, Fernández C, Gatica D, Toro B, Quiroga C, Rodriguez AE, Sanchez G, Gillette TG, Hill JA, Donoso P, Lavandero S (2013) Cardiomyocyte ryanodine receptor degradation by chaperone-mediated autophagy. Cardiovasc Res 98(2):277–285. https://doi.org/10.1093/cvr/cvt029
Fidziańska A, Walczak E, Walski M (2007) Abnormal chaperone-mediated autophagy (CMA) in cardiomyocytes of a boy with Danon disease. Folia Neuropathol 45(3):133–139
Endo Y, Furuta A, Nishino I (2015) Danon disease: a phenotypic expression of LAMP-2 deficiency. Acta Neuropathol 129(3):391–398. https://doi.org/10.1007/s00401-015-1385-4
Métrailler S, Schorderet DF, Cottet S (2012) Early apoptosis of rod photoreceptors in Rpe65(-/-) mice is associated with the upregulated expression of lysosomal-mediated autophagic genes. Exp Eye Res 96(1):70–81. https://doi.org/10.1016/j.exer.2011.12.019
Li Y, Lu L, Luo N, Wang YQ, Gao HM (2017) Inhibition of PI3K/AKt/mTOR signaling pathway protects against d-galactosamine/lipopolysaccharide-induced acute liver failure by chaperone-mediated autophagy in rats. Biomed Pharmacother 92:544–553. https://doi.org/10.1016/j.biopha.2017.05.037
Das S, Seth RK, Kumar A, Kadiiska MB, Michelotti G, Diehl AM, Chatterjee S (2013) Purinergic receptor X7 is a key modulator of metabolic oxidative stress-mediated autophagy and inflammation in experimental nonalcoholic steatohepatitis. Am J Physiol Gastrointest Liver Physiol 305(12):G950-963. https://doi.org/10.1152/ajpgi.00235.2013
Lee CH, Lee KH, Jang AH, Yoo CG (2017) The impact of autophagy on the cigarette smoke extract-induced apoptosis of bronchial epithelial cells. Tuberc Respir Dis (Seoul) 80(1):83–89. https://doi.org/10.4046/trd.2017.80.1.83
Handa K, Kanno H, Matsuda M, Sugaya T, Murakami T, Prudnikova M, Ozawa H, Itoi E (2020) Chaperone-mediated autophagy after spinal cord injury. J Neurotrauma. https://doi.org/10.1089/neu.2019.6820
Su M, Guan H, Zhang F, Gao Y, Teng X, Yang W (2016) HDAC6 regulates the chaperone-mediated autophagy to prevent oxidative damage in injured neurons after experimental spinal cord injury. Oxid Med Cell Longev 2016:7263736. https://doi.org/10.1155/2016/7263736
Dohi E, Tanaka S, Seki T, Miyagi T, Hide I, Takahashi T, Matsumoto M, Sakai N (2012) Hypoxic stress activates chaperone-mediated autophagy and modulates neuronal cell survival. Neurochem Int 60(4):431–442. https://doi.org/10.1016/j.neuint.2012.01.020
Hu MM, Yang Q, Xie XQ, Liao CY, Lin H, Liu TT, Yin L, Shu HB (2016) Sumoylation promotes the stability of the DNA sensor cGAS and the adaptor STING to regulate the kinetics of response to DNA virus. Immunity 45(3):555–569. https://doi.org/10.1016/j.immuni.2016.08.014
Singh V, Finke-Isami J, Hopper-Chidlaw AC, Schwerk P, Thompson A, Tedin K (2017) Salmonella co-opts host cell chaperone-mediated autophagy for intracellular growth. J Biol Chem 292(5):1847–1864. https://doi.org/10.1074/jbc.M116.759456
Napolitano G, Johnson JL, He J, Rocca CJ, Monfregola J, Pestonjamasp K, Cherqui S, Catz SD (2015) Impairment of chaperone-mediated autophagy leads to selective lysosomal degradation defects in the lysosomal storage disease cystinosis. EMBO Mol Med 7(2):158–174. https://doi.org/10.15252/emmm.201404223
Funding
This work was supported by Qingdao Applied Basic Research Program Youth Project: 19-6-2-59-cg and China Postdoctoral Science Foundation Funded Project: 2015M57074, 2016T90612.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that this article content has no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
#Zhaozhong Liao and Bin Wang have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Liao, Z., Wang, B., Liu, W. et al. Dysfunction of chaperone-mediated autophagy in human diseases. Mol Cell Biochem 476, 1439–1454 (2021). https://doi.org/10.1007/s11010-020-04006-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-020-04006-z