Abstract
Structural Genomics has been successful in determining the structures of many unique proteins in a high throughput manner. Still, the number of known protein sequences is much larger than the number of experimentally solved protein structures. Homology (or comparative) modeling methods make use of experimental protein structures to build models for evolutionary related proteins. Thereby, experimental structure determination efforts and homology modeling complement each other in the exploration of the protein structure space. One of the challenges in using model information effectively has been to access all models available for a specific protein in heterogeneous formats at different sites using various incompatible accession code systems. Often, structure models for hundreds of proteins can be derived from a given experimentally determined structure, using a variety of established methods. This has been done by all of the PSI centers, and by various independent modeling groups. The goal of the Protein Model Portal (PMP) is to provide a single portal which gives access to the various models that can be leveraged from PSI targets and other experimental protein structures. A single interface allows all existing pre-computed models across these various sites to be queried simultaneously, and provides links to interactive services for template selection, target-template alignment, model building, and quality assessment. The current release of the portal consists of 7.6 million model structures provided by different partner resources (CSMP, JCSG, MCSG, NESG, NYSGXRC, JCMM, ModBase, SWISS-MODEL Repository). The PMP is available at http://www.proteinmodelportal.org and from the PSI Structural Genomics Knowledgebase.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Structural genomics has been successful in determining the structures of many unique proteins in a high throughput manner. Since 2001, these efforts have resulted in more than 6,772 structure depositions to the PDB [1], 3,251 of which are from the NIH-sponsored Protein Structure Initiative (PSI) Centers. Nevertheless, the number of known protein sequences is much larger than the number of experimentally solved protein structures, and this number is growing at an unprecedented rate due to large-scale genome sequencing and meta-genomics projects [2]. Homology (or comparative) modeling methods make use of experimental protein structures to build models for evolutionarily related proteins. This technique predicts the three-dimensional structure of a given protein sequence (target) based primarily on its alignment to one or more proteins of known structure (templates). For every experimentally determined structure, often models for hundreds of proteins can be derived using a variety of established methods for comparative protein structure modeling methods, which dramatically increases the structural coverage of protein sequences. Structural genomics and homology modeling thereby complement each other in the exploration of protein structure space [3].
The quality of a comparative protein structure model depends on the evolutionary distance between the sequence of the target protein to be modeled and the template structure. In general comparative models sharing more than 40% of sequence identity with the template are considered high-quality models. Comparative protein structure models are routinely used in a widespread range of biomedical applications such as the rational design of mutagenesis experiments, the interpretation of disease-related mutations, or structure-based virtual screening studies [4–6]. However, despite this tremendous growth in protein structure data in recent years, structural information is often not used to its full extent in biomedical research projects. One reason is that experimental protein structures are only available for a small subset of all known protein sequences. A significant impediment in using 3D-models effectively is that model information on a specific protein is distributed over different web sites in heterogeneous formats, using incompatible accession code systems. We have developed the Protein Model Portal (PMP, http://www.proteinmodelportal.org/) as part of the PSI Structural Genomics Knowledgebase (http://kb.psi-structuralgenomics.org/KB/) in order to provide a single portal which gives access to the various models that can be leveraged from PSI targets and other experimental protein structures [7, 8]. The Portal has been presented at the NIGMS PSI Bottlenecks Workshop. Here, we describe the challenges that exist for building such a model portal, present the technical implementation, show specific examples for accessing the portal, briefly report on the community workshop held on “Applications of Protein Models in Biomedical Research”, and discuss future developments of the project.
Bottlenecks
While information about experimental structures is maintained within a single resource, the world wide PDB [9], access to protein model information is by far more complicated. The major reasons are listed here:
-
Various algorithmic approaches [10–16] with different strengths and weaknesses have been developed for building 3-dimensional models of proteins. Also the quality of individual models highly depends on the evolutionary proximity to the selected structural templates. Finally the update frequency of different services varies. For these reasons, a consensus view of the results obtained from different modeling resources is very often helpful in identifying the most reliable solution.
-
The content of the PDB database is organized around experimental information (structure centric), whereas model information is based on sequence databases (sequence centric); difficulties arise due to the fact that sequence database content and accession codes are often transient and frequently incompatible between different databases.
-
Models often cover only fragments of a protein sequence. The different segments modeled for a given target protein are usually based on various alternative alignments, alternative templates, or result from diverse modeling procedures.
-
Models are not stable information by itself, but reflect the current status of sequence and structure databases at time of modeling. Models have to be revised, as new, more suitable template structures become available. Also, changes in the primary sequence of target proteins in the sequence databases necessitate remodeling.
-
Models have typically few intrinsic annotations (in comparison to UniProt [17] or PDB databases) which go beyond the alignment to the template structure. Changes in the functional annotation of a protein database entry (e.g., UniProt; InterPro [18]) occur independent from and more frequently than changes of the primary amino acid sequence, which would require rebuilding of the model.
-
Models are not experimental observations, but the results of theoretical predictions. While standards for describing the reliability and limitations of the most commonly used experimental structure determination techniques have been established, the spectrum of reliability and applicability of current modeling methods is broad and the level of uncertainty is significantly higher [19, 20]. Therefore, detailed information about each individual model is crucial for assessing its expected accuracy, and thereby determining its scope of applicability [3].
-
The number of currently available models is orders of magnitudes larger than that of experimental structures (ca. 7 million vs. 50,000) and, therefore, poses technical challenges in efficient data handling.
-
As model information is complex—one needs to estimate the expected accuracy and reliability of a specific model or for parts thereof; users of models are often unsure to which extent a model can be used for a given application.
To summarize, model information is heterogeneous, highly context dependent, dynamic, and consists of large volumes of data. Consequently, a community workshop held at Rutgers on archiving structural models of biological macromolecules has recommended no longer accepting models as part of the PDB archive, and to establish a portal for protein model information [21]. Following this recommendation, and taking into account the challenges mentioned before, we have developed the PMP as a web portal which federates resources from model providers, experimental protein structures (PDB), and functional annotation databases.
The aim of the PMP is to foster the effective usage of molecular model information in biomedical research by providing unified access independent of individual sequence nomenclature and accession code system and by supporting the development of data standards to facilitate exchange of information and algorithms. Furthermore, PMP aims to provide a forum for discussions between developers of modeling methods and applied biomedical researchers on best-practices, including methods for quality assessment, guidelines for the publication of theoretical models, and educational resources on usage of models for different biological applications.
Architecture of the protein model portal
A common reference system for target proteins is established based on the UniProt database [17] by calculating cryptographic md5 hashes for all full length amino acid sequences. This approach combines database entries with identical amino acid sequences, while proteins sequences which differ by at least one amino acid are kept separate (Fig. 1). This reference system is continuously updated with every new UniProt release. The protein modeling portal federates protein model data from different providers (CSMP, JCSG, MCSG, NESG, NYSGXRC, JCMM, ModBase [10], and SWISS-MODEL Repository [22, 23]) by integrating model meta information: the segment of a target protein for which a model is available, template structure (PDB ID and chain), and sequence identity between the target and the template sequences. Searchable indices for matching amino acid sequences, sequence based similarity searches (using BLAST [24]), and accession code database queries are generated by mapping the model data into the md5-based reference system. Mappings between various database accession code systems are derived from iProClass [25]. The Protein Model Portal database containing this information has been implemented using MySQL (Fig. 2).
Reference system based on md5 cryptographic hash sums for UniProt full-length target sequences. In this system, identical target protein sequences are grouped together independent from their individual database accession codes (e.g., Hemoglobin beta chain from Human, Chimpanzee, and Bonobo), while entries which differ in at least one amino acid position are kept separate (e.g., 7E → V variant of Human sickle cell anemia hemoglobin)
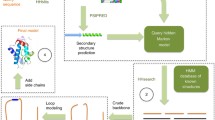
Schematic flow of data in Protein Model Portal. Meta information about the available models, i.e., the target protein, template structure, and sequence identity, is retrieved from each partner resource. The UniProt database is used to generate a reference system based on md5 cryptographic hash sums of the full-length primary sequences. Searchable indices are generated for all proteins with model information, allowing for accession code-based queries, matching of amino acid sequence fragments, and sequence similarity searches. The portal communicates with all partner resources and the PSI structural genomics knowledge base via Web services. The three-dimensional coordinates of a model, as well as functional annotation information from UniProt and InterPro is retrieved dynamically in real time when required to generate the web page
The PMP is queried from the PSI structural genomics knowledgebase using web services based on SOAP (Simple Object Access Protocol) and REST (Representational State Transfer). Users can also access the Portal directly using the web-based graphical user interface implemented with PHP at http://www.proteinmodelportal.org/. Functional annotation for individual target proteins is retrieved in real-time from the respective annotation providers using web services: information about individual domains is retrieved from the InterPro database using DAS [26], whereas sequence annotations from the UniProt database are retrieved using REST from the UniProt server (http://www.uniprot.org). The three-dimensional coordinates of the individual models are stored at the different model providers and retrieved by the portal in real-time when required for the visualization of the model overview page. Preview images of the structural model, generated using Molscript [27], Render, and Raster3D [28], provide the first quick preview of the protein model. The alignment between the target sequence and the template is inferred dynamically on-the-fly by structural superposition of the final model to the template structure using MAMMOTH [29]. The consequent use of portal technologies allows federating a set of heterogeneous resources into a single portal while at the same time ensuring consistency of the exchanged data.
Web access to PMP
The direct entry point for the PMP is the website http://www.proteinmodelportal.org. The query form allows the user to search using database accession codes such as UniProt, IPI, GenBank, RefSeq, or Entrez identifiers, to search for models built on a specific template structure, or to query the portal database with the amino acid sequence of a protein of interest. For a specific target protein, all models available from the participating resources as well as experimental structures in the PDB are displayed in graphical and tabular form (Fig. 3). When a specific model is selected, the model detail page shows information on the template which was used for building the model, provides a graphic preview of the model structure, as well as the target-template sequence alignment (Fig. 4). The model can be displayed as ribbon representation inside the webpage using the Astex Viewer plugin [30]. A download link points to the original home page of the model provider, where additional information about the model building procedure can be found. Information about the target primary sequence is retrieved dynamically from the UniProt database, listing all entries that share the same primary sequence. The domain structure of the target sequence is displayed based on PFAM domain annotation [31] which is retrieved dynamically from the InterPro database. Finally, the PMP provides links to services for template selection, model building and quality assessment.
Typical view of a model detail page. Information about the model provider, the segment of the target protein (e.g., MLP-like protein 34; Arabidopsis thaliana) covered by the model, and the template structure used for model building, are stored in the portal database. All other information required for building the webpage, such as the coordinates of the model, the PFAM domain structure, and UniProt annotation of the protein sequence, is retrieved dynamically
Model portal content
The current release of the portal allows searching 7.6 million model structures provided by the different partner sites: CSMP, JCSG, MCSG, NESG, NYSGXRC, JCMM, ModBase [32], SWISS-MODEL Repository [33]. At least one model is available for 3.0 million unique sequences out of the 7.1 million distinct sequences of the current UniProt release (14.4). The distribution of chain lengths of the models shows a maximum around 150 residues, indicating that the majority of models consist of single domains. However, more than one quarter of the models have significantly longer chains of more than 300 residues (Fig. 5). As model quality is correlated with sequence similarity between target and template, we have analyzed the best available model (i.e., the one with highest sequence identity) for each residue in the model portal database (Fig. 6). As expected, for the majority of modeled residues (41%) the templates shared between 20% and 40% sequence identity with the target.
Distribution of chain length. The histogram shows the length distribution of models provided by the model portal. The maximum around 150 residues indicates that the majority of models consist of single domains. However, more than one quarter of the models have significantly longer chains of more than 300 residues
Model quality on residue level. For each residue, the model with the highest sequence identity between target and template is considered. The pie chart shows the percentage of residues which can be modeled at a certain identity level. For the majority of modeled residues (41%) the targets shares between 20% and 40% sequence identity with the templates
Community workshop
A ‘‘Workshop on Applications of Protein Models in Biomedical Research’’ was held in San Francisco in July 2008, where protein structure modelers explored how models are used in biomedical research, and which requirements and challenges exist for the different applications. The workshop program involved a first day of 16 presentations on topics that ranged from the coverage of protein sequence-structure space to the uses of modeling in medicine. On the second day, open questions were addressed by four independent discussion groups. The participants discussed the state-of-the-art in applying molecular modeling to biomedical problems, requirements and challenges for various applications, as well as ways to strengthen the collaboration between the modeling and experimental communities. The PMP has been recognized as an important means for increasing the impact of molecular modeling on biology and medicine, and specific recommendations for the further development of the PMP were made. The outcome of the workshop will be made available for the community in the form of a white paper.
Conclusion and outlook
We have established a single portal that allows users to simultaneously and transparently perform searches against all model information made available by the participating sites, using various protein identifiers or amino acid sequence as the query input. This is achieved by federating the distributed resources via web services. The next step in the development of the PMP will include a common interface for submitting interactive modeling requests to different automated modeling services, tools for comparing and assessing the quality of protein structural models, and the possibility to map a wide variety of functional annotations to the protein models. The consequent usage of Web services will not only allow to coordinate information provided by other services within the portal, but also to complement sequence-based resources such as genome browsers, with model-based information. The development of widely accepted standards for data exchange and of tools for quality assessment of models would be one of the challenges in the future, which are expected to be addressed during a second PMP workshop in 2009.
Abbreviations
- CSMP:
-
Center for structures of membrane proteins
- JCMM:
-
Joint center for molecular modeling
- JCSG:
-
Joint center for structural genomics
- MCSG:
-
Midwest center for structural genomics
- NESG:
-
Northeast structural genomics consortium
- NYSGXRC:
-
New York SGX research center for structural genomics
- PDB:
-
Protein Data Bank
- PMP:
-
Protein Model Portal
- PSI SGKB:
-
PSI structural genomics knowledgebase
- PSI:
-
Protein structure initiative
- SIB:
-
Swiss Institute of Bioinformatics
References
Berman H et al (2007) The worldwide Protein Data Bank (wwPDB): ensuring a single, uniform archive of PDB data. Nucleic Acids Res 35:D301–D303. doi:10.1093/nar/gkl971
Yooseph S et al (2007) The sorcerer II global ocean sampling expedition: expanding the universe of protein families. PLoS Biol 5:e16. doi:10.1371/journal.pbio.0050016
Baker D, Sali A (2001) Protein structure prediction and structural genomics. Science 294:93–96. doi:10.1126/science.1065659
Hillisch A, Pineda LF, Hilgenfeld R (2004) Utility of homology models in the drug discovery process. Drug Discov Today 9:659–669. doi:10.1016/S1359-6446(04)03196-4
Peitsch MC (2002) About the use of protein models. Bioinformatics 18:934–938. doi:10.1093/bioinformatics/18.7.934
Tramontano A (2008) The biological applications of protein models. In: Schwede T, Peitsch MC (eds) Computational structural biology. World Scientific Publishing, Singapore
Berman HM (2008) Harnessing knowledge from structural genomics. Structure 16:16–18. doi:10.1016/j.str.2007.12.003
Berman HM et al (2008) The protein structure initiative structural genomics knowledgebase. Nucleic Acids Res. Database Issue (in press)
Berman H, Henrick K, Nakamura H (2003) Announcing the worldwide Protein Data Bank. Nat Struct Biol 10:980. doi:10.1038/nsb1203-980
Pieper U et al (2006) MODBASE: a database of annotated comparative protein structure models and associated resources. Nucleic Acids Res 34:D291–D295. doi:10.1093/nar/gkj059
Schwede T et al (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385. doi:10.1093/nar/gkg520
Yeats C et al (2006) Gene3D: modelling protein structure, function and evolution. Nucleic Acids Res 34:D281–D284. doi:10.1093/nar/gkj057
Mirkovic N et al (2007) Strategies for high-throughput comparative modeling: applications to leverage analysis in structural genomics and protein family organization. Proteins 66:766–777. doi:10.1002/prot.21191
Schwede T et al (2008) Protein structure modeling. In: Schwede T, Peitsch MC (eds) Computational structural biology. World Scientific Publishing, Singapore
Zhang Y (2007) Template-based modeling and free modeling by I-TASSER in CASP7. Proteins 69(Suppl 8):108–117. doi:10.1002/prot.21702
Chivian D, Baker D (2006) Homology modeling using parametric alignment ensemble generation with consensus and energy-based model selection. Nucleic Acids Res 34:e112. doi:10.1093/nar/gkl480
Bairoch A et al (2005) The universal protein resource (UniProt). Nucleic Acids Res 33:D154–D159. doi:10.1093/nar/gki070
Mulder NJ, Apweiler R (2008) The InterPro database and tools for protein domain analysis. Curr Protoc Bioinformatics. Chapter 2:Unit 2.7
Battey JN et al (2007) Automated server predictions in CASP7. Proteins 69(Suppl 8):68–82. doi:10.1002/prot.21761
Kopp J et al (2007) Assessment of CASP7 predictions for template-based modeling targets. Proteins 69(Suppl 8):38–56. doi:10.1002/prot.21753
Berman HM et al (2006) Outcome of a workshop on archiving structural models of biological macromolecules. Structure 14:1211–1217. doi:10.1016/j.str.2006.06.005
Kopp J, Schwede T (2004) The SWISS-MODEL repository of annotated three-dimensional protein structure homology models. Nucleic Acids Res 32:D230–D234. doi:10.1093/nar/gkh008
Kopp J, Schwede T (2006) The SWISS-MODEL repository: new features and functionalities. Nucleic Acids Res 34:D315–D318. doi:10.1093/nar/gkj056
Altschul SF et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. doi:10.1093/nar/25.17.3389
Huang H et al (2007) Integration of bioinformatics resources for functional analysis of gene expression and proteomic data. Front Biosci 12:5071–5088. doi:10.2741/2449
Jenkinson AM et al (2008) Integrating biological data—the Distributed Annotation System. BMC Bioinformatics 9(Suppl 8):S3. doi:10.1186/1471-2105-9-S8-S3
Kraulis PJ (1991) MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J Appl Crystallogr 24:946–950
Merritt EA, Murphy ME (1994) Raster3D Version 2.0. A program for photorealistic molecular graphics. Acta Crystallogr D Biol Crystallogr 50:869–873. doi:10.1107/S0907444994006396
Ortiz AR, Strauss CE, Olmea O (2002) MAMMOTH (matching molecular models obtained from theory): an automated method for model comparison. Protein Sci 11:2606–2621. doi:10.1110/ps.0215902
Hartshorn MJ (2002) AstexViewer: a visualisation aid for structure-based drug design. J Comput Aided Mol Des 16:871–881. doi:10.1023/A:1023813504011
Finn RD et al (2008) The Pfam protein families database. Nucleic Acids Res 36:D281–D288. doi:10.1093/nar/gkm960
Pieper U et al (2008) MODBASE, a database of annotated comparative protein structure models and associated resources. Nucleic Acids Res. doi:10.1093/nar/gkn791
Kiefer F et al (2008) The SWISS-MODEL Repository and associated resources. Nucleic Acids Res. doi:10.1093/nar/gkn750
Acknowledgments
We are grateful to Rainer Pöhlmann ([BC]2 Basel Computational Biology Center & Biozentrum University of Basel) for professional systems support, Eric Jain for the swift implementation of md5 based REST queries on the UniProt server and Michael Künzli for the computation of the statistics of the PMP models. We would like to thank all teams at the different partner sites for the great collaboration on the PSI SGKB Protein Model Portal integration, especially Wendy Tao (RCSB-PDB), Andrej Sali and Ursula Pieper (UCSF/NYSGXRC), Christine Orengo and David Lee (MCSG and UCL), Adam Godzik (JCMM), and Diana Murray (NESG). We are indebted to Roland Dunbrack Jr. (FCCC/NMHRCM) for invaluable advice on the development of PMP. The PSI SGKB Protein Model Portal was supported by the National Institutes of Health NIH as a sub-grant with Fox Chase Cancer Center grant 3 P20 GM076222-02S1, and as a sub-grant with Rutgers University, under Prime Agreement Award Number: 3U54GM074958-04S2.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Arnold, K., Kiefer, F., Kopp, J. et al. The Protein Model Portal. J Struct Funct Genomics 10, 1–8 (2009). https://doi.org/10.1007/s10969-008-9048-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10969-008-9048-5