Abstract
Nocardithiocin is a thiopeptide compound produced by the pathogenic actinomycete Nocardia pseudobrasiliensis that displays activity against Gram-positive bacteria including drug-resistant Mycobacterium tuberculosis. However, the clinical use of nocardithiocin is hampered by light sensitivity and low water solubility. To improve these properties, the sixth amino acid of core peptide was substituted for other 19 amino acids by modifying the gene sequence of precursor peptide. Ten amino acids were successfully substituted to produce 1–3 novel compounds, of which six were the expected nocardithiocin derivatives. The compounds were tested for their stability, solubility, and antimicrobial activity. Of the 17 compounds produced, ten displayed antibiotic activity and two of them improved MIC value to bacteria. Furthermore, nocardithiocin and all derivatives were stable to the light. The substitution of amino acid sequence in precursor peptide could lead the successful production of nocardithiocin derivatives and alter its property. This report represents the first step of nocardithiocin utilization, and amino acid substitution technique can be used to create desirable nocardithiocin derivatives.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thiopeptides are natural products derived from ribosomally synthesized post-translationally modified peptides (RiPPs) that possess a variety of biological properties including antimicrobial, antiparasitic, and anticancer activities (Li and Kelly 2010; Just-Baringo et al. 2014a). However, their poor pharmacokinetic properties, especially low water solubility, hampers their clinical use. To improve their properties, derivatives have been generated by several ways such as chemical synthesis (Nicolaou et al. 2003; Gross et al. 2013; Just-Baringo et al. 2014b), semi-synthesis (Naidu et al. 2006; Xu et al. 2013), and genetic modification (Bowers et al. 2010; Li et al. 2011, 2012; Young et al. 2012; Zhang and Kelly 2015) (Just-Baringo et al. 2014c). Based on the biosynthetic pathway of RiPPs compounds, DNA sequence of precursor peptide is directly related to their peptide structure (Arndt et al. 2009; Kelly et al. 2009; Morris et al. 2009). So the derivatives production of thiopeptides were achieved relatively easy by modifying the gene sequence of core peptide region among precursor peptide. The increasing information of biosynthetic genes and recent progress in the development of genetic tools and techniques have made thiopeptides more attractive as novel lead compounds for pharmaceutical agents.
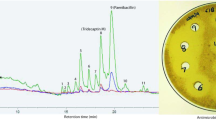
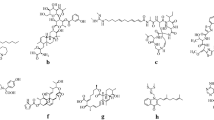
Nocardithiocin is a thiopeptide compound produced by Nocardia pseudobrasiliensis that displays antimicrobial activity against Gram-positive bacteria, especially acid-fast bacteria (Fig. 1a) (Mukai et al. 2009). Impressive antimicrobial activity against drug-resistant Mycobacterium tuberculosis has resulted in significant attention to the gene cluster responsible for nocardithiocin biosynthesis (Sakai et al. 2015). Gene disruption and expression analysis revealed a 15.2 kb cluster of 12 genes including precursor peptide (notG), as well as post-transcriptional modification enzymes that modify the peptide structure (Fig. 2a).
Like other thiopeptides, nocardithiocin is poorly soluble in water and light sensitivity was reported (Mukai et al. 2009). Herein, we attempted to improve its properties by genetic modification without diminishing its antibiotic activity. Although the actual mode of action of nocardithiocin is not elucidated yet, a number of 26-membered macrocycles thiopeptides including thiostrepton, noshiheptide, and micrococcin have been reported that interaction with ribosomal protein L11 as well as 23S rRNA through a Threonine residue (highlighted in red in Fig. 1a) cause antimicrobial activity (Bowers et al. 2010; Acker et al. 2009; Tran et al. 2017). Residues S1, C9, S10, C11, and T3 of nocardithiocin precursor peptide were thought to be related to retention of the macrocyclic structure and antimicrobial activity, respectively (Fig. 2b). In the present study, the sixth amino acid (V6) of the core peptide which exist at distal position from pyridine ring was selected to replace with the other 19 amino acids and the properties and antimicrobial activity of the derivatives were assessed.
Materials and Methods
Strains and Cultivation Conditions
Nocardithiocin producer N. pseudobrasiliensis IFM 0761 was obtained from Medical Mycology Research Center (MMRC), Chiba University, Japan. Escherichia coli DH5α was used for construction and propagation of vectors. N. pseudobrasiliensis strains were normally cultivated at 37 °C in brain–heart infusion (Becton Dickinson and Company, Tokyo, Japan) supplemented with 1% glucose and 1% glycerol (BHIgg) broth or 2% agar plates. For the nocardithiocin production, strains were cultivated at 30 °C in Czapek-Dox (CD) medium consisting of 0.6% NaNO3, 0.052% KCl, 0.152% KH2PO4, 1% glucose, 2 mM MgSO4 and trace elements.
Deletion of notG in Nocardia pseudobrasiliensis
The notG gene was removed from N. pseudobrasiliensis genome by the double cross-over method, leaving the start and stop codon to retain in-frame translation of the rest of the biosynthetic genes. The notG disruption vector was constructed using upstream and downstream notG fragments that were amplified by PCR using notG-upF (tgcctgcaggtcgagcgtcccaggtgacc) and notG-upR (acgggactactacatgacactatccttcct), and notG-downF (gatagtgtcatgtagtagtcccgtcgtggg) and notG-downR (tcctctagagtcgacgaacaacctggcatc), primer pairs, with N. pseudobrasiliensis IFM 0761genomic DNA as a template. Underlined sequences correspond to overlaps used for recombination. Both up- and down-stream fragments were ligated into the HincII site of the pNV18.1 vector using the GeneArt Seamless Cloning and Assembly Enzyme Mix (Life Technologies, Tokyo, Japan) according to the manufacturer’s protocol.
Transformation of N. pseudobrasiliensis was conducted as described previously (Sakai et al. 2015). The first cross-over strains were grown on BHIgg plates containing 100 µg/ml neomycin, and further cultivated in BHIgg liquid medium without neomycin for 3 days. Cultures were spread onto BHIgg agar medium without neomycin, and resulting colonies were picked and inoculated into BHIgg plates with and without neomycin. Colonies that grew only on BHIgg plates without neomycin were selected as double cross-over strains. Deletion of notG in the double cross-over strains was confirmed by PCR, Southern blotting, and DNA sequencing.
Expression of Site-Directed notG Mutants
Native and mutant notG expression vectors were constructed by joining PCR fragment of the notG fragment to the native promoter and nnrB terminator. Site-directed mutagenesis of notG was conducted by PCR using notG-nPF/V60X-R and V60X-F/notG-R-TrrnB primer pairs with N. pseudobrasiliensis genomic DNA as the template (X indicates any amino acid). The terminator sequence was amplified using the notG-TeenBF/TrrnB-R primer pair with the pRU1701 vector as template. All fragments were ligated into the HincII site of pNV18.1 using the GeneArt Seamless Cloning and Assembly Enzyme Mix. All expression vectors were confirmed by sequencing, and primers used for construction are listed in Table S1.
Detection and Purification of Nocardithiocin Derivatives
Constructed strains (the ΔnotG strain containing a native or mutant notG expression vector) were pre-cultured in BHIgg medium containing neomycin for 3 days at 37 °C, and 1% of each pre-culture was inoculated into CD medium containing neomycin and cultivated for 7 days at 30 °C. After cultivation, the same volume of methanol was added to the culture and incubated for at least 1 h. Samples were filtered to remove bacterial cells, and filtrates were evaporated using a rotary evaporator. The remaining water layer was extracted with ethyl acetate, and the ethyl acetate extract was dried in vacuo.
Crude extracts were analyzed to examine the production of nocardithiocin and its derivatives by HPLC (RF-20A/RF-20Ax, Shimadzu Corp., Kyoto, Japan) with a WakoPack Wakosil-II 3C18 HG column and a gradient elution of 10–100% acetonitrile in water over 50 min at a flow rate of 1 ml/min. Eluates were monitored using a fluorescence detector at an extinction wavelength of 350 nm and an emission wavelength of 450 nm (RF-20A/RF-20Axs, Shimadzu Corp.) and a mass spectrometer (LCMS-2020, Shimadzu Corp.). Nocardithiocin derivatives were purified from crude extracts by preparative HPLC using a Cosmosil 5C18 MS-II column (10 × 150 mm, NACALAI TESQUE, Inc., Kyoto, Japan) with a gradient elution of 30–60% acetonitrile in water over 20 min followed by 100% acetonitrile for 5 min, monitored by a fluorescence detector.
Structural Confirmation
The crude extracts were first analyzed by LC–MS and the structure of nocardithiocin derivatives were determined by LC–MS/MS analysis. LC–MS/MS was conducted using TripleTOF 6600 hybrid quadrupole-time-of-flight tandem mass spectrometer (Sciex, Concord, Canada) equipped with an electrospray ionization interface. An ODS-80Ts column (5 µm, 2.0 × 150 mm, TOSOH, Tokyo, Japan) was used with mobile phases 0.1% formic acid in water (A) and 0.1% formic acid in acetonitrile (B). LC gradient condition was initially 10% B to 100% B (30 min) and 10% (40 min) at flow rate of 0.2 ml/min. Information dependent acquisition was performed in positive ion mode. MS spectra were acquired from m/z 100 to m/z 2000. The 10 most abundant ions were selected for subsequent fragmentation, and MS/MS spectra were acquired from m/z 100 to m/z 2000.
Testing Light and Temperature Stability of Nocardithiocin and Its Derivatives
Purified compounds were dissolved in DMSO at a final concentration of 0.5 mg/ml and divided into three parts. One part was covered with foil and kept in the dark, one part was placed under a fluorescent lamp (light) for 4 days at room temperature, and the final part was stored at 4 °C in the dark. Samples were analyzed by HPLC to quantify the amount of remaining nocardithiocin derivatives. Light stability was calculated by comparing the HPLC peak area of samples stored in the light and dark at room temperature. Similarly, temperature stability was calculated by comparing the dark samples stored at room temperature (30 °C, in summer) and 4 °C for 4 days.
Testing Solubility of Nocardithiocin and Its Derivatives
Purified dried samples were dissolved in DMSO or water at a final concentration of 1 mg/ml and incubated overnight. Insoluble matter was removed by centrifugation, and soluble compounds were quantified using HPLC by comparing the peak areas of compounds in DMSO (completely dissolved 100%) and water.
Testing Antimicrobial Activity
Antimicrobial activity against several bacteria and fungi (Escherichia coli, Staphylococcus aureus, Bacillus subtilis, Micrococcus luteus, Aspergillus niger, Trichophyton mentagrophytes, Candida albicans, and Cryptococcus neoformans) was determined as described previously (Nagai et al. 1993) using the microbroth dilution method. These strains were obtained from the Medical Mycology Research Center (MMRC), Chiba University. The minimum inhibitory concentration (MIC) against mycobacteria was determined using the Clinical Laboratory and Standards Institute (CLSI) protocol (M24-A2). Mycobacterial strains were obtained from the National Institute of Infectious Diseases (NIID). MIC and IC50 (half maximal inhibitory concentration) values were determined for antimicrobial activity against bacteria and fungi, respectively.
Results
Deletion of notG in N. pseudobrasiliensis
Before expressing the mutant notG gene, native notG which encode the precursor peptide was removed from the N. pseudobrasiliensis IFM 0761 genome (Fig. 2). Deletion of notG in ΔnotG strains was confirmed by PCR and sequencing (data not shown). HPLC analysis confirmed the absence of nocardithiocin production in ΔnotG strains (Fig. 3). Three ΔnotG strains were checked and all of them exhibited the same morphology, so one of them was selected as the host for production of nocardithiocin derivatives.
Production of Nocardithiocin Derivatives
The native notG expression vector and 19 mutant notG expression vectors with the sixth amino acid (V6) replaced by the other 19 amino acids (Fig. 2b) were each transformed into the ΔnotG strain, and transformants were cultivated on agar plates. Several colonies were picked to confirm the production of nocardithiocin or its derivatives by HPLC, and the highest producer was selected for subsequent experiments. As shown in Fig. 4, several fluorescence peaks were detected that appeared to be nocardithiocin derivatives. Peaks were significant for ten mutants (A, F, L, I, M, Y, S, T, N, and Q substitutions), indicating successful production of nocardithiocin derivatives. Substitution with A, L, M, S, T, or Q resulted in two or three clear fluorescence peaks with slightly different m/z values, were numbered in order of elution time in Fig. 4. The information of replaced amino acid and MS data from the fluorescence peaks are summarized in Table 1.
Structural Confirmation of Nocardithiocin Derivatives
Nocardithiocin and its derivatives were purified by preparative HPLC. As shown in Table 1, peptides in which the sixth amino acid was replaced by F, M, Y, and N did not yield the expected m/z values, whereas peptides containing A, L, I, S, T, and Q matched the expected m/z values, although peptides with A, L, S, T, and Q produced additional unknown peaks. To further confirm the structure of these compounds, LC-MS/MS analysis was performed and the MS/MS results and proposed structures are shown in Fig. S1 and Table S2. Compounds with the expected m/z values (A3, L2, I, S2, T2, and Q2) produced fragmentation patterns comparable to that of nocardithiocin (i.e., fragment ions were observed with the expected mass shifts). Therefore, these compounds were assumed to be expected nocardithiocin derivatives. The different fragmentation pattern of the other compounds with that of nocardithiocin indicating that they have other structures than expected one amino acid substituted derivatives. These non-expected derivatives were divided into two groups based on their MS/MS fragmentation patterns. Group 1 consists of peptides A1, F, L1, M1, M2, T1, and N, all of which keep the similar structure to nocardithiocin and its expected derivatives. However for example, A1 and M1 have + 6 and + 4 m/z value different from expected derivative, respectively. This may be caused by the failure of thiazole ring formation next to the substituted amino acid. And the other F, L1, Y, T1 and N showed − 12 m/z value difference with expected derivatives and M2 showed − 12 m/z value difference with M1 may indicating the defective of methyltransferase activity at S4. Group 2 is composed of A2, S1, and Q1 peptides whose fragment patterns were completely different from nocardithiocin and expected derivatives, so we failed to determine their structures but the compounds in this group showed same fragment at lower m/z values (≤ 736).
Properties of Nocardithiocin Derivatives
The light sensitivity of purified compounds was tested and compared with that of native nocardithiocin. In a previous study, nocardithiocin was reported to be unstable in the light (Mukai et al. 2009). However, the results of the present study (Fig. 5a) suggest that neither nocardithiocin nor its derivatives (including unexpected compounds) were light sensitive. Additionally, the stability at room temperature was also examined, and nocardithiocin and its derivatives were shown to be stable (Fig. 5b).
Analysis of stability tests. a Stability of nocardithiocin and purified compounds in the presence of light. The percentage of the compound remaining was calculated by comparing the peak area following storage in the light and dark. b Stability of nocardithiocin and purified compounds stored at room temperature. The percentage of the compound remaining was calculated by comparing the peak area following storage at °C and room temperature (30 °C, in summer) for 4 days. Nocardithiocin and its derivatives are indicated by orange bars, and compounds belonging to groups 1 and 2 are indicated by green and blue bars, respectively
Since the solubility of thiopeptide compounds is very low (Li and Kelly 2010; Just-Baringo et al. 2014b), we tested the solubility of nocardithiocin and its derivatives. Compared the solved amount in DMSO (completely soluble), only 0.14% (1.2 µM) of nocardithiocin was soluble in water. All nocardithiocin derivatives showed a slight improvement in solubility in water (Fig. 6), and Q2 was the most soluble among expected derivative (3.07%; 25.8 µM).
Solubility of nocardithiocin and purified compounds in water. a The level of solubility in DMSO was assigned a value of 100%, and peak areas of compounds dissolved in water were compared with peak areas of compounds dissolved in DMSO. b The concentration of dissolved compound was calculated using m/z values. Nocardithiocin and its derivatives are indicated by orange bars, and compounds belonging to groups 1 and 2are indicated by green and blue bars, respectively
Antimicrobial Activity of Nocardithiocin Derivatives
The antimicrobial activity of nocardithiocin derivatives was tested against several bacteria and fungi (Table 2). Expected nocardithiocin derivatives A3, L2, I, S2, T2, and Q2 retained antimicrobial activity, but compounds with unexpected structures lost its activity, except for compound M1. Furthermore, expected derivatives and compounds M1 and M2 exhibited activity against drug-resistant Mycobacterium tuberculosis strains (Table 3). In most cases, native type of nocardithiocin showed stronger antimicrobial activity than its derivatives, but compound I displayed four times and two times greater activity against Bacillus subtilis and Micrococcus luteus, respectively. And compound L2 showed two times higher activity against M. luteus (Table 2).
Discussion
Nocardithiocin is a thiopeptide compound produced by N. pseudobrasiliensis (Mukai et al. 2009) that displays powerful activity against drug-resistant M. tuberculosis, making it a promising novel antibacterial drug. However, its clinical use is hampered by low water solubility and light sensitivity, so the property should be improved for further application. Identification of the nocardithiocin biosynthesis gene cluster (Sakai et al. 2015) revealed that it is synthesized through modification of a ribosomally translated precursor peptide, similar to other thiopeptides (Wieland Brown et al. 2009; Liao et al. 2009). This biosynthetic feature prompted us to modify the nocardithiocin structure by altering the amino acid sequence of the precursor peptide to improve its properties.
Firstly, we removed the native notG gene encoding the precursor peptide from the genome of N. pseudobrasiliensis, and the notG deletion strain was used as the host for nocardithiocin derivative production. The mutant fragment of notG were prepared by PCR-based site directed mutagenesis method and expressed under the notG native promoter, because production of derivatives under the control of constitutive promoter PermE (Labes et al. 1997) was much lower (Fig. S2) than native notG promoter. Expression of notG under the control of PermE was actually higher than the native promoter, suggesting that coordinated regulation with other genes in the cluster is important for optimal nocardithiocin production. Therefore, further improvement of nocardithiocin production may be achieved by modifying the expression levels of the transcriptional regulator notD in the gene cluster.
Among the 19 amino acid substituted derivatives, 10 amino acid produced clear HPLC fluorescent peaks under nocardithiocin detection condition, but nine (P, W C, G, D, E, H, K, and R) did not show any significant compound production (Fig. 3). The imino acid structure of P, the large side chain of W, and the small side chain of G may prevent the enzymatic reaction(s) required to synthesize nocardithiocin derivatives. Similarly, a negative (D and E) or positive (H, K, and R) charge could disrupt derivative synthesis. Finally, the sulfur-containing side chain of C could form a thiazole ring following the action of cyclodehydratase, resulting in three adjacent thiazoles at positions C5 and C7, which could explain the failure to synthesize this derivative. Several reports showed the preference of amino acids depending on the replaced position in thiopeptides (Young et al. 2012; Zhang et al. 2012; Tran et al. 2017).
Substitution with A, L, M, S, T, or Q resulted in the production of two or three major compounds, implying that nocardithiocin biosynthetic enzymes or other enzymes might have the ability to produce diverse compounds from nocardithiocin precursor peptides with altered amino acid sequences. Other than the compound M2, expected nocardithiocin derivatives eluted later in the elution profile (Table 1). Comparison of the MS/MS fragmentation patterns with that of nocardithiocin (V) confirmed that A3, L2, I, S2, T2, and Q2 were the expected derivatives (Fig. S1). Although the exact structures have not been determined, unexpected compounds were classified into two groups based on their MS/MS fragmentation patterns. The fragmentation pattern of group 1 compounds suggested that their similar structure with nocardithiocin derivatives because several MS/MS fragments were coincident with those of nocardithiocin (Table S2). The speculated structure of group 1 compounds from MS/MS spectrum indicated the partial or defective reaction of thiazole formation or addition of methyl group (Fig. S1). The size of amino acid residue may cause this incomplete reaction. The properties of the purified compounds were tested, and none of them displayed the previously reported light sensitivity (Mukai et al. 2009). Although nocardithiocin and its derivatives in crude samples appeared as single peaks in the HPLC analysis, purified samples contained two separated peaks (Fig. S3). After single peaks were collected by HPLC, collected samples showed two separate peaks with similar proportions in the 1st HPLC. This phenomenon could have been interpreted as light-induced degradation in the previous report. Furthermore, the previous study used a glass tubes for storing nocardithiocin samples, and nocardithiocin may be adsorbed to glass tubes due to its hydrophobic and basic nature.
Nocardithiocin and produced compounds were not degraded under light irradiation, so light sensitivity remains as major limitation of clinical use. However the produced derivatives remained poorly soluble in water (Fig. 5). In previous report, water-soluble analogues of nocathiacin prepared by semi-synthesis retained antimicrobial activity (Xu et al. 2013). In that report, modification of side chains at the tail end of the peptide improved its water solubility, and similar modifications could also enhance the water solubility of nocardithiocin derivatives without diminishing antimicrobial activity.
The antimicrobial activity of nocardithiocin and its derivatives was tested against several microbes. Among the 17 successfully purified compounds, 10 showed some antimicrobial activity. While native nocardithiocin displayed higher activity than most of its derivatives, compounds L2 and I displayed slightly higher antimicrobial activity against Gram-positive bacteria (Table 2). Young et al. (2012) also reported the amino acid replacement of G37468 (T2C) increased twofold improved activity against B. subtilis implying the potential of amino acid replacement of thiopeptide compounds for improvement of its antimicrobial activity. Compounds L2, I, and T2 that exhibited relatively strong antimicrobial activity share similar side chains at the sixth position to that present in nocardithiocin (V6), and similarly low water solubility. This result indicated that a branched methyl group at the sixth position of nocardithiocin makes some contribution to its antimicrobial activity. Although the sixth amino acid was chosen for mutation based on the assumption that it is not crucial for antimicrobial activity, substitution clearly gave a significant impact to antimicrobial activity. The prepared compounds which have different structure with expected nocardithiocin derivatives did not display clear antimicrobial activity may implying the importance of methyl group on S4 or proper thiazole ring formation for nocardithiocin activity.
In this report, among 13 amino acid residue of nocardithiocin, only the sixth amino acid was substituted, and many more derivatives could be prepared by replacing other position of amino acid residue(s) and/or by using unusual amino acids other than the 19 canonical amino acids found in native proteins (Luo et al. 2016). The relationship of structure–activity could not be deeply analyzed yet, increased information will facilitate the rational design of nocardithiocin derivatives with desirable properties in future studies (Wolf et al. 2014).
References
Acker MG, Bowers AA, Walsh CT (2009) Generation of thiocillin variants by prepeptide gene replacement and in vivo processing by Bacillus cereus. J Am Chem Soc 131(48):17563–17565
Arndt HD, Schoof S, Lu JY (2009) Thiopeptide antibiotic biosynthesis. Angew Chem Int Ed 48(37):6770–6773
Bowers AA, Acker MG, Koglin A, Walsh CT (2010) Manipulation of thiocillin variants by prepeptide gene replacement: structure, conformation, and activity of heterocycle substitution mutants. J Am Chem Soc 132(21):7519–7527
Gross S, Nguyen F, Bierschenk M, Sohmen D, Menzel T, Antes I, Wilson D, Bach T (2013) Amythiamicin D and related thiopeptides as inhibitors of the bacterial elongation factor EF-Tu: modification of the amino acid at carbon atom C2 of ring C dramatically influences activity. ChemMedChem 8:1954–1962
Just-Baringo X, Bruno P, Pitart C, Vila J, Albericio F, Álvarez M (2014a) Dissecting the structure of thiopeptides: assessment of thiazoline and tail moieties of baringolin and antibacterial activity optimization. J Med Chem 57:4185–4195
Just-Baringo X, Albericio F, Álvarez M (2014b) Thiopeptide antibiotics: retrospective and recent advances. Mar Drugs 12(1):317–351
Just-Baringo X, Albericio F, Álvarez M (2014c) Thiopeptide engineering: a multidisciplinary effort towards future drugs. Angew Chem Int Ed Engl 53(26):6602–6616
Kelly WL, Pan L, Li C (2009) Thiostrepton biosynthesis: prototype for a new family of bacteriocins. J Am Chem Soc 131(12):4327–4334
Labes G, Bibb M, Wohlleben W (1997) Isolation and characterization of a strong promoter element from the Streptomyces ghanaensis phage I19 using the gentamicin resistance gene (aacC1) of Tn 1696 as reporter. Microbiology 143:1503–1512
Li C, Kelly WL (2010) Recent advances in thiopeptide antibiotic biosynthesis. Nat Prod Rep 27:153–164
Li C, Zhang F, Kelly WL (2011) Heterologous production of thiostrepton A and biosynthetic engineering of thiostrepton analogs. Mol Biosyst 7(1):82–90
Li C, Zhang F, Kelly WL (2012) Mutagenesis of the thiostrepton precursor peptide at Thr7 impacts both biosynthesis and function. Chem Commun 48(4):558–560
Liao R, Duan L, Lei C, Pan H, Ding Y, Zhang Q, Chen D, Shen B, Yu Y, Liu W (2009) Thiopeptide biosynthesis featuring ribosomally synthesized precursor peptides and conserved posttranslational modifications. Chem Biol 16(2):141–147
Luo X, Zambaldo C, Liu T, Zhang Y, Xuan W, Wang C, Reed SA, Yang PY, Wang RE, Javahishvili T, Schultz PG, Young TS (2016) Recombinant thiopeptides containing noncanonical amino acids. Proc Natl Acad Sci USA 113(13):3615–3620
Morris RP, Leeds JA, Naegeli HU, Oberer L, Memmert K, Weber E, LaMarche MJ, Parker CN, Burrer N, Esterow S, Hein AE, Schmitt EK, Krastel P (2009) Ribosomally synthesized thiopeptide antibiotics targeting elongation factor Tu. J Am Chem Soc 131(16):5946–5955
Mukai A, Fukai T, Hoshino Y, Yazawa K, Harada K, Mikami Y (2009) Nocardithiocin, a novel thiopeptide antibiotic, produced by pathogenic Nocardia pseudobrasiliensis IFM 0757. J Antibiot 62(11):613–619
Nagai H, Mikami Y, Yazawa K, Gonoi T, Yasumoto T (1993) Biological activities of novel polyether antifungals, gambieric acids A and B from a marine dinoflagellate Gambierdiscus toxicus. J Antibiot 46(3):520–522
Naidu BN, Sorenson ME, Matiskella JD, Li W, Sausker JB, Zhang Y, Connolly TP, Lam KS, Bronson JJ, Pucci MJ, Yang H, Ueda Y (2006) Synthesis and antibacterial activity of nocathiacin I analogues. Bioorg Med Chem Lett 16(13):3545–3549
Nicolaou KC, Nevalainen M, Zak M, Bulat S, Bella M, Safina BS (2003) Synthetic studies on thiostrepton: construction of thiostrepton analogues with the thiazoline-containing macrocycle. Angew Chem Int Ed Engl 42(29):3418–3424
Sakai K, Komaki H, Gonoi T (2015) Identification and functional analysis of the Nocardithiocin Gene cluster in Nocardia pseudobrasiliensis. PLoS ONE 10(11):e0143264
Tran HL, Lexa KW, Julien O, Young TS, Walsh CT, Jacobson MP, Wells JA (2017) Structure-activity relationship and molecular mechanics reveal the importance of ring entropy in the biosynthesis and activity of a natural product. J Am Chem Soc 139(7):2541–2544
Wieland Brown LC, Acker MG, Clardy J, Walsh CT, Fischbach MA (2009) Thirteen posttranslational modifications convert a 14-residue peptide into the antibiotic thiocillin. Proc Natl Acad Sci USA 106(8):2549–2553
Wolf A, Schoof S, Baumann S, Arndt HD, Kirschner KN (2014) Structure-activity relationships of thiostrepton derivatives: implications for rational drug design. J Comput Aided Mol Des 28(12):1205–1215
Xu L, Farthing AK, Dropinski JF, Meinke PT, McCallum C, Hickey E, Liu K (2013) Synthesis and antibacterial activity of novel water-soluble nocathiacin analogs. Bioorg Med Chem Lett 23(1):366–369
Young TS, Dorrestein PC, Walsh CT (2012) Codon randomization for rapid exploration of chemical space in thiopeptide antibiotic variants. Chem Biol 19(12):1600–1610
Zhang F, Kelly WL (2015) Saturation mutagenesis of TsrA Ala4 unveils a highly mutable residue of thiostrepton A. ACS Chem Biol 10(4):998–1009
Acknowledgements
We thank the National Institute of Infectious Diseases (NIID) for providing M. tuberculosis strains.
Funding
This study was funded by a JSPS KAKENHI award (Grant No. JP25305012).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Research Involving Human Participants and/or Animals
This article does not contain any studies with human participants or animal performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Sakai, K., Hara, Y., Ishibashi, M. et al. Characterization of Nocardithiocin Derivatives Produced by Amino Acid Substitution of Precursor Peptide notG. Int J Pept Res Ther 26, 281–290 (2020). https://doi.org/10.1007/s10989-019-09836-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-019-09836-0