Abstract
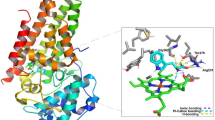
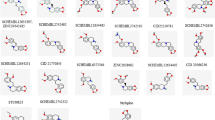
AKR1B1 and AKR1B10 are important members of aldo–keto reductase family which plays a significant role in cancer progression by modulating cellular metabolism. These enzymes are involved in various metabolic processes, including the synthesis and metabolism of hormones, detoxification of reactive aldehydes, and the reduction of various endogenous and exogenous compounds. This study aimed to explore the potential of strychnine as an anticancer agent by targeting AKR1B1 and AKR1B10 via drug repurposing approach. To assess the drug-like properties of strychnine, a physiologically based pharmacokinetic (PKPB) model and High Throughput Pharmacokinetics (HTPK) approach were employed. The obtained results fell within the expected range for drug molecules, confirming its suitability for further investigation. Additionally, density functional theory (DFT) studies were conducted to gain insight into the electronic properties contributing to the drug molecule’s reactivity. Building upon the promising DFT results, molecular docking analysis using the AutoDock tool was performed to examine the binding interactions between strychnine and the proposed targets, AKR1B1 and AKR1B10. Findings from the molecular docking studies suggested a higher probability of strychnine acting as an inhibitor of AKR1B1 and AKR1B10 with docking scores of − 30.84 and − 29.36 kJ/mol respectively. To validate the stability of the protein–ligand complex, Molecular Dynamic Simulation (MDS) studies were conducted, revealing the formation of a stable complex between the enzymes and strychnine. This comprehensive approach sheds light on the potential effectiveness of strychnine as a treatment for breast, lung, liver, and pancreatic cancers, as well as related malignancies. The novel insights gained from the physiologically based pharmacokinetic modeling, density functional theory, molecular docking, and molecular dynamics simulations collectively support the prospect of strychnine as a promising molecule for anticancer therapy. Further investigations are warranted to validate these findings and explore the therapeutic potential of strychnine in preclinical and clinical settings.




















Similar content being viewed by others
Availability of Data and Materials
Data will be available on request by the corresponding author.
References
Juliusson G, Lazarevic V, Hörstedt AS, Hagberg O, Höglund M (2012) Acute myeloid leukemia in the real world: why population-based registries are needed. Blood J Am Soc Hematol 119(17):3890–3899
Hussen BM, Hidayat HJ, Salihi A, Sabir DK, Taheri M, Ghafouri-Fard S (2021) MicroRNA: a signature for cancer progression. Biomed Pharmacother 138:111528
Penning TM (2015) The aldo-keto reductases (AKRs): overview. Chem Biol Interact 234:236–246
Giménez-Dejoz J, Weber S, Fernández-Pardo Á, Möller G, Adamski J, Porté S, Parés X, Farrés J (2019) Engineering aldo-keto reductase 1B10 to mimic the distinct 1B15 topology and specificity towards inhibitors and substrates, including retinoids and steroids. Chem Biol Interact 307:186–194
Munkácsy G, Santarpia L, Győrffy B (2023) Therapeutic potential of tumor metabolic reprogramming in triple-negative breast cancer. Int J Mol Sci 24(8):6945
Sun T, Liu Z, Yang Q (2020) The role of ubiquitination and deubiquitination in cancer metabolism. Mol Cancer 19:1–19
Taskoparan B, Seza EG, Demirkol S, Tuncer S, Stefek M, Gure AO, Banerjee S (2017) Opposing roles of the aldo-keto reductases AKR1B1 and AKR1B10 in colorectal cancer. Cell Oncol 40:563–578
Kropotova ES, Tychko RA, Zinov’eva OL, Zyryanova AF, Khankin SL, Cherkes VL, Aliev VA, Beresten SF, Oparina NY, Mashkova TD (2010) Downregulation of AKR1B10 expression in colorectal cancer. Mol Biol 44:216–222
Chen B, Garmire L, Calvisi DF, Chua MS, Kelley RK, Chen X (2020) Harnessing big ‘omics’ data and AI for drug discovery in hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol 17(4):238–251
Sravanthi TV, Manju SL (2016) Indoles—a promising scaffold for drug development. Eur J Pharm Sci 91:1–10
Sharma V, Kumar P, Pathak D (2010) Biological importance of the indole nucleus in recent years: a comprehensive review. J Heterocycl Chem 47(3):491–502
Lauria A, Delisi R, Mingoia F, Terenzi A, Martorana A, Barone G, Almerico AM (2014) 1, 2, 3-Triazole in heterocyclic compounds, endowed with biological activity, through 1, 3-dipolar cycloadditions. Eur J Org Chem 2014(16):3289–3306
Saraswati S, Mathur R, Agrawal SS (2010) 653 Evaluation of strychnine, a plant alkaloid for in vitro antiangiogenesis, apoptosis and antioxidant potential in MCF-7 cancer cells. EJC Suppl 7(8):204
Song Y, Yang J, Yu J, Li J, Yuan J, Wong NK, Ju J (2020) Chlorinated bis-indole alkaloids from deep-sea derived Streptomyces sp. SCSIO 11791 with antibacterial and cytotoxic activities. J Antibiot 73(8):542–547
Ruiz FX, Cousido-Siah A, Mitschler A, Farres J, Pares X, Podjarny A (2013) X-ray structure of the V301L aldo–keto reductase 1B10 complexed with NADP+ and the potent aldose reductase inhibitor fidarestat: Implications for inhibitor binding and selectivity. Chem Biol Interact 202(1–3):178–185
Zhang L, Zhang H, Zhao Y, Li Z, Chen S, Zhai J, Chen Y, Xie W, Wang Z, Li Q, Zheng X (2013) Inhibitor selectivity between aldo–keto reductase superfamily members AKR1B10 and AKR1B1: role of Trp112 (Trp111). FEBS Lett 587(22):3681–3686
Huang L, He R, Luo W, Zhu YS, Li J, Tan T, Zhang X, Hu Z, Luo D (2016) Aldo-keto reductase family 1 member B10 inhibitors: potential drugs for cancer treatment. Recent Pat Anti-Cancer Drug Discovery 11(2):184–196
Foulani AAE, Hammoudan I, Byoud F, Jamal-eddine J, Lekhlif B (2022) Synthesis, characterization, and evaluation of new composites coagulants polyaluminum chloride-sodium alginate. Water Air Soil Pollut 233(8):301
Gaussian RA (2009) 1, mj frisch, gw trucks, hb schlegel, ge scuseria, ma robb, jr cheeseman, g. Scalmani, v. Barone, b. Mennucci, ga petersson et al., gaussian. Inc Wallingford CT 121:150–166
Chem 3D pro 12.0 (Copyright) 1986 to 2009 by CambridgeSoft Corp. [Cambridge, Mass., U.S.A.]
Castellví A, Crespo I, Crosas E, Cámara-Artigas A, Gavira JA, Aranda MA, Parés X, Farrés J, Juanhuix J (2019) Efficacy of aldose reductase inhibitors is affected by oxidative stress induced under X-ray irradiation. Sci Rep 9(1):3177
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791
Khan SU, Ahemad N, Chuah LH, Naidu R and Htar TT (2020) Illustrated step by step protocol to perform molecular docking: human estrogen receptor complex with 4-hydroxytamoxifen as a case study. Progress Drug Discovery Biomed Sci 3(1):a0000054–a0000081
Visualizer DS (2005) Accelrys software Inc. Discovery Studio Visualizer
Wang Z, Sun H, Yao X, Li D, Xu L, Li Y, Tian S, Hou T (2016) Comprehensive evaluation of ten docking programs on a diverse set of protein–ligand complexes: the prediction accuracy of sampling power and scoring power. Phys Chem Chem Phys 18(18):12964–12975
Channar PA, Aziz M, Ejaz SA, Chaudhry GES, Saeed A, Ujan R, Hasan A, Ejaz SR, Saeed A (2023) Structural and functional insight into thiazolidinone derivatives as novel candidates for anticancer drug design: in vitro biological and in-silico strategies. J Biomol Struct Dyn 41(3):942–953
Bowers KJ, Chow E, Xu H, Dror RO, Eastwood MP, Gregersen BA, Klepeis JL, Kolossvary I, Moraes MA, Sacerdoti FD and Salmon JK (2006) November. Scalable algorithms for molecular dynamics simulations on commodity clusters. In: Proceedings of the 2006 ACM/IEEE Conference on Supercomputing. pp 84-es
Ferreira LG, Dos Santos RN, Oliva G, Andricopulo AD (2015) Molecular docking and structure-based drug design strategies. Molecules 20(7):13384–13421
Aziz M, Ejaz SA, Rehman HM, Alsubaie AS, Mahmoud KH, Siddique F, Al-Buriahi MS, Alrowaili ZA (2023) Identification of NEK7 inhibitors: structure based virtual screening, molecular docking, density functional theory calculations and molecular dynamics simulations. J Biomol Struct Dyn 41(14):6894–6908
Aziz M, Ejaz SA, Tamam N, Siddique F, Riaz N, Qais FA, Chtita S, Iqbal J (2022) Identification of potent inhibitors of NEK7 protein using a comprehensive computational approach. Sci Rep 12(1):6404
Ejaz SA, Alsfouk AA, Batiha GES, Aborode AT, Ejaz SR, Umar HI, Aziz M, Saeed A, Mahmood HMK, Fayyaz A (2023) Identification of N-(4-acetyl-4, 5-dihydro-5-(7, 8, 9-substituted-tetrazolo [1, 5-a]-quinolin-4-yl)-1, 3, 4-thiadiazol-2-yl) acetamide derivatives as potential caspase-3 inhibitors via detailed computational investigations. Struct Chem 34(2):425–438
Funding
The research conducted in this study was financially supported by Princess Nourah bint Abdulrahman University through Project Number PNURSP2023R142, Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. The funding agency significantly contributed to several aspects of the investigation, including study design, procurement, sample characterization, data processing, interpretation, and paper preparation. The authors are grateful and acknowledge the Simulations Plus, Inc., Lancaster, California for providing the software to Al Ain University, United Arab Emirates to predict ADMET and PK properties.
Author information
Authors and Affiliations
Contributions
MS: methodology, investigations. MA: methodology, experimental material design, Investigations. SA: methodology, investigations. PAC: methodology, experimental material design, investigations. BAA: funding, characterization, resources, validation, visualization, writing—original draft. GAK: investigations, writing—review and editing. SH: methodology, formal analysis. AS: funding, characterization, resources, validation, visualization, writing-original draft. AH: investigation, writing—review and editing. MA (Mosab Arafat): investigation, writing—review and editing. MNQ: methodology, formal analysis. AA: methodology, experimental material design, investigations. FS: methodology, experimental material design, investigations. SAE: conceptualization, methodology, supervision, investigation, writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of Interest
There is no any financial or personal competing interest among authors.
Ethical Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sarfraz, M., Aziz, M., Afzal, S. et al. Repurposing of Strychnine as the Potential Inhibitors of Aldo–keto Reductase Family 1 Members B1 and B10: Computational Modeling and Pharmacokinetic Analysis. Protein J (2023). https://doi.org/10.1007/s10930-023-10163-z
Accepted:
Published:
DOI: https://doi.org/10.1007/s10930-023-10163-z