Abstract
Fungal infections are becoming a serious problem due to their high morbidity and mortality combined with the rise in drug resistance and dearth of new antimycotic drugs. The scorpion venom-derived peptide BmKn2, and its synthetic analogue Kn2–7, were previously observed to have antibacterial activity. These peptides and their d-amino acid analogues (dBmKn2 and dKn2–7) were tested for antifungal activity against drug resistant and clinical isolates of Candida albicans. In planktonic susceptibility studies, dKn2–7 had greater activity than the other three peptides against 6 out of 7 fungal strains, with no apparent correlation between drug resistance and minimum fungicidal concentrations (MFCs). Time kill experiments demonstrated that the fungicidal activity of dKn2–7 began within the first hour and killing rates were dose dependent at ≥ 1 × MFC. Against biofilms, the d-analogues were the most effective, while the l-analogues had low efficacy in most strains even at 10 times the planktonic MFC. Stability testing suggests that this increased efficacy of the d-analogues may be due to increased resistance to protease degradation compared to the l-analogues. Peptides were also assessed for mammalian cell toxicity. BmKn2 and dBmKn2 induced significant hemolysis at levels similar to their MFCs, whereas Kn2–7 and dKn2–7 caused hemolysis at 4–16 times their MFCs. The 50% inhibitory concentration (IC50) for dKn2–7 against murine fibroblasts was greater than or equal to the planktonic MFCs and biofilm IC50s for dKn2–7 in all C. albicans strains tested. These results support the potential for dKn2–7 to be further investigated as a novel antifungal therapeutic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasive fungal wound infections have emerged as a notable cause of severe morbidity and high mortality rates among injured U.S. military personnel (Murray et al. 2011; Warkentien et al. 2012). Among civilian populations, candidemia is the fourth most common nosocomial bloodstream infection, with recent surgery and long-term catheterization as common risk factors for developing systemic candidiasis (Kullberg and Arendrup 2015; Leroy et al. 2009). Systemic fungal infections are associated with mortality rates as high as 40% and cost the U.S. healthcare system an estimated $2.6 billion in 1998 (Kullberg and Arendrup 2015; Wilson et al. 2002). Use of existing antifungal therapies is limited because they attack targets similar to structures in mammalian tissues and, therefore, commonly induce severe toxicity (Lewis 2011). This situation is further complicated by the limited number of FDA-approved antifungal agents, emergence of drug resistance, and extreme recalcitrance of fungal biofilms, which can lead to recurrent infections (Bachmann et al. 2002; Chandra et al. 2001; Desai et al. 2014; Ellis 2002). Therefore, there is an urgent and unmet need to develop novel antifungal therapies that are effective against drug resistant fungi, can kill fungal biofilms, and are better tolerated by patients.
Antimicrobial peptides (AMPs) are naturally produced by a wide range of organisms as an innate defense mechanism and have become a recent source of new antimicrobial drugs (Gordon et al. 2005; Ramesh et al. 2016). AMPs offer a number of important advantages over traditional small molecule treatments, such as broad spectrum activity against bacteria, fungi, and viruses; rapid onset of microbial killing; lower propensity for inducing microbial resistance; and anti-inflammatory properties (Brogden 2005; Gordon et al. 2005; Zasloff 2002). Most of the published reports on AMPs involve investigation of antibacterial effects, but antifungal peptides with varying structural characteristics, modes of action, and spectra of activity against microbes have been identified from multiple sources including insects, aquatic invertebrates, and mammals (van der Weerden et al. 2013). Indeed, the specific peptide sequences and secondary structures required for optimal antifungal activity and the associated mechanisms of action have not been fully elucidated (van der Weerden et al. 2013). Despite the focus on bacterial activity of AMPs in most studies, a few antifungal peptides have reached clinical trials, although none are currently approved by the FDA for human use (Mahlapuu et al. 2016).
BmKn2 is an AMP isolated from Mesobuthus martensii scorpion venom that exhibits antibacterial activity, but is only weakly active against Gram-negative bacteria (Cao et al. 2012). Three uncharged amino acid residues in BmKn2 were substituted for positively-charged residues to create Kn2–7. With this higher charge density, Kn2–7 is moderately more effective at killing Gram-positive bacteria and significantly more effective at killing Gram-negative bacteria than BmKn2. The differences in antibacterial efficacy of BmKn2 and Kn2–7 were correlated to association of the peptides with different bacterial surface components in binding affinity assays. Furthermore, transmission electron microscopy analysis indicated that binding of BmKn2 and Kn2–7 to bacterial surface molecules led to rapid (< 2 min) cell wall and membrane disruption. In an in vivo mouse model, Kn2–7 was able to resolve topical Staphylococcus aureus infections with minimal host toxicity, indicating that it has an acceptable safety profile at therapeutic levels. In addition to its antibacterial effects, Kn2–7 has also been found to have anti-HIV activity (Chen et al. 2012).
Here, we evaluated the possible in vitro antifungal properties of BmKn2 and Kn2–7 for the purpose of identifying broad spectrum small molecules with a low risk of inducing host toxicity as candidates for drug development. One potential limitation of peptide therapeutics is degradation by endogenous proteases (Gordon et al. 2005). Therefore, d-amino acid analogs of BmKn2 and Kn2–7 (dBmKn2 and dKn2–7) were also tested for efficacy against planktonic and biofilm cultures of clinical and drug resistant laboratory strains of C. albicans and toxicity against host cells to determine if such analogs demonstrated increased antifungal activity without adversely affecting host toxicity compared to the l-amino acid forms of the peptides (Zhao et al. 2016).
Materials and Methods
Candida albicans Strains and Culture Conditions
C. albicans ATCC® MYA-2876 as a wild-type, drug sensitive strain, and the ATCC® drug resistant strains 64124, 76485, and 200955 were purchased from American Type Culture Collection (Manassas, VA, USA). Clinical C. albicans strains CA 494, CA 526, and CA 2519 were obtained from a cell repository at Brooke Army Medical Center, Joint Base San Antonio-Fort Sam Houston, Texas. Information on drug susceptibilities of these strains is given in Supplementary Table S1 in Online Resource 1. Prior to experiments, colonies isolated from Sabouraud dextrose agar (SDA) plates were grown overnight in yeast peptone dextrose medium. Cells were collected via centrifugation and washed twice with phosphate buffered saline (PBS) prior to suspension in RPMI 1640 medium (pH 7.0) buffered with 165 mM 3-(N-morpholino) propanesulfonic acid at the specified density for each assay.
Peptides
BmKn2 (FIGAIARLLSKIF), dBmKn2, Kn2–7 (FIKRIARLLRKIF), and dKn2–7 were synthesized using solid-phase methodology and purified to greater than 95% purity using reverse-phase HPLC by GenScript (Piscataway, NJ, USA). AMP 1018 (VRLIVAVRIWRR) was synthesized and purified to 98% purity by Genemed Synthesis (San Antonio, TX, USA). Melittin was purchased from Sigma-Aldrich (Saint Louis, MO, USA). Stock solutions of melittin, Kn2–7, and dKn2–7 were prepared in water at 100 mg/mL. Stock solutions of BmKn2 and dBmKn2 were prepared at 100 mg/mL in dimethyl sulfoxide.
Planktonic Growth Inhibition
Minimum inhibitory concentrations (MICs) were determined for the seven C. albicans strains as previously described (Clinical and Laboratory Standards Institute (CLSI) 2008) with slight modifications. In brief, 100 µL aliquots of C. albicans at 2.0 × 103 cells/mL were added to a 96-well microtiter plate containing AMPs in RPMI 1640 medium to achieve final drug concentrations of 0–100 µg/mL. Wells containing media alone were also included as sterility controls. Plates were incubated at 37 °C for 24 h then a BioTek Synergy HT plate reader (Winooski, VT, USA) was used to measure the optical density of each well at 600 nm. MIC was reported as the lowest concentration of peptide without detectable growth compared to the media sterility control; detectible growth was determined by a significant change (P ≤ 0.05 in student’s t-test) in OD600 from the sterility control. The plates were incubated for an additional 24 h at 37 °C before a 2.5- or 5-µL aliquot was taken from the well and plated on SDA. Agar plates were incubated for 24 h, and minimum fungicidal concentrations (MFCs) were reported as the lowest concentration of a peptide that resulted in no growth from any of the wells. Three independent experiments were performed with triplicate samples for each condition.
Time-Kill Assay
The fungicidal activity of dKn2–7 (0, 6.25, 12.5, 25 and 50 µg/mL) in RPMI 1640 was measured using the C. albicans MYA-2876 strain (1.0 × 106 cells/mL) treated in 15-mL tissue culture tubes. Treated cells were incubated at 37 °C with shaking (250 rpm). At 0, 1, 2, 3, 4, 20, and 24 h, 20 µL aliquots were removed, and 10-, 100-, and 1000-fold dilutions were prepared. Aliquots of these dilutions were added to SDA plates. Plates were incubated for 24 h at 37 °C, after which CFUs were enumerated. CFU counts were normalized for each sample against the 0 h count. Data are from three independent experiments with three replicate CFU counts per condition in each experiment.
Biofilm Susceptibility
Susceptibility of C. albicans biofilms to BmKn2, dBmKn2, Kn2–7, and dKn2–7 was determined using a metabolic assay as previously described (Pierce et al. 2008). In brief, biofilms were formed by adding 100 µL aliquots of 1.0 × 106 cells/mL in RPMI 1640 into 96-well flat bottomed plates that were then incubated for 24 h at 37 °C. Biofilms were rinsed twice with 100 µL of PBS to remove any planktonic cells. In a separate plate, AMP dilutions (0–1000 µg/mL) were prepared in RPMI 1640, and 100 µL was added to each biofilm. The biofilms were incubated for 24 h at 37 °C and rinsed twice with 100 µL of PBS. Freshly prepared XTT reagent (100 µL; Sigma Aldrich, St Louis, MO, USA) was added to each well, and samples were incubated for 2 h at 37 °C. XTT-only wells (no biofilms) were used as the positive control for complete metabolic inhibition. After incubation, 80 µL of the supernatant was transferred to a 96-well transparent flat-bottomed plate and absorbance was measured at 490 nm using a BioTek Synergy HT plate reader. Data presented is an average of three independent experiments performed with triplicate samples for each condition. Susceptibility values were normalized to the metabolic activity of biofilms grown in peptide-free media for 24 h. Due to weak biofilm formation, strains ATCC 76485 and ATCC 200955 were not used in this assay.
Confocal Microscopy Analysis
C. albicans MYA-2876 biofilms were grown on glass-bottomed 96-well plates pre-coated for 24 h with human fibrinogen using methods described in the previous section. Biofilms were cultured for 24 h, rinsed with PBS, incubated in the presence of dKn2–7 at 0–500 µg/mL in RPMI 1640 for 24 h, and then rinsed twice with PBS. Treated biofilms were simultaneously stained with propidium iodide (PI, 40 µM) and SYTO® 9 (6.7 µM; Life Technologies, Carlsbad, CA, USA) for 30 min in PBS with 2% glucose (wt/vol). Staining solution was aspirated, and biofilms were rinsed twice with 100 µL of PBS. The stained biofilms were immediately imaged with a CFI Plan Fluor 40 × magnification objective (0.75 NA) on a Nikon Eclipse C1 confocal microscope (Melville, NY, USA), and images were processed using NIS-Elements (v. 4.20.02).
Stability of Peptides Against Protease Activity
The stability of Kn2–7 and dKn2–7 peptides (50 µg/mL) in trypsin (10 µg/mL; Sigma Aldrich) was tested in PBS. Prior to the start of the reaction, all solutions were prepared on ice. Solutions were then placed in a 37 °C water bath for the entirety of the experiment after the 0 h samples were collected. At 0, 4, and 24 h, 100 µL from each sample was aspirated and thoroughly mixed with 200 µL of 96% ethanol. The mixture was incubated on ice for 15 min to allow trypsin to precipitate before being centrifuged for 5 min at 9300 relative centrifugal force to pellet precipitated proteins. Supernatant (200 µL) was carefully aspirated and stored at − 20 °C until chromatographic analysis.
Liquid chromatography mass spectrometric (LC–MS) analysis to quantify uncleaved peptide in the samples was performed on an LC-20AD HPLC system (Shimadzu Corporation, Kyoto, Japan) and API 4000 mass spectrometer with an electrospray ionization interface and triple quadrupole mass analyzer (AB Sciex, Framingham, MA, USA). Samples were diluted to 8 times the original volume with 5% acetonitrile (ACN)/0.1% formic acid (FA) and spiked with 1 µg/mL AMP 1018 as an internal standard. Ten µL aliquots were injected for LC–MS analysis. A gradient elution was carried out on an Agilent ZORBAX Eclipse XDB 80 Å C18 column (2.1 × 50 mm, 5 µm) at 40 °C with mobile phase A (10% ACN/0.1% FA in water) and mobile phase B (100% ACN/0.1% FA) with a total flow rate of 0.4 mL/min. Peptides were bound to the column for 0.5 min with 100% mobile phase A and then eluted with a linear gradient from 0 to 90% mobile phase B for 2.5 min. Under these chromatographic conditions, the retention time for Kn2–7/dKn2–7 and AMP 1018 were 2.46 and 2.42 min, respectively. Multiple reaction monitoring measurements were performed in the positive electrospray tandem MS mode using two transitions for each analyte, (m/z 558.71/120.0 and m/z 558.71/103.0 for Kn2–7/dKn2–7, m/z 385.18/239.1 and m/z 385.18/115.1 for AMP 1018). A least-squares linear regression analysis of observed peptide-to-internal standard peak-area ratios was determined for each sample, and 4 and 24 h samples were normalized to the 0 h samples to determine peptide stability over time. Data in the graph represent the average of three individual experiments.
Hemolysis Assay
Hemolytic activity of BmKn2, dBmKn2, Kn2–7, dKn2–7, and melittin was assessed using red blood cells (RBCs) prepared from human whole blood (BioreclamationIVT, Westbury, NY, USA) as previously described (Evans et al. 2013). Peptides (6.25–400 µg/mL) were incubated with RBCs (2% vol/vol in PBS) for 1 h at 37 °C in a 96-well microtiter plate and subsequently centrifuged at 500×g for 5 min. The supernatant (100 µL) was transferred to a 96-well, transparent flat-bottomed plate. Hemoglobin release was quantified based on absorbance readings measured at 490 nm using a BioTek Synergy HT plate reader. Negative controls consisted of RBCs in PBS without peptide. Positive controls consisted of RBCs treated with 1% Triton X-100 (Sigma Aldrich). The assay was performed in triplicate for three independent experiments using RBCs collected from different donors. Results from each assay were averaged, and % hemolysis was determined using the following equation:
Fibroblast Toxicity
Cytotoxic effects of dKn2–7 against fibroblasts was evaluated using a metabolic assay. In brief, mouse fibroblast 3T3/NIH cells were plated in a 96-well microtiter plate at 1.0 × 104 cells/well in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% calf serum and allowed to attach overnight. Cells were treated with dKn2–7 (0–125 µg/mL) in DMEM with 10% calf serum for 24 h at 37 °C. After treatment, AMP solutions were removed, 100 µL of a 10:1 DMEM with serum:WST-1 cell proliferation reagent (Sigma Aldrich) solution was added to each well, and samples were incubated for 2 h at 37 °C. The cellular metabolic activity was quantified based on absorbance readings at 450 nm (detection wavelength) minus absorbance at 630 nm (reference wavelength) using a BioTek Synergy HT plate reader. Cells treated with 0.1% Triton X-100 in DMEM were used as a positive control. Toxicity assays were performed in triplicate for three independent experiments. Normalized metabolic activity was determined using the following equation:
Statistics
Data are presented as the mean ± standard deviation (SD) unless otherwise stated. Statistical analysis was performed using GraphPad Prism v6.07 (GraphPad Software, Inc., La Jolla, CA, USA). One-way analysis of variance (ANOVA) and Dunnett’s multiple comparisons test or two-way ANOVA and Tukey’s multiple comparison tests were used to compare groups, and P values ≤ 0.05 were considered statistically significant. For biofilm susceptibility and fibroblast toxicity, 50% inhibitory concentrations (IC50s) and their corresponding 95% confidence intervals were calculated using GraphPad Prism’s nonlinear fit function for inhibitor vs. response.
Results
AMPs Exhibit Antifungal Activity Against Clinical and Drug Resistant C. albicans Strains In Vitro
As the most common cause of fungal infections in developed countries, C. albicans was selected for these antifungal assays (Chandra et al. 2001). MICs of the four AMPs were determined using the standard broth microdilution technique and a 24-h treatment time. MFC determination was made using 48 h treated samples. The most potent antifungal peptide tested was dKn2–7 with MICs ranging from 12.5 to 100 µg/mL across all seven C. albicans strains tested (Table 1). Kn2–7 had MIC values double those of dKn2–7 for most strains (25–100 µg/mL). BmKn2 and dBmKn2 were the least effective growth inhibitors and had similar MICs to each other in all strains. Peptide fungicidal efficacy, as determined by MFCs, followed the same trend. dKn2–7 was the most potent peptide against all strains with MFCs of 25–100 µg/mL. The other three peptides had similar MFCs to each other in most strains, with BmKn2 being the least effective overall. A standard, drug susceptible laboratory strain, MYA-2876, had identical MIC and MFC profiles to that of the multi-drug resistant strain ATCC 64124. All peptides exhibited the lowest growth inhibitory and fungicidal activity against one of the clinical strains, CA 494. AMP activity did not appear to be affected by strain resistance to commonly used antifungal drugs (strain susceptibilities to fluconazole, amphotericin B, and caspofungin can be found in Supplementary Table 1 in Online Resource 1).
dKn2–7 Exhibits Dose-Dependent Killing Rates at Concentrations Above the MFC
A major advantage of AMPs over traditional antifungals is their ability to rapidly kill planktonic fungi before they can form biofilms (Cao et al. 2012; Chandra et al. 2001). As the most potent antifungal of the four peptides tested, dKn2–7 was used to determine the rate of killing of planktonic C. albicans MYA-2876 over 24 h. dKn2–7 was tested at concentrations of ½ × MIC (6.25 µg/mL), 1 × MIC (12.5 µg/mL), 1 × MFC (25 µg/mL), and 2 × MFC (50 µg/mL). No substantial differences in CFU/mL were observed between the controls and ½ × MIC samples at any time point (Fig. 1). The 1 × MIC samples initially showed reduced CFUs compared to the controls (60% reduction by 4 h), but no significant differences from the controls were observed at later time points, indicating a regrowth of remaining cells after the initial period of cell killing. The discrepancy between this finding and the 24 h results from the MIC studies (Table 1) may be due to the fact that the time-kill assay had a 103 higher fungal cell concentration at 0 h compared to the MIC studies. Both 1 × MFC and 2 × MFC reduced CFUs gradually over time, with > 99.9% killing by 20 and 4 h, respectively. These results indicate a dose-dependent response of planktonic C. albicans to dKn2–7. They also demonstrate the need for at least 1 × MFC levels of dKn2–7 to maintain growth inhibitory effects over time at high initial cell densities.
Time-kill kinetics of dKn2–7 against planktonic C. albicans MYA-2876 over 24 h. Data were normalized to the associated CFU/mL at time 0 and are from three independent experiments with 3 CFU counts per experimental condition; results are presented as the mean ± SD with white symbols indicating averaged normalized values from the individual triplicate experiments
C. albicans Biofilms are More Susceptible to d-enantiomers than l-enantiomers of the Peptides
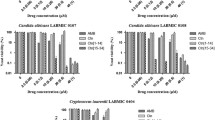
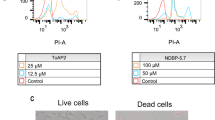
Five of the seven C. albicans strains tested for planktonic susceptibility produced biofilms robust enough for susceptibility testing. Biofilms of these five strains were exposed to all four peptides at concentrations up to 1000 µg/mL for 24 h then tested for metabolic activity using the XTT assay. dKn2–7 exhibited the greatest anti-biofilm potency with IC50s ranging from 25 to 80 µg/mL (Table 2) and complete inhibition of metabolic activity at concentrations of ≥ 125 µg/mL in all strains (Fig. 2). dBmKn2 was the second most potent AMP, reducing metabolic activity to that of the positive control at ≥ 250–500 µg/mL in all strains. l-amino acid analogues were less potent than the d-amino acid forms of the peptides in all five strains. Kn2–7 eradicated biofilms in only two of the five strains and BmKn2 did not eradicate biofilms in any of the strains at the highest concentration tested. Interestingly, CA 494 was the most susceptible clinical strain in these biofilm studies, despite being the most resistant strain in planktonic form. Overall, the clinical strains produced more resistant biofilms than the two laboratory strains tested.
Efficacy of BmKn2 (red), dBmKn2 (orange), Kn2–7 (green), and dKn2–7 (blue) peptides against pre-formed C. albicans biofilms. Biofilm viability after 24-h AMP treatment was measured using the XTT assay and normalized to metabolic activity of untreated biofilms; each data point represents the average of three independent experiments performed with triplicate samples for each experimental condition
To confirm that results obtained from the metabolic activity assay were indicative of cell death rather than cell quiescence, MYA-2876 biofilms were stained and imaged via confocal microscopy after 24-h dKn2–7 treatment. SYTO® 9 stain was used to visualize all cells, and PI was used to visualize dead cells. Overlaid images of the confocal micrographs (Fig. 3) support the XTT assay results. Control biofilms with no peptide were almost entirely green (indicative of live cells). Biofilms exposed to 62.5 µg/mL of dKn2–7 showed > 50% cell death (indicated by the orange cells in the overlay picture) with substantial numbers of live cells still present. This correlates well with the 84% decrease in metabolic activity detected for this peptide concentration in the XTT assay (Fig. 2). The overlay image from biofilms exposed to 125 µg/mL were completely orange, correlating closely with the negligible metabolic activity levels detected in biofilms treated with dKn2–7 at that concentration. Taken together, these findings confirmed that AMPs were fungicidal and caused cell death in a dose-dependent manner.
Confocal microscopy images of MYA-2876 biofilms treated with dKn2–7 corroborate the XTT results (Fig. 2). C. albicans MYA-2876 biofilms were treated with dKn2–7 for 24 h and then stained with SYTO® 9 (labels all cells green) and propidium iodide (labels cells with permeable membranes red); the ratio of green to red or orange cells in the overlay images is indicative of the level of biofilm viability
dKn2–7 is Less Prone to Protease Degradation Than Kn2–7
To test whether the increased anti-biofilm efficacy of the d-peptides might be due to a reduced susceptibility to degradation by proteases compared to the l-peptide enantiomers, Kn2–7 and dKn2–7 concentrations were determined over 24 h in the presence of trypsin, which was expected to cleave after any of the 5 lysine or arginine residues in those peptide sequences. Significant degradation of Kn2–7 (21% remaining after 4 h and 16% remaining after 24 h) was observed. In contrast, no significant drop in dKn2–7 concentrations was observed after 24 h (Fig. 4). These findings demonstrate that d-peptides were less prone to protease degradation, potentially mediating their increased efficacy against C. albicans biofilms.
Stability of Kn2–7 and dKn2–7 against protease activity. Kn2–7 and dKn2–7 at 50 µg/mL were incubated in PBS alone or 10 µg/mL of trypsin in PBS. Peptide concentration at 0, 4, and 24 h was determined by LC-MS and normalized to the concentration detected at 0 h (t0). *Indicates a statistically significant difference with P ≤ 0.05 between the two samples indicated by brackets as determined by two-way ANOVA with Tukey’s multiple comparisons test
Kn2–7 and dKn2–7 Do Not Induce Significant Hemolysis at Fungicidal Levels
AMPs are often observed to induce significant hemolysis, which can lead to dose limitations in clinical use (Matsuzaki 2009). The hemolytic activities of Kn2–7, dKn2–7, BmKn2, and dBmKn2 were assessed against human RBCs and compared to that of melittin, an AMP with known hemolytic toxicity (Mojsoska and Jenssen 2015). Melittin exhibited significant hemolysis at the lowest tested dose (6.25 µg/mL) and caused complete hemolysis at 25 µg/mL (Fig. 5). BmKn2 and dBmKn2 first caused significant hemolysis at 100 µg/mL and complete hemolysis at 200 µg/mL. This minimum concentration for hemolytic effect (100 µg/mL) is within the range of MICs and MFCs for some of the fungal strains treated with those two peptides. Kn2–7 and dKn2–7 only exhibited statistically significant hemolysis at 400 µg/mL, and hemolysis was < 40% at this highest tested concentration. This minimum hemolytic concentration is 4–8 times higher than all but one of the MFCs for Kn2–7 and 4–16 times higher than all of the MFCs for dKn2–7. This hemolytic concentration is also higher than the biofilm eradication concentrations for dKn2–7 (125 µg/mL). No significant differences in hemolysis were observed between the l- and d-enantiomers of each peptide.
Hemolytic activity of BmKn2, dBmKn2, Kn2–7, and dKn2–7. Human RBCs were exposed to AMPs for 1 h, and hemolysis was quantified using absorbance readings at 490 nm to measure hemoglobin in the supernatant. Data were normalized to hemolysis in positive controls that consisted of RBCs treated with 1% Triton X-100. Melittin was included as an AMP with known hemolytic toxicity. *Indicates a statistically significant difference with P ≤ 0.05 between the sample and the 0 µg/mL control (brackets indicate multiple samples were different from the 0 µg/mL controls)
Fibroblast Toxicity
Fibroblasts are an important cell type in the wound healing process and, therefore, excessive AMP toxicity against fibroblasts could impede healing in infected wounds. As dKn2–7 was the only peptide to exhibit potent antifungal efficacy against both planktonic and biofilm forms of C. albicans and low hemolytic activity, it was selected for further toxicity testing against NIH/3T3 fibroblasts. Metabolic activity of the fibroblasts after 24-h exposure to dKn2–7 was determined using the WST-1 assay. No significant toxicity was observed at or below 70 µg/mL, and complete cell death was observed at 100 µg/mL and higher concentrations (Fig. 6). The IC50 was 79 µg/mL (95% confidence interval of 76–81 µg/mL), which is at or above dKn2–7 biofilm IC50s in the five C. albicans strains tested.
Toxicity of dKn2–7 on fibroblast cells. Mouse fibroblasts (3T3/NIH cell line) were exposed to dKn2–7 for 24 h. Metabolic activity was measured using the WST-1 assay and normalized to the metabolic activity of untreated cells; nonlinear regression was used to determine a best fit curve for dose response
Discussion
The discovery of penicillin and subsequent development of commercial antibiotics against infectious diseases is widely hailed as one of the most significant health advancements in human history. However, the widespread use of commercial antibiotics has led to the development of microbial resistance and a recent increase in morbidity and mortality from infections. As microbes develop resistance against many of these classes of drugs, it has become harder to find and develop new drug classes that will target different microbial functions without causing excessive host toxicity (Bassetti et al. 2013). This is especially true for pathogenic fungi, which are more structurally and metabolically similar than bacteria to mammalian cells (Lewis 2011).
To address this problem, AMPs derived from a wide array of organisms have been investigated as a potential source of new antibiotics (Mahlapuu et al. 2016; Zasloff 2002). Much of the research on AMP activity has centered on membrane disruption as the predominant antimicrobial effect, but recent data has revealed effects of AMPs on intracellular targets, AMPs with multiple mechanisms of action that differ for bacteria and fungi, and target cell specificities that may vary for peptides with similar sequences and conformations (Amaral et al. 2012; van der Weerden et al. 2013; Yeaman and Yount 2003). Hence, one cannot directly extrapolate the previous mechanistic data obtained for BmKn2 and Kn2–7 in bacteria (Cao et al. 2012) to infer specific antifungal mechanisms of action for these peptides. Indeed, to the best of our knowledge, the antifungal mechanisms of BmKn2 and Kn2–7 and their d-enantiomers have not been previously proposed or characterized.
As many chronic fungal infections are mediated by polymicrobial biofilms containing a mixture of bacterial and fungal cells, it is advantageous to treat these infections with broad-spectrum drugs capable of reducing the growth of multiple pathogens (Harriott and Noverr 2011). The fungal MIC values determined for BmKn2 and Kn2–7 are higher than their reported MICs against Gram-positive bacteria (reported MICs of 6.25–12.5 and 3.13–6.25 µg/mL, respectively) (Cao et al. 2012). Kn2–7 appears to be equally effective against C. albicans and Gram-negative bacteria (reported MICs of 6.25–100 µg/mL), whereas BmKn2 appears to be more effective against C. albicans than against Gram-negative bacteria (reported MICs of > 100 µg/mL for 7 of 8 species and strains tested). These results indicate that Kn2–7 may be more effective in polymicrobial fungal infections than mainstay antifungal drugs that are strictly antimycotic.
Efficacy against biofilm infections is especially important as biofilms often increase drug resistance and lead to recurrent infections (Desai et al. 2014). Candida spp. biofilms are highly resistant to amphotericin B and triazoles, even when the planktonic forms of that strain are not drug resistant, leaving echinocandins as the most efficacious drug class to treat biofilm infections (Chandra et al. 2001; Desai et al. 2014; Marcos-Zambrano et al. 2016). The best-case scenario is if a therapeutic can prevent biofilms from forming by killing planktonic fungal cells quickly as drug resistance is lower in the early stages of biofilm development (Chandra et al. 2001). Our time kill assay showed that for dKn2–7, a rapid onset of cell death (within 1–2 h) occurred at ≥ 1 × MIC, and 99.9% cell death was induced within 4 h at 2 × MFC. The rapid onset of antifungal activity of dKn2–7 is similar to amphotericin B killing kinetics for C. albicans that show significant cell death in the initial hours after exposure to drug concentrations above the MFC (Canton et al. 2004). This similarity indicates dKn2–7, like amphotericin B, acts at the cell surface, and provides a distinct advantage over triazoles and echinocandins which may take a day or longer to induce significant killing as their targets are directed at metabolic processes (Canton et al. 2013; Moosa et al. 2003).
In the majority of cases, biofilms form in wounds from pathogens introduced to the patient on surfaces (such as shrapnel and particulate matter in battlefield wounds) or form on indwelling medical devices (such as catheters or dental implants) before a person is placed on an antifungal therapeutic regimen (Chandra et al. 2001; Murray et al. 2011). At this point, the patient must rely on drugs to combat the fully-formed biofilm. Therefore, the AMPs were tested against pre-formed biofilms in the current study. Biofilms of all 5 strains were highly resistant to the l-enantiomer peptides (Fig. 2). The laboratory strains were the most susceptible with a slow decline in biofilm metabolic activity as l-peptide levels surpassed 1 × MFC, but the biofilms still maintained some viability even at 20 × MFC. For the clinical strains, BmKn2 had no observable effect except against CA 494 at the highest concentration tested. CA 494 was the only clinical strain that Kn2–7 was able to eradicate. Conversely, the biofilms were much more susceptible to the d-enantiomer peptides. Decreases in biofilm metabolic activity were seen at levels below the MFC for these peptides, and complete eradication was observed at 5–10 × MFC. For the clinical strains, eradication was observed at 1–5 × MIC. Interestingly, the fungicidal effect of both l- and d-peptides against biofilms appeared to be gradual and highly dose dependent in the two laboratory strains, while the transition from no killing to significant killing was more pronounced in the clinical strains. Despite this, the minimum concentrations of d-peptides needed for complete biofilm eradication were more uniform between strains than the planktonic MFCs.
The discrepancy between the relative efficacies of Kn2–7 and dKn2–7 against planktonic and biofilm forms of C. albicans may be explained by multiple factors including the stability of the peptides against protease degradation, involvement of receptor-mediated interactions of the AMP at the fungal cell surface, and differences in binding putative intracellular targets (de la Fuente-Núñez et al. 2015; Yeaman and Yount 2003) and has important implications. The decreased degradation of dKn2–7 compared to Kn2–7 in the presence of trypsin (Fig. 4) suggests resistance to proteases and, at least in part, explains the increased efficacy of dKn2–7 against C. albicans. Our results are in agreement with previous studies that have observed d-amino acid peptides to be more efficacious against certain strains of bacterial biofilms than their l-enantiomers not only in vitro, but also in vivo (Andrea et al. 2018; de la Fuente-Núñez et al. 2015). This may have been due to the ability of the d-enantiomers to resist pathogen and host proteases, thus increasing the retention time of the peptides in vivo.
Many AMPs studied for antimicrobial therapies cause high local and systemic toxicity (Gordon et al. 2005). It can be difficult to predict which mammalian cell types will be susceptible to a particular AMP, so the route of administration is often a deciding factor in which host cell types to test (Bacalum and Radu 2014). For use of AMPs in infected wounds, we determined that hemolysis and fibroblast toxicity were important to investigate. Our hemolysis results were promising as Kn2–7 and dKn2–7 induced significant hemolysis only at 4–16 × MFC. However, as has been reported for other AMPs (Bacalum and Radu 2014), the minimum concentration of dKn2–7 that induced hemolysis (400 µg/mL) was much higher than levels that caused toxicity in murine fibroblasts. The IC50 for dKn2–7 against fibroblasts was 79 µg/mL, and this value is slightly above or within the range of all C. albicans planktonic MFCs and biofilm IC50s, indicating that therapeutic levels of the peptide may be toxic in vivo. However, a previous study found Kn2–7 IC50s for HeLa and human lymphoblast cell lines to be 38.46 and 43.18 µg/mL, respectively (Chen et al. 2012). Despite these low in vitro IC50s, a topical gel containing 5 mg/mL of Kn2–7 was able to resolve in vivo S. aureus mouse skin infections to result in full healing with no significant toxicity compared to non-peptide treated controls. Discrepancies between in vitro and in vivo toxicities of AMPs are not without precedent (Wu et al. 2014), and thus, additional testing is warranted to more fully characterize the effects of dKn2–7 on host cells.
In conclusion, this study investigated four peptides for antifungal potential against multiple C. albicans strains. Overall, dKn2–7 exhibited the most potent antifungal efficacy against all strains in planktonic and biofilm states compared to the other three peptides. Additionally, dKn2–7 toxic concentrations toward a mammalian cell line were at or above the highest concentrations needed for fungicidal activity. These results, combined with the previously reported efficacy and low toxicity of its l-enantiomer in an in vivo S. aureus-infected wound model, suggest that dKn2–7 could be a viable option for treating infectious fungal wounds. Also, the low levels of hemolysis induced by dKn2–7 at concentrations capable of eradicating planktonic and biofilm C. albicans cultures may be an advantage for use of this peptide against systemic fungal infections or biofilms associated with implanted medical devices such as central venous catheters. However, future studies involving a wider range of fungal isolates and more comprehensive analysis of peptide effects on host tissues are required to better determine dKn2–7’s ability to provide antifungal activity without undue toxicity in vivo.
References
Amaral AC et al (2012) Predicting antimicrobial peptides from eukaryotic genomes: in silico strategies to develop antibiotics. Peptides 37:301–308. https://doi.org/10.1016/j.peptides.2012.07.021
Andrea A, Molchanova N, Jenssen H (2018) Antibiofilm peptides and peptidomimetics with focus on surface immobilization. Biomolecules 8:27. https://doi.org/10.3390/biom8020027
Bacalum M, Radu M (2014) Cationic antimicrobial peptides cytotoxicity on mammalian cells: an analysis using therapeutic index integrative concept. Int J Pept Res Ther 21:47–55. https://doi.org/10.1007/s10989-014-9430-z
Bachmann SP, VandeWalle K, Ramage G, Patterson TF, Wickes BL, Graybill JR, Lopez-Ribot JL (2002) In vitro activity of caspofungin against Candida albicans biofilms. Antimicrob Agents Chemother 46:3591–3596. https://doi.org/10.1128/AAC.46.11.3591-3596.2002
Bassetti M, Merelli M, Temperoni C, Astilean A (2013) New antibiotics for bad bugs: where are we? Ann Clin Microbiol Antimicrob 12:1–15
Brogden KA (2005) Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3:238–250. https://doi.org/10.1038/nrmicro1098
Canton E, Peman J, Gobernado M, Viudes A, Espinel-Ingroff A (2004) Patterns of amphotericin B killing kinetics against seven Candida species. Antimicrob Agents Chemother 48:2477–2482. https://doi.org/10.1128/AAC.48.7.2477-2482.2004
Canton E, Peman J, Hervas D, Espinel-Ingroff A (2013) Examination of the in vitro fungicidal activity of echinocandins against Candida lusitaniae by time-killing methods. J Antimicrob Chemother 68:864–868. https://doi.org/10.1093/jac/dks489
Cao L et al (2012) Antibacterial activity and mechanism of a scorpion venom peptide derivative in vitro and in vivo. PLoS ONE 7:e40135. https://doi.org/10.1371/journal.pone.0040135
Chandra J, Kuhn DM, Mukherjee PK, Hoyer LL, McCormick T, Ghannoum MA (2001) Biofilm formation by the fungal pathogen Candida albicans: development, architecture, and drug resistance. J Bacteriol 183:5385–5394. https://doi.org/10.1128/jb.183.18.5385-5394.2001
Chen Y et al (2012) Anti-HIV-1 activity of a new scorpion venom peptide derivative Kn2–7. PLoS ONE 7:e34947. https://doi.org/10.1371/journal.pone.0034947
Clinical and Laboratory Standards Institute (CLSI) (2008) Reference method for broth dilution antifungal susceptibility testing of yeasts; approved standard—Third edition. Clinical and Laboratory Standards Institute, 950 West Valley Road, Suite 2500, Wayne, Pennsylvania 19087 USA
de la Fuente-Núñez C et al (2015) enantiomeric peptides the eradicate wild-type and multidrug-resistant biofilms and protect against lethal Pseudomonas aeruginosa infections. Chem Biol 22:196–205. https://doi.org/10.1016/j.chembiol.2015.01.002
Desai JV, Mitchell AP, Andes DR (2014) Fungal biofilms, drug resistance, and recurrent infection. Cold Spring Harb Perspect Med 4:a019729. https://doi.org/10.1101/cshperspect.a019729
Ellis D (2002) Amphotericin B: spectrum and resistance. J Antimicrob Chemother 49:7–10. https://doi.org/10.1093/jac/49.suppl_1.7
Evans BC et al (2013) Ex vivo red blood cell hemolysis assay for the evaluation of pH-responsive endosomolytic agents for cytosolic delivery of biomacromolecular drugs. J Vis Exp 73:e50166. https://doi.org/10.3791/50166
Gordon YJ, Romanowski EG, McDermott AM (2005) A review of antimicrobial peptides and their therapeutic potential as anti-infective drugs. Curr Eye Res 30:505–515
Harriott MM, Noverr MC (2011) Importance of Candida–bacterial polymicrobial biofilms in disease. Trends Microbiol 19:557–563. https://doi.org/10.1016/j.tim.2011.07.004
Kullberg BJ, Arendrup MC (2015) Invasive Candidiasis. N Engl J Med 373:1445–1456. https://doi.org/10.1056/NEJMra1315399
Leroy O et al (2009) Epidemiology, management, and risk factors for death of invasive Candida infections in critical care: a multicenter, prospective, observational study in France (2005–2006). Crit Care Med 37:1612–1618. https://doi.org/10.1097/CCM.0b013e31819efac0
Lewis RE (2011) Current concepts in antifungal pharmacology. Mayo Clin Proc 86:805–817. https://doi.org/10.4065/mcp.2011.0247
Mahlapuu M, Hakansson J, Ringstad L, Bjorn C (2016) Antimicrobial peptides: an emerging category of therapeutic agents. Front Cell Infect Microbiol 6:194. https://doi.org/10.3389/fcimb.2016.00194
Marcos-Zambrano LJ, Escribano P, Bouza E, Guinea J (2016) Susceptibility of Candida albicans biofilms to caspofungin and anidulafungin is not affected by metabolic activity or biomass production. Med Mycol 54:155–161. https://doi.org/10.1093/mmy/myv094
Matsuzaki K (2009) Control of cell selectivity of antimicrobial peptides. Biochim Biophys Acta 1788:1687–1692
Mojsoska B, Jenssen H (2015) Peptides and peptidomimetics for antimicrobial drug design. Pharmaceuticals 8:366–415. https://doi.org/10.3390/ph8030366
Moosa MYS, Sobel JD, Elhalis H, Du W, Akins RA (2003) Fungicidal activity of fluconazole against Candida albicans in a synthetic vagina-simulative medium. Antimicrob Agents Chemother 48:161–167. https://doi.org/10.1128/aac.48.1.161-167.2004
Murray CK et al (2011) Infections complicating the care of combat casualties during operations Iraqi freedom and enduring freedom. J Trauma 71:S62–S73. https://doi.org/10.1097/TA.0b013e3182218c99
Pierce CG, Uppuluri P, Tristan AR, Wormley FL Jr, Mowat E, Ramage G, Lopez-Ribot JL (2008) A simple and reproducible 96-well plate-based method for the formation of fungal biofilms and its application to antifungal susceptibility testing. Nat Protoc 3:1494–1500. https://doi.org/10.1038/nport.2008.141
Ramesh S, Govender T, Kruger HG, de la Torre BG, Albericio F (2016) Short antimicrobial peptides (SAMPs) as a class of extraordinary promising therapeutic agents. J Pept Sci 22:438–451. https://doi.org/10.1002/psc.2894
van der Weerden NL, Bleackley MR, Anderson MA (2013) Properties and mechanisms of action of naturally occurring antifungal peptides. Cell Mol Life Sci 70:3545–3570. https://doi.org/10.1007/s00018-013-1260-1
Warkentien T et al (2012) Invasive mold infections following combat-related injuries. Clin Infect Dis 55:1441–1449. https://doi.org/10.1093/cid/cis749
Wilson LS, Reyes CM, Stolpman M, Speckman J, Allen K, Beney J (2002) The direct cost and incidence of systemic fungal infections. Value Health 5:26–34. https://doi.org/10.1046/j.1524-4733.2002.51108.x
Wu X et al (2014) In vitro and in vivo activities of antimicrobial peptides developed using an amino acid-based activity prediction method. Antimicrob Agents Chemother 58:5342–5349. https://doi.org/10.1128/AAC.02823-14
Yeaman MR, Yount NY (2003) Mechanisms of antimicrobial peptide action and resistance. Pharmacol Rev 55:27–55. https://doi.org/10.1124/pr.55.1.2
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415:389–395. https://doi.org/10.1038/415389a
Zhao Y et al (2016) Antimicrobial activity and stability of the d-amino acid substituted derivatives of antimicrobial peptide polybia-MPI. AMB Express 6:122. https://doi.org/10.1186/s13568-016-0295-8
Acknowledgements
The views expressed in this article are those of the authors and do not necessarily reflect the official policy or position of the Department of the Navy, Department of Defense, nor the U.S. Government. Some of the authors are employees of the U.S. Government. This work was prepared as part of our official duties. Title 17 U.S.C. Section 105 provides that ‘Copyright protection under this title is not available for any work of the United States Government.’ Title 17 U.S.C. Section 101 defines a U.S. Government work as a work prepared by a military Service member or employee of the U.S. Government as part of that person’s official duties. CA 494, CA 526, and CA 2519 were obtained from a repository at the Brooke Army Medical Center at JBSA-Fort Sam Houston, TX. The authors thank Ms. Jennifer Talackine for performing the hemolysis assays.
Funding
This work was supported by Naval Medical Research Center's Advanced Medical Development Program using work unit number G1723 and, in part, by an appointment to the Postgraduate Research Participation Program at the Naval Medical Research Unit San Antonio (NAMRU-SA) administered by the Oak Ridge Institute for Science and Education through an interagency agreement between the U.S. Department of Energy and NAMRU-SA.
Author information
Authors and Affiliations
Contributions
Conceptualization: JWG DK NJM, SSS; Methodology: JWG, SSS; Formal analysis and investigation: JWG, SSS; Writing—original draft preparation: JWG, SSS; Writing—review and editing: WC, JWG, DK, NJM, SSS; Funding acquisition: JWG, DK, NJM, SSS; Supervision: NJM.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
The Naval Medical Research Unit San Antonio Institutional Review Board reviewed this study and determined that it does not meet the definition of human subjects research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Snyder, S.S., Gleaton, J.W., Kirui, D. et al. Antifungal Activity of Synthetic Scorpion Venom-Derived Peptide Analogues Against Candida albicans. Int J Pept Res Ther 27, 281–291 (2021). https://doi.org/10.1007/s10989-020-10084-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-020-10084-w