Abstract
An mRNA transcript that codes for a Disintegrin and a Metalloprotease from a venom gland of the viper Bothrops ammodytoides was isolated. Vector pCR®2.1-TOPO was used to clone the Disintegrin transcript and then subcloned in the pET-28a vector to express the protein in the E. coli strain BL21. The recombinant disintegrin, HisrDisintegrin, comprises 38 residues at the N-terminal and 75 residues of the Disintegrin, including 14 cysteines or 7 disulfide bonds. HisrDisintegrin was obtained soluble in the intracellular fraction and separated by affinity chromatography. The experimental molecular mass of HisrDisintegrin, 11,750.8 Da, agreed with its theoretical value, including the cleavage of a Met residue caused by a bacterial post-translational modification. HisrDisintegrin was folded in in vitro conditions and then purified, finding a main fraction with a comparable molecular mass. The HisrDisintegrin contains an alpha/beta structure, as observed by circular dichroism. Its biological activity demonstrated the growth reduction of the human endothelial (HMEC-1) cells and the human mammary gland adenocarcinoma (MDA-MB-231). Furthermore, HisrDisintegrin inhibits the adhesion of HMEC-1 and MDA-MB-231 cells to the proteins of the extracellular matrix (laminin (LN), fibronectin (FN), and vitronectin (VN)).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bothrops ammodytoides is the southernmost situated viper in the world that populates from the cold Patagonia of Argentina to the warm desert regions of “El Gran Chaco” and the dry “Pampeana” region. It is a small viper that could cause the typical complications of Bothrops envenomations, which causes hemorrhage, dermonecrosis, and inflammatory-edematogenic effects in mice. It also exhibits procoagulant activity on human plasma but little or no thrombin-like activity on bovine fibrinogen (de Roodt et al. 2000). The venom of B. ammodytoides contains metallo-proteases, and other enzymes such as serine-proteases and phospholipases type A2 (PLA2), which contribute to the toxic effects of this viper (Clement et al. 2012a, 2019a, b). On the other hand, disintegrins are a large family of polypeptides generally obtained from viperid and crotalid venoms. They inhibit the integrin activity involved in fibrinogen-dependent platelet aggregation and in cell adhesion (Vasconcelos et al. 2021). Disintegrins have typical motifs, such as the triad ArgGlyAsp (RGD), which has been exploited as an active structure for the production of anticancer drugs such as Eptifibatide (Phillips and Scarborough 1997). Here, we report a C-terminal region of a metallo-protease from the venom gland of B. ammodytoides containing the disintegrin motif which was cloned from cDNA and heterologously expressed. The recombinant disintegrin had coagulant activity; although its primary structure does not contain the typical RGD motif, it was able to inhibit human endothelial (HMEC-1) and human mammary gland adenocarcinoma (MDA-MB-231) cell lines. Hence, we provide the proof of concept that this recombinant disintegrin could be used as a protein lead for anticancer drug development.
Materials and Methods
Venom Gland
Adult specimens of B. ammodytoides were maintained in good health in plastic cages (27 ºC and 12 h light-dark cycle). Venom was collected by manual extraction, dried under vacuum, and kept at -20ºC. An anesthetized (ketamine-xylazine) B. ammodytoides was subjected to a surgical extraction of one of the two venom glands. RNAlater→ (Thermofisher, Asheville, NC, USA) was used to treat the extracted venom gland and stored at -20ºC. The treated animal stayed healthy and recovered.
Plasmids, Bacterial Strains, and Enzymes
Plasmid pCR®2.1-TOPO® (Invitrogen, CA, USA) was used for cloning the Disintegrin gene and pET28a (Invitrogen, CA, USA) for production of the 6His-tagged recombinant HisrDisintegrin. DNA cloning and plasmid propagation were conducted in Escherichia coli strain XL1-Blue. The expression of the recombinant Disintegrin was performed in E. coli strain BL21. Taq polymerase, T4 DNA ligase, restriction enzymes, and Factor Xa protease (FXa) were provided by New England Biolabs (NEB, MA, USA).
RNA Extraction and Gene Assembly
A single venom gland of B. ammodytoides was used to extract the total RNA using the “Total RNA Isolation System” (Quiagen, CA, USA). cDNA was obtained with 4 µg of Total RNA (3’RACE kit, Invitrogen, Carlsbad, CA, U.S.A.). The specific oligonucleotides (Disintegrin-dir: GAG GTG GGA GAA GAA TGT G, Tm 58 ºC and Disintegrin-Rv: GAG TGT CCT GCA GAT GTC TTC, Tm 54 ºC) were conceived to obtain the Disintegrin transcript (Clement et al., 2012) in a PCR reaction (0.4 µM of each oligonucleotide, 200 µM dNTPs, 1X polymerase buffer, and 1 U of Taq polymerase. PCR conditions: 94 °C/30 s, 55 °C/2 min, and 72 °C/2min. Final step 72 °C/7 min). The amplified DNA fragment (~ 225 bp) was extracted from the agarose gel using the High Pure PCR Product Purification Kit (Roche®, Basel, Switzerland).
The vector pCR 2.1-TOPO® (TOPO TA Cloning® Kits, Invitrogen; Carlsbad, CA, U.S.A.) was used to clone the purified DNA fragment. E. coli XL1Blue cells were transformed with the new plasmid, grown in Petri dishes with Luria Broth (LB) agar added with ampicillin (100 µg/mL), and incubated at 37 ºC, 14 h. Colonies were evaluated by colony PCR with oligonucleotides M13 forward (5’- GTAAAACGACGGCCAGT-3’) and M13 reverse (5’- CAGGAAACAGCTATGAC-3’). Plasmids from positive colonies with the expected DNA size were purified and sequenced at the Institute of Biotechnology, UNAM.
Expression Plasmid Construction
A TOPO recombinant plasmid with the confirmed sequence was used to amplify the Disintegrin peptide by PCR with designed oligonucleotides, containing the BamHI site and FXa cleavage site at the 5’ end and the HindIII site and stop codons at the 3’ end (Dis-pET28-F: 5’-GAGAGGATCCATCGAGGGAAGGGAGGTG-3’; Dis-pET28-Rev: 5’-CAGATGTCTTCTAGTAACTGCAGCCAAGCTTTCTC-3’). The amplified gene and the pET28a expression plasmid (Invitrogen, Carlsbad, CA, U.S.A.) were digested with BamHI and HindIII enzymes. After digestion, the Disintegrin gene and vector were run and purified from the agarose gel. The digested DNAs were mixed in a ratio of 1:5, and then T4 DNA Ligase Buffer (1X) and T4 ligase (1 Weiss U) (Thermo-Fisher Co.; San José, CA, U.S.A.) were added. The ligation reaction was kept at 16 ºC 16 h. Chemically competent E. coli XL1-Blue cells were transformed with the ligation product (pET28a-Disintegrin). Transformed cells were grown on LB agar and added with 50 µg/mL of kanamycin (Sigma, St. Louis, MO, U.S.A.). After incubation (37 ºC, 14 h), some colonies were evaluated by colony PCR using the oligonucleotides 1522-Fwd (5’-AGATCTCGATCCCGC-3’) and 1522-Rev (5’-GACCCGTTTAGAGGC-3’). Positive colonies were seeded in 3 mL of LB, incubated overnight at 37ºC and 180 rpm, and then subjected to plasmid purification and sequencing at the Institute of Biotechnology, UNAM, Mexico.
Expression and Purification of HisrDisintegrin
E. coli strain BL21 harboring the pET28a-HisrDisintegrin expression vector was grown at 37ºC in LB medium (50 µg/mL kanamycin). When the culture absorbance (600 nm) was close to 0.8 units, 1 mM IPTG (isopropyl-β-D-thiogalactopyranoside) was added as the inductor and the culture was incubated for 18 h at 16ºC, 200 rpm. Afterward, the culture was centrifuged at 8,000 g for 20 min (JA-10 rotor, Beckman model J2-21). Cells were resuspended in wash buffer (0.05 M Tris-HCl, pH 8.0) and disrupted at 35 kpsi (One Shot cell disruptor, Constant Systems®). The mixture was centrifuged at 12,000 rpm for 15 min (JA–20 rotor, Beckman Coulter Avanti J-30I®). Soluble and insoluble fractions were analyzed by SDS-PAGE.
The recombinant HisrDisintegrin, found in the soluble fraction, was purified by Ni-nitrilotriacetic acid (Ni-NTA) affinity chromatography (Qiagen, CA, USA) using wash buffer and elution buffer B (0.05 M Tris-HCl buffer, 300 mM imidazole, pH 8.0). The elution buffer was removed by reverse-phase HPLC (RP-HPLC) using a C18 analytical reverse-phase column (Supelco 4.6 × 250 mm, 5 μm). The HPLC ran a separation program from 0 to 60% solvent B (0.1% TFA in acetonitrile) for 60 min at 1 mL/min. The fractions were detected at 230 nm. Collected fractions were dried under vacuum. Molecular masses were determined in a Thermo Scientific LCQ Fleet ion trap mass spectrometer (San Jose, CA) using a Surveyor MS syringe pump delivery system and the data acquisition and deconvolution were performed on the Xcalibur Windows NT PC data system.
Circular Dichroism
The secondary structure of the recombinant HisrDisintegrin was analyzed by Circular dichroism (CD) spectroscopy. The spectrum was obtained at room temperature in 1 mm path quartz cells with a wavelength range of 185 to 260 nm (Jasco J-1500 spectrometer, Japan). Data were gathered at a 50 nm/min rate with 0.1 nm intervals. A 0.25 mg/mL protein solution was prepared in 60% trifluoroethanol (TFE) because it enhanced the secondary structure. The average CD corresponded to three CD measures. The secondary structure percentages were then analyzed using online algorithms provided by BeStSel (Beta Structure Selection, webserver http://bestsel.elte.hu/index.php (accessed on 30 January 2023) (Micsonai et al. 2018).
Cell Lines and Culture Conditions
The American Type Culture Collection (ATCC, Manassas, VA, USA) sold the cell lines HMEC-1 (human endothelial) and MDA-MB-231 (human mammary gland adenocarcinoma). HMEC-1 cells were grown in MCDB-131 medium with 10% Fetal Bovine Serum (FBS), 10 ng/mL epidermal growth factor (Sigma-Aldrich), and 1 µg/mL hydrocortisone, then incubated at 37 °C in a humid chamber containing 5% CO2. MDA-MB-231 cells were grown in L-15 Leibovitz medium with 10% fetal bovine serum (FBS) and placed in a CO2-free incubator at 37 °C with humid air.
Cytotoxicity Assay
MDA-MB-231 and HMEC-1 cells were subjected to different concentrations of HisrDisintegrin, and its cytotoxicity was assessed by the MTT assay (3-(4,5-dimethylthiazol-2-yl)-2,5-diphe-nyltetrazolium bromide, Sigma-Aldrich) (Mosmann 1983). Briefly, 100 µL of trypsinized cells (2.0 × 105 cells/mL) in complete media were seeded in each well of a 96-well plate. The cells were treated with HisrDisintegrin at different concentrations (from 0.06 to 16 µM or PBS pH 7.4 as a negative control). The plate was incubated at 37 °C for 24 h. Afterward, 100 µL of MTT reagent (0.5 mg/mL) prepared in each cell culture medium were added to each well with cells and kept at 37 °C for 3 h. 200 µL of isopropyl alcohol and 0.4 N HCl were added to each well after the MTT remotion. The plate was kept in the dark for 30 min at 25 °C. Absorbance was measured at 570 nm (HTS 7000 Bioassay Reader, Perkin Elmer, Waltham, MA, USA). Viable cells, as a percentage, were determined by relating the number of viable cells of the treated cells to the control group (untreated cells) (Zakraoui et al. 2017). The experiment was repeated three times in triplicate at each HisrDisintegrin concentration.
Cell Adhesion Inhibition Assay
The binding inhibition of HMEC-1 and MDA-MB-231 cells to laminin, vitronectin, and fibronectin, caused by HisrDisintegrin was assessed using a previously established protocol (Hammouda et al. 2016; Lima-Dos-Santos et al. 2015; Wierzbicka-Patynowski et al. 1999). 100 µL (1 µg/mL) of either fibronectin, vitronectin, or laminin were added to each well of a 96-well plate, in triplicate, and left at 4 °C overnight. The wells were then blocked with 200 µL of a culture medium supplemented with 2% bovine serum albumin (BSA) at 37 °C for 1.5 h. The cells were harvested, counted, and resuspended in a medium containing 1% BSA at 5 × 105 cells/mL. Various concentrations of HisrDisintegrin (0.03, 0.06, 0.125, 0.25, 0.5, and 1 µM) were added to the cell suspension and incubated at 37 °C for 2 h; PBS was used as the negative control. After removing the blocking solution, 100 µL of the cell/HisrDisintegrin mixture and the negative control were added to the fibronectin-, vitronectin-, and laminin-coated wells and incubated at 37 °C for 2 h. The wells were washed three times, and 200 µL of L15 or MCDB-131 medium containing 1.5 mg/mL MTT and 1% BSA were added to the wells and incubated at 37 °C for 3 h. Cell lysis was achieved by adding 100 µL of dimethyl sulfoxide, and gently mixed. Absorbance was read at 570 nm. [(absorbance of negative control - absorbance of cell/HisrDisintegrin sample)/absorbance of negative control] × 100 was used to determine the percentage of inhibition. The experiment was repeated three times, and each HisrDisintegrin concentration for both cell lines was evaluated in triplicate.
Wound Healing Assay
MDA-MB-231 or HMEC-1 cells were plated (2 × 105 cells/mL) in a 96-well microtiter plate and incubated at 37 °C overnight. The confluent monolayer was scratched with a sterile 200 µL pipet tip at the midline of each well (Galan et al. 2008; Ren et al. 2006). A specific culture medium was used to wash the detached cells. The remaining cells were treated with 200 µL of fresh culture medium containing 10% FBS. 100 µL of PBS 0.01 M, pH 7.4, or HisrDisintegrin 0.04 µM prepared in the same PBS were added to the control and treatment groups to stimulate cell migration. The cells were then incubated at 37 °C in a humidified chamber, and images were captured at 0, 12, and 24 h using a microscope. The percentage of cell migration was calculated using the equation: [(C – E)/C] X 100, where C is the distance (µm) between the cell edges at t = 0 h of the control, and E is the distance between the cell edges of the monolayer at the final time (t = 24 h) of the incubation. The images were analyzed using Image J software, and the data were plotted as the percentage of cell migration of the control group vs. the treatment with HisrDisintegrin. The experiment was repeated three times, and the assays were performed in triplicate.
Statistics
Results were expressed as mean and standard deviation, or as mean with 95% confidence intervals. For all the statistical methods the software Prism 4.0 (Graph Pad Inc., San Diego, CA) was used.
Results and Discussion
Isolation and Sequence Determination of HisrDisintegrin
The cDNA for HisrDisintegrin with the expected size (225 bp) was amplified by PCR, cloned in pCR®2.1-TOPO® plasmid, and then in the expression vector pET-28a. The amino acid sequence of HisrDisintegrin is 100% identical to the disintegrin fractions of the ADAM (a disintegrin and metalloproteinase) proteins named bothropasin and catrocollastatin isolated from the venoms of B. jajaraca and Crotalus atrox, respectively (Table 1).
The disintegrin from B. ammodytoides seems to belong to the group 1 of the a disintegrin and metalloproteinase (ADAM) family of proteins including the snake venom metalloproteinases (ADAM/SVPM). They are also classified according to the number of residues and disulfide bridges, in a large group that consist of more than 70 residues cross-linked by seven disulfide bridges (Vasconcelos et al. 2021). The disintegrin from B. ammodytoides is distinguished from other snake disintegrin structures such as rhodostomin and bitistatin from groups 3 and 2, respectively, because of their amino acid identity and their content in disulfide connections (Vasconcelos et al. 2021).
Expression, Purification, and Protein Folding of HisrDisintegrin
The gene encoding HisrDisintegrin was assembled and subcloned into the expression vector pET-28a (Fig. S1), which produces N-terminal 6His-tagged proteins, for a straightforward purification of the recombinant products by affinity column chromatography Ni-NTA (Ni-nitrilotriacetic acid). A cleavage site for FXa was added between the 6His-tag and the HisrDisintegrin, to separate the 6His-tag from the HisrDisintegrin if necessary (Fig. S1). HisrDisintegrin was found in soluble fraction after expression in E. coli BL21 strain, and it was recovered by Ni-NTA (Fig. 1).
Expression of HisrDisintegrin in E. coli BL21 cells and its purification. (A) SDS-PAGE Expression. First lane: molecular weight markers (kDa); Lane 1: cells before IPTG induction; Lane 2: cells induced with IPTG; Lane 3: soluble fraction; Lane 4: insoluble fraction. (B) SDS-PAGE Purification. First Lane: molecular weight marker (kDa); Lane 1: recirculating; Lane 2: 35 mM Imidazole wash; Lane 3: 300 mM imidazole elution
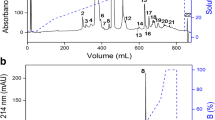
The heterologous expression of HisrDisintegrin in the soluble fraction and its purification process was corroborated by a western-blot assay (anti-6His-tag antibody coupled to alkaline phosphatase). HisrDisintegrin was purified by RP-HPLC, and a fraction with retention time around 35 min or 35% acetonitrile was collected (Fig. 2). The fraction also showed a similar apparent molecular mass SDS-PAGE (15/4%). The protein yield of the HisrDisintegrin fraction was 0.5 mg/L.
Chromatographic separation of HisrDisintegrin. RP-HPLC purification of HisrDisintegrin from the soluble fraction, after Ni-NTA affinity chromatography, using a C18 analytical reverse-phase column (Supelco 4.6 × 250 mm, 5 μm). The separation gradient used (dotted line) was from 0 to 60% solvent B (0.1% TFA in acetonitrile) for 60 min at 1 mL/min. The fractions were detected at 230 nm
Furthermore, the experimental molecular mass of the fraction obtained by mass spectrometry was 11,750.8 Da (oxidized form) (Fig. S2) and agreed to the theoretical molecular mass of HisrDisintegrin after the cleavage of a Met residue caused by a bacterial post-translational modification. Therefore, the fraction obtained from RP-HPLC corresponded to the oxidized cystine isoform of the HisrDisintegrin. Multiple cysteine-rich protein isoforms generated during heterologous expression have been reported (Clement et al. 2012b, 2015, 2016; Estrada et al. 2007). HisrDisintegrin contains 14 cysteines and could form up to 135,135 isoforms considering scrambling disulfide pairing possibilities. Clearly, in the snake venom gland, the molecular machinery in charge of the disintegrin expression directs the formation of a correct folding.
Secondary Structure of HisrDisintegrin
Here the CD spectrum of HisrDisintegrin was obtained in solution using 60% TFE, which promotes hydrogen bond formation mimicking in-cell solvation for protein folding (Fig. 3). According to the CD deconvolution program BeStSel (Micsonai et al. 2018), the secondary structure content (%) of this recombinant disintegrin was α-helix 18.6, β-antiparallel 16.3, β-parallel 0, β-turns 14.2 and random structures 51. So far, this recombinant disintegrin contain a significative proportion of α-helix and antiparallel structures, here characterized for having negative ellipticities at 203–229 nm giving a characteristics CD spectrum of an α/β protein. The root-mean-square deviation (RMSD) of the HisrDisintegrin CD experimental data compared to the fitted data predicted by the model was 0.22.
Although a reliable estimation of secondary structure content from the CD spectra cannot be calculated for proteins with the mixed α and β elements such secondary structure estimation, it could be used as a relative measure of structural changes of an unfolded protein (Khrapunov 2009). So that, the CD of a protein could be a reliable mean for determination of secondary structure of proteins that has been obtained using recombinant techniques (Greenfield 2006). The CD analysis of HisrDisintegrin showed elements of α/β protein, and we can consider that it is folded, utmost if it contains disulfide connections that tightly restrict internal changes. Furthermore, to compare the secondary structure of HisrDisintegrin, the CD spectrum of a classical α/β protein, the scorpion neurotoxin AaH2 (Fig. 3). Similarly, according to the BeStSel deconvolution program (Micsonai et al. 2018), the secondary structure content (%) of AaH2 was α-helix 86.8, β-antiparallel 8.5, β-parallel 0, β-turns 4.7, and random structures 0. AaH2 has 66 amino acids and four disulfide bridges, compared to the 75 amino acids and the seven disulfide connections of Disintegrin. Although AaH2 and Disintegrins are different in their mode of action, they share common structural patterns, such as a large content of α-helix 86.8 and β-antiparallel 8.5; however, in the case of the recombinant Disintegrin, it is less structured for the large content of random structures, which in most analyzed Disintegrins have shown disordered structures; some examples are the 3D structures of rhodostomin (2PJF) and bothropasin (3DSL), obtained by NMR and X-ray diffraction methods, respectively. The CD data of proteins is challenging to compare with X-ray crystallography or NMR (Greenfield 2006).
Cytotoxicity
The MTT assay was used to assess the effect of HisrDisintegrin on HMEC-1 and MDA-MB-231 cells. Cells treated with different concentrations for 24 h of HisrDisintegrin (from 0.0625 to 16 µM) decreased their cell viability in a dose-dependent manner. HisrDisintegrin at 16, 8, and 4 µM significantly decreased cell viability by 70.8, 67.3, and 54.9% of MDA-MB-231 and 52.5, 38.6, and 36.4% of HMEC-1 cells, respectively (p < 0.001). HisrDisintegrin resulted in 80% cell viability at 1 µM and lower concentrations (Fig. 4). Consequently, the highest selected concentration to evaluate the inhibitory activities of HisrDisintegrin was 1 µM.
Commonly, isolated Disintegrins are known as components without toxicity (Walsh and Marcinkiewicz 2011). However, at cellular level, HisrDisintegrin showed higher cytotoxicity compared to other isolated snake venom disintegrins such as totonacin (Rivas Mercado et al. 2020), tzabcanin (Saviola et al. 2016), vicrostatin (Minea et al. 2012). Non cytotoxic HisrDisintegrin concentrations selected for biological activities were similar to those used for other Bothrops disintegrins, such as Neuwiedin from B. neuwiedi (Lima-Dos-Santos et al. 2015), Colombistatin from B. colombiensis (Sanchez et al. 2009), Bothrasperin from B. asper (Pinto et al. 2003), and DisBa-01 from B. alternatus venom (Kauskot et al. 2008).
Effect of HisrDisintegrin from B. ammodytoides on HMEC-1 and MDA-MB-231 Cell Adhesion to Extracellular Matrix Proteins (Laminin, Vitronectin, and Fibronectin)
Inhibition of MDA-MB-231 cells adhesion to the extracellular matrix (ECM) proteins laminin (LN), vitronectin (VN), and fibronectin (FN) after 24 h of exposure to HisrDisintegrin at 1 µM was 33.2 ± 2.7, 64.3 ± 1.6, and 76.3 ± 1.6%, respectively. On the other hand, endothelial HMEC-1 cells showed increased susceptibility to the same extracellular matrix proteins blockade. HisrDisintegrin at 1 µM blocked HMEC-1 cells’ adhesion to FN, VN, and LN at 90.4 ± 2.2, 93 ± 2, and 36.3 ± 2%, respectively, and even showed a high inhibitory effect higher than 80% on VN also at 0.5, 0.25 and 0.125 µM followed by FN.
Disintegrins can impede the bonding of cells to surfaces covered with ECM proteins (Lima-Dos-Santos et al. 2015); thus HisrDisintegrin avoided the cell adhesion of MDA-MB-231 and HMEC-1 to the ECM proteins FN, VN, and LN. MDA-MB-231 cells are currently studied in cancer and express integrins in high levels, such as αv and β1, β5, and αvβ5. On the other hand, HMEC-1 cells are used in angiogenesis studies, expressing crucial integrins (v. g. αv, α1, 2, 3, 4, 5, 6, and β1, 3, 4, and 5) (Lucena et al. 2012; Minea et al. 2012; Taherian et al. 2011). HisrDisintegrin at concentrations below 1 µM presented a low cytotoxicity effect. HisrDisintegrin concentrations ranging from 1 to 0.03 µM were selected for adhesion inhibition assay. Our results demonstrated a significant inhibitory effect of HisrDisintegrin on the adhesion of MDA-MB-231 and HMEC-1 cells to VN and FN. A higher HMEC-1 cells inhibitory effect compared to other reported snake venom disintegrins such as tzabcanin, which inhibits about 70% A-375 cell adhesion to VN (Saviola et al. 2016), totonacin inhibits 60 and 44% MDA-MB-231 and HMEC-1 cell adhesion to VN (Rivas Mercado et al. 2020), more differences are noted when it is compared to other Bothrops disintegrins as DisBa-01 known by its high affinity on FN binding receptor αVβ3 integrin (Kauskot et al. 2008), or Colombistatin that has a higher inhibitory effect on FN and collagen I (Montenegro et al. 2017; Sanchez et al. 2009). Similarly, Bothrasperin inhibits endothelial t-End and melanoma B16 cells to VN, FN, and collagen 70 to 80% (Pinto et al. 2003), with lower inhibitory activity if it is compared to HisrDisintegrin inhibitory effect on VN and FN on HMEC-1 cells (Fig. 5). As VN and FN were more inhibited ligands in both HMEC-1 and MDA-MB-231 cells, and considering VN targets (integrins αVβ3, α8β1, αVβ5, αVβ1, and αIIbβ3) (Barczyk et al. 2010), and FN targets (integrins: αIIbβ3, α4β1, α4β7, α5β1, α8β1, αvβ1, αvβ3, αvβ6, and αvβ8) (Saviola et al. 2016), the results suggest HisrDisintegrin antagonized one or more of the five mentioned integrin VN receptors, and one or more of the nine integrins FN receptors. Finally, LN stands as the ECM less affected protein as HisrDisintegrin inhibitory effect on MDA-MB-231 and HMEC-1 cells was just about 30% at the highest disintegrin concentration proved, 1 µM. LN targets seven integrin receptors: α1β1, α2β1, α3β1, α6β1, α7β1, α10β1, and α6β4 (Saviola et al. 2016).
Evaluation of the Effect of HisrDisintegrin from B. ammodytoides on MDA-MB-231 and HMEC-1 Cell Migration
An in vitro wound healing assay assessed cell migration, where the cells were scraped from the wells’ bottom. Post a 24-hour incubation period, HisrDisintegrin at a concentration of 0.04 µM effectively restrained MDA-MB-231 and HMEC-1 cell migration by 87.8 ± 4% and 76.5 ± 2.5%, respectively (Fig. 6).
The observation that HisrDisintegrin inhibits cell migration proposes that αvβ3 integrin is blocked, although it doesn’t exclude the possibility that other integrins are targeted. For example, αvβ5 is prominently expressed in MDAMB-231 cells and is also known to facilitate cancer cell migration (Dang et al. 2006; Pecheur et al. 2002; Rolli et al. 2003; Sloan et al. 2006; Taherian et al. 2011). Similar results on cell migration, especially regarding to high 80–90% inhibition level have been reported for few other viperid snake venom disintegrins, such as crotatroxin 2, vicrostatin r-viridistatin 2, and r-mojastin 1 (Galan et al. 2008; Lucena et al. 2011; Minea et al. 2012).
Concluding Remarks
This work is the first approach to demonstrate the use of the recombinant HisrDisintegrin from B. ammodytoides with the capacity to inhibit cell migration and cell adhesion to ECM proteins. The use of recombinant proteins is becoming more common nowadays in the search for novel leads with clinical applications (Alvarenga et al. 2014; Espino-Solis et al. 2009). The recombinant HisrDisintegrin could be used as a model to find short protein domains with anti-angiogenesis and anti-metastasis effects to have affordable synthetic precursors for drug synthesis and low-cost medicaments.
Data Availability
Data will be made available on request.
References
Alvarenga LM, Zahid M, di Tommaso A, Juste MO, Aubrey N, Billiald P, Muzard J (2014) Engineering venom’s toxin-neutralizing antibody fragments and its therapeutic potential. Toxins 6:2541–2567. https://doi.org/10.3390/toxins6082541
Barczyk M, Carracedo S, Gullberg D (2010) Integrins Cell Tissue Res 339:269–280. https://doi.org/10.1007/s00441-009-0834-6
Clement H, Costa de Oliveira V, Zamudio FZ, Lago NR, Valdez-Cruz NA, Valle B, Hajos M, Alagon SE, Possani A, de Roodt LD, A.R (2012a) Isolation, amino acid sequence and biological characterization of an “aspartic-49” phospholipase A(2) from Bothrops (Rhinocerophis) ammodytoides venom. Toxicon 60:1314–1323. https://doi.org/10.1016/j.toxicon.2012.08.019
Clement H, Olvera A, Rodriguez M, Zamudio F, Palomares LA, Possani LD, Odell GV, Alagon A, Sanchez-Lopez R (2012b) Identification, cDNA cloning and heterologous expression of a hyaluronidase from the tarantula Brachypelma vagans venom. Toxicon 60:1223–1227. https://doi.org/10.1016/j.toxicon.2012.08.018
Clement H, Flores V, Diego-Garcia E, Corrales-Garcia L, Villegas E, Corzo G (2015) A comparison between the recombinant expression and chemical synthesis of a short cysteine-rich insecticidal spider peptide. J Venom Anim Toxins Incl Trop Dis 21:19. https://doi.org/10.1186/s40409-015-0018-7
Clement H, Flores V, De la Rosa G, Zamudio F, Alagon A, Corzo G (2016) Heterologous expression, protein folding and antibody recognition of a neurotoxin from the mexican coral snake Micrurus laticorallis. J Venom Anim Toxins Incl Trop Dis 22:25. https://doi.org/10.1186/s40409-016-0080-9
Clement H, Corrales-Garcia LL, Bolanos D, Corzo G, Villegas E (2019a) Immunogenic Properties of recombinant enzymes from Bothrops ammodytoides towards the generation of neutralizing antibodies against its own venom. Toxins 11:702. https://doi.org/10.3390/toxins11120702
Clement H, Corzo G, Neri-Castro E, Arenas I, Hajos S, de Roodt AR, Villegas E (2019b) cDNA cloning, heterologous expression, protein folding and immunogenic properties of a phospholipase A2 from Bothrops ammodytoides venom. Protein Expr Purif 154:33–43. https://doi.org/10.1016/j.pep.2018.09.004
Dang D, Bamburg JR, Ramos DM (2006) Alphavbeta3 integrin and cofilin modulate K1735 melanoma cell invasion. Exp Cell Res 312:468–477. https://doi.org/10.1016/j.yexcr.2005.11.011
de Roodt AR, Dolab JA, Hajos SE, Gould E, Dinapoli H, Troiano JC, Gould J, Dokmetjian JC, Carfagnini JC, Fernandez T, Amoroso M, Segre L, Vidal JC (2000) Some toxic and enzymatic activities of Bothrops ammodytoides (yarara nata) venom. Toxicon 38:49–61
Espino-Solis GP, Riano-Umbarila L, Becerril B, Possani LD (2009) Antidotes against venomous animals: state of the art and prospectives. J Proteom 72:183–199. https://doi.org/10.1016/j.jprot.2009.01.020
Estrada G, Garcia BI, Schiavon E, Ortiz E, Cestele S, Wanke E, Corzo LDP, G (2007) Four disulfide-bridged scorpion beta neurotoxin CssII: heterologous expression and proper folding in vitro. Biochim Biophys Acta 1770:1161–1168
Galan JA, Sanchez EE, Rodriguez-Acosta A, Soto JG, Bashir S, McLane MA, Paquette-Straub C, Perez JC (2008) Inhibition of lung tumor colonization and cell migration with the disintegrin crotatroxin 2 isolated from the venom of Crotalus atrox. Toxicon 51:1186–1196. https://doi.org/10.1016/j.toxicon.2008.02.004
Greenfield NJ (2006) Using circular dichroism collected as a function of temperature to determine the thermodynamics of protein unfolding and binding interactions. Nat Protoc 1:2527–2535. https://doi.org/10.1038/nprot.2006.204
Hammouda MB, Montenegro MF, Sanchez-Del-Campo L, Zakraoui O, Aloui Z, Riahi-Chebbi I, Karoui H, Rodriguez-Lopez JN, Essafi-Benkhadir K (2016) Lebein, a snake venom disintegrin, induces apoptosis in human melanoma cells. Toxins 8. https://doi.org/10.3390/toxins8070206
Kauskot A, Cominetti MR, Ramos OH, Bechyne I, Renard JM, Hoylaerts MF, Crepin M, Legrand C, Selistre-de-Araujo HS, Bonnefoy A (2008) Hemostatic effects of recombinant DisBa-01, a disintegrin from Bothrops alternatus. Front Biosci 13:6604–6616. https://doi.org/10.2741/3176
Khrapunov S (2009) Circular dichroism spectroscopy has intrinsic limitations for protein secondary structure analysis. Anal Biochem 389:174–176. https://doi.org/10.1016/j.ab.2009.03.036
Lima-Dos-Santos I, Della-Casa MS, Portes-Junior JA, Calabria PA, Magalhaes GS, Moura-da-Silva AM (2015) Characterization of Neuwiedin, a new disintegrin from Bothrops neuwiedi venom gland with distinct cysteine pattern. Toxicon 104:57–64. https://doi.org/10.1016/j.toxicon.2015.08.006
Lucena S, Sanchez EE, Perez JC (2011) Anti-metastatic activity of the recombinant disintegrin, r-mojastin 1, from the Mohave rattlesnake. Toxicon 57:794–802. https://doi.org/10.1016/j.toxicon.2011.02.014
Lucena SE, Jia Y, Soto JG, Parral J, Cantu E, Brannon J, Lardner K, Ramos CJ, Seoane AI, Sanchez EE (2012) Anti-invasive and anti-adhesive activities of a recombinant disintegrin, r-viridistatin 2, derived from the Prairie rattlesnake (Crotalus viridis viridis). Toxicon 60:31–39. https://doi.org/10.1016/j.toxicon.2012.03.011
Micsonai A, Wien F, Bulyaki E, Kun J, Moussong E, Lee YH, Goto Y, Refregiers M, Kardos J (2018) BeStSel: a web server for accurate protein secondary structure prediction and fold recognition from the circular dichroism spectra. Nucleic Acids Res 46:W315–W322. https://doi.org/10.1093/nar/gky497
Minea R, Helchowski C, Rubino B, Brodmann K, Swenson S, Markland F Jr (2012) Development of a chimeric recombinant disintegrin as a cost-effective anti-cancer agent with promising translational potential. Toxicon 59:472–486. https://doi.org/10.1016/j.toxicon.2011.02.020
Montenegro CF, Casali BC, Lino RLB, Pachane BC, Santos PK, Horwitz AR, Selistre-de-Araujo HS, Lamers ML (2017) Inhibition of alphavbeta3 integrin induces loss of cell directionality of oral squamous carcinoma cells (OSCC). PLoS ONE 12:e0176226. https://doi.org/10.1371/journal.pone.0176226
Mosmann T (1983) Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods 65:55–63. https://doi.org/10.1016/0022-1759(83)90303-4
Pecheur I, Peyruchaud O, Serre CM, Guglielmi J, Voland C, Bourre F, Margue C, Cohen-Solal M, Buffet A, Kieffer N, Clezardin P (2002) Integrin alpha(v)beta3 expression confers on tumor cells a greater propensity to metastasize to bone. FASEB J 16:1266–1268. https://doi.org/10.1096/fj.01-0911fje
Phillips DR, Scarborough RM (1997) Clinical pharmacology of eptifibatide. Am J Cardiol 80. https://doi.org/10.1016/s0002-9149(97)00572-9. 11B-20B
Pinto A, Angulo Y, Jimenez R, Lomonte B (2003) Isolation of bothrasperin, a disintegrin with potent platelet aggregation inhibitory activity, from the venom of the snake Bothrops asper. Rev Biol Trop 51:253–259
Ren A, Wang S, Cai W, Yang G, Zhu Y, Wu X, Zhang Y (2006) Agkistin-s, a disintegrin domain, inhibits angiogenesis and induces BAECs apoptosis. J Cell Biochem 99:1517–1523. https://doi.org/10.1002/jcb.20859
Rivas Mercado E, Neri Castro E, Benard Valle M, Rucavado-Romero A, Olvera Rodriguez A, Zamudio Zuniga F, Alagon Cano A, Ocanas G, L (2020) Disintegrins extracted from totonacan rattlesnake (Crotalus totonacus) venom and their anti-adhesive and anti-migration effects on MDA-MB-231 and HMEC-1 cells. Toxicol In Vitro 65:104809. https://doi.org/10.1016/j.tiv.2020.104809
Rolli M, Fransvea E, Pilch J, Saven A, Felding-Habermann B (2003) Activated integrin alphavbeta3 cooperates with metalloproteinase MMP-9 in regulating migration of metastatic breast cancer cells. Proc Natl Acad Sci U S A 100:9482–9487. https://doi.org/10.1073/pnas.1633689100
Sanchez EE, Rodriguez-Acosta A, Palomar R, Lucena SE, Bashir S, Soto JG, Perez JC (2009) Colombistatin: a disintegrin isolated from the venom of the south american snake (Bothrops colombiensis) that effectively inhibits platelet aggregation and SK-Mel-28 cell adhesion. Arch Toxicol 83:271–279. https://doi.org/10.1007/s00204-008-0358-y
Saviola AJ, Burns PD, Mukherjee AK, Mackessy SP (2016) The disintegrin tzabcanin inhibits adhesion and migration in melanoma and lung cancer cells. Int J Biol Macromol 88:457–464. https://doi.org/10.1016/j.ijbiomac.2016.04.008
Sloan EK, Pouliot N, Stanley KL, Chia J, Moseley JM, Hards DK, Anderson RL (2006) Tumor-specific expression of alphavbeta3 integrin promotes spontaneous metastasis of breast cancer to bone. Breast Cancer Res 8:R20. https://doi.org/10.1186/bcr1398
Taherian A, Li X, Liu Y, Haas TA (2011) Differences in integrin expression and signaling within human breast cancer cells. BMC Cancer 11:293. https://doi.org/10.1186/1471-2407-11-293
Vasconcelos AA, Estrada JC, David V, Wermelinger LS, Almeida FCL, Zingali RB (2021) Structure-function relationship of the disintegrin family: sequence signature and Integrin Interaction. Front Mol Biosci 8:783301. https://doi.org/10.3389/fmolb.2021.783301
Walsh EM, Marcinkiewicz C (2011) Non-RGD-containing snake venom disintegrins, functional and structural relations. Toxicon 58:355–362. https://doi.org/10.1016/j.toxicon.2011.07.004
Wierzbicka-Patynowski I, Niewiarowski S, Marcinkiewicz C, Calvete JJ, Marcinkiewicz MM, McLane MA (1999) Structural requirements of echistatin for the recognition of alpha(v)beta(3) and alpha(5)beta(1) integrins. J Biol Chem 274:37809–37814. https://doi.org/10.1074/jbc.274.53.37809
Zakraoui O, Marcinkiewicz C, Aloui Z, Othman H, Grepin R, Haoues M, Essafi M, Srairi-Abid N, Gasmi A, Karoui H, Pages G, Essafi-Benkhadir K (2017) Lebein, a snake venom disintegrin, suppresses human colon cancer cells proliferation and tumor-induced angiogenesis through cell cycle arrest, apoptosis induction and inhibition of VEGF expression. Mol Carcinog 56:18–35. https://doi.org/10.1002/mc.22470
Acknowledgements
The mass spectrometry determination conducted by Dr. Fernando Zamudio is greatly acknowledged. Also, we acknowledge Dr. Adolfo de Roodt, BSc., DVM Vanessa Costa de Oliveira, and DVM Pablo Regner for their help during the extraction of the venom gland and B. ammodytoides venom supply. We additionally acknowledge Dr. Paul Gaytán, M.C. Eugenio López-Bustos and Q.I. Santiago Becerra from Unidad de Síntesis y Secuenciación de ADN at Instituto de Biotecnología. This research was funded by DGAPA-UNAM (grant number IT200321) and by CONACyT (grant number PRONAII 303045).
Author information
Authors and Affiliations
Contributions
Conceptualization, HC and GC; methodology, HC, LLCG, ERV and LGO; formal analysis, HC, ERV, LGO and GC; investigation, HC and GC; resources, GC; writing—original draft preparation, HC, LLCG and GC; writing—review and editing, HC, LLCG, ERV, LGO and GC; funding acquisition, GC.
Corresponding author
Ethics declarations
Compliance with Ethical Standards
Animal experimental procedures were in accordance with international recommendations and the guidelines of the Good Experimental Practices, under the supervision of the Ethical and Animal Welfare Committee of the Institute of Biotechnology (Ethical approval IT/IBt/Project #411). In brief, proper animal handling, to minimize distress and discomfort, was always conducted towards maximizing the animal welfare during experimentation according also to the Mexican legislation for the use of laboratory animals (Norma Oficial Mexicana, 1999, NOM-062-ZOO-1999).
Competing interest
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Clement, H., Corrales-García, L.L., Rivas-Mercado, E. et al. cDNA Cloning, Heterologous Expression, Cytotoxicity, and Inhibitory Effects of a Disintegrin from Bothrops ammodytoides Venom. Int J Pept Res Ther 29, 54 (2023). https://doi.org/10.1007/s10989-023-10530-5
Accepted:
Published:
DOI: https://doi.org/10.1007/s10989-023-10530-5