Abstract
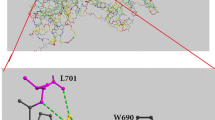
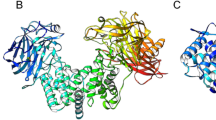
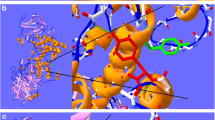
Chondroitinase ABC I (cABC I) from Proteus vulgaris is an important enzyme in medicinal biotechnology due to its ability to help axon regeneration after spinal cord injury. Its practical application involves solving several problems at the molecular and cellular levels. Structurally, most residues at the C-terminal domain of cABC I are arranged as organized strands, and only a small fraction of residues have helical conformation. The structural and functional features of modified residues on two specific helix fragments have previously been reported. The single mutant M889K has been combined with L679S and L679D mutants to make enzyme variants containing simultaneously modified helix. Here, the pH stability and temperature-based analysis of the transition state structure for the catalysis reaction were investigated. We found that double mutant L679D/M889K is the better choice to use in physiological conditions due to its higher pH stability at physiological pH as well as its different optimum temperature as compared with the (wild-type) WT protein. According to Arrhenius’s analysis, the values of the Gibbs free energy of the transition state (∆G#) are not changed upon mutation. However, the relative contribution and absolute values of the enthalpy and entropy change to the total value of ∆G#, varied between the WT and mutants.





Similar content being viewed by others
References
Ward OP, Moo-Young M (1988) Thermostable enzymes. Biotechnol Adv 6:39–69. https://doi.org/10.1016/0734-9750(88)90573-3
Basheer SM, Chellappan S (2017) Enzyme engineering. In: Sugathan S, Pradeep NAS (eds) Bioresources and bioprocess in biotechnology. Springer, Singapore, pp 151–158
Niehaus F, Niehaus F, Bertoldo C et al (1999) Extremophiles as a source of novel enzymes for industrial application. Appl Microbiol Biotechnol 51:711–729. https://doi.org/10.1007/s002530051456
Platis D, Labrou N (2008) Chemical and genetic engineering strategies to improve the potency of pharmaceutical proteins and enzymes. Curr Med Chem 15:1940–1955. https://doi.org/10.2174/092986708785132924
Yuan YM, He C (2013) The glial scar in spinal cord injury and repair. Neurosci Bull 29:421–435. https://doi.org/10.1007/s12264-013-1358-3
Wilhelmsson U, Bushong EA, Price DL et al (2006) Redefining the concept of reactive astrocytes as cells that remain within their unique domains upon reaction to injury. Proc Natl Acad Sci USA 103:17513–17518. https://doi.org/10.1073/pnas.0602841103
F M, A O, (2004) Proteoglycans and injury of the central nervous system. Congenit Anom 44:181–186
Rhodes KE, Fawcett JW (2004) Chondroitin sulphate proteoglycans: Preventing plasticity or protecting the CNS? J Anat 204:33–48. https://doi.org/10.1111/j.1469-7580.2004.00261.x
Silver J, Miller JH (2004) Regeneration beyond the glial scar. Nat Rev Neurosci 5:146–156. https://doi.org/10.1038/nrn1326
Shirdel A, Khalifeh K (2022) Chondroitinase ABC I as a novel candidate for reducing damage in spinal cord injury. In: Rajendram R, Preedy V, Martin C (eds) Diagnosis and treatment of spinal cord injury. Academic Press, pp 325–335
Prabhakar V, Capila I, Bosques CJ et al (2005) Chondroitinase ABC I from Proteus vulgaris: cloning, recombinant expression and active site identification. Biochem J 386:103–112. https://doi.org/10.1042/BJ20041222
Wang H, Zhang L, Wang Y et al (2021) Engineering a thermostable chondroitinase for production of specifically distributed low-molecular-weight chondroitin sulfate. Biotechnol J. https://doi.org/10.1002/biot.202000321
Cantarel BI, Coutinho PM, Rancurel C et al (2009) The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37:233–238. https://doi.org/10.1093/nar/gkn663
Huang W, Lunin VV, Li Y et al (2003) Crystal structure of Proteus vulgaris chondroitin sulfate ABC lyase I at 1.9 Å resolution. J Mol Biol 328:623–634. https://doi.org/10.1016/S0022-2836(03)00345-0
Hettiaratchi MH, O’Meara MJ, Teal CJ et al (2019) Local delivery of stabilized chondroitinase ABC degrades chondroitin sulfate proteoglycans in stroke-injured rat brains. J Control Release 297:14–25. https://doi.org/10.1016/j.jconrel.2019.01.033
García-Alías G, Lin R, Akrimi SF et al (2008) Therapeutic time window for the application of chondroitinase ABC after spinal cord injury. Exp Neurol 210:331–338. https://doi.org/10.1016/j.expneurol.2007.11.002
Barritt AW, Davies M, Marchand F et al (2006) Chondroitinase ABC promotes sprouting of intact and injured spinal systems after spinal cord injury. J Neurosci 26:10856–10867. https://doi.org/10.1523/JNEUROSCI.2980-06.2006
Zhao RR, Fawcett JW (2013) Combination treatment with chondroitinase ABC in spinal cord injury—breaking the barrier. Neurosci Bull 29:477–483. https://doi.org/10.1007/s12264-013-1359-2
Hu HZ, Granger N, Balakrishna Pai S et al (2018) Therapeutic efficacy of microtube-embedded chondroitinase ABC in a canine clinical model of spinal cord injury. Brain 141:1017–1027. https://doi.org/10.1093/brain/awy007
Tester NJ, Plaas AH, Howland DR (2007) Effect of body temperature on chondroitinase ABC’s ability to cleave chondroitin sulfate glycosaminoglycans. J Neurosci Res 85:1110–1118. https://doi.org/10.1002/jnr.21199
Jamshidi N, Shirdel A, Pourahmadi M et al (2020) Bioinformatics and experimental studies on the structural roles of a surface-exposed α-helix at the C-terminal domain of Chondroitinase ABC I. Int J Biol Macromol 163:1572–1578. https://doi.org/10.1016/j.ijbiomac.2020.07.165
Chen Z, Li Y, Feng Y et al (2015) Enzyme activity enhancement of chondroitinase ABC I from Proteus vulgaris by site-directed mutagenesis. RSC Adv 5:76040–76047. https://doi.org/10.1039/c5ra15220h
Naderi MS, Moghadam TT, Khajeh K, Ranjbar B (2018) Improving the stability of chondroitinase ABC I via interaction with gold nanorods. Int J Biol Macromol 107:297–304. https://doi.org/10.1016/j.ijbiomac.2017.08.167
Shirdel SA, Khalifeh K, Golestani A et al (2015) Critical role of a loop at C-terminal domain on the conformational stability and catalytic efficiency of chondroitinase ABC I. Mol Biotechnol 57:727–734. https://doi.org/10.1007/s12033-015-9864-3
Shamsi M, Akram Shirdel S, Jafarian V et al (2016) Optimization of conformational stability and catalytic efficiency in chondroitinase ABC Ι by protein engineering methods. Eng Life Sci 16:690–696. https://doi.org/10.1002/elsc.201600034
Moradi K, Shirdel SASA, Shamsi M et al (2017) Investigating the structural and functional features of representative recombinants of chondroitinase ABC I. Enzyme Microb Technol 107:64–71. https://doi.org/10.1016/j.enzmictec.2017.08.006
Mohammadyari H, Shirdel SA, Jafarian V, Khalifeh K (2019) Designing and construction of novel variants of Chondroitinase ABC I to reduce aggregation rate. Arch Biochem Biophys 668:46–53. https://doi.org/10.1016/j.abb.2019.05.013
Shen M-Y, Sali A (2006) Statistical potential for assessment and prediction of protein structures. Protein Sci 15:2507–2524. https://doi.org/10.1110/ps.062416606
Fisher CL, Pei GK (1997) Modification of a PCR-based site directed mutagenesis method. Biotechniques 23:570–574
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Eyring H (1935) The activated complex and the absolute rate of chemical reactions. Chem Rev 17:65–77. https://doi.org/10.1021/cr60056a006
Hettiaratchi MH, O’Meara MJ, O’Meara TR et al (2020) Reengineering biocatalysts: computational redesign of chondroitinase ABC improves efficacy and stability. Sci Adv 6:1–9. https://doi.org/10.1126/sciadv.abc6378
Rypniewski WR, Perrakis A, Vorgias CE, Wilson KS (1994) Evolutionary divergence and conservation of trypsin. Protein Eng Des Sel 7:57–64. https://doi.org/10.1093/protein/7.1.57
Acknowledgements
We appreciate the Tarbiat Modares University of Tehran for the technical support of the research. The authors have declared no conflict of interest. Dr. Khosro Khajeh and Dr. Abolfazl Golestani are acknowledged for their generous donation of bacterium.
Funding
This work was supported by the research council of the University of Zanjan.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that influence the data of the current work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10930_2023_10093_MOESM1_ESM.tif
Supplementary file1 (TIF 90 KB)Supplementary image 1. SDS-PAGE analysis of purified recombinant proteins. Lane 1, marker; lane2, WT; lane3, M889K; lane 4, M889L; lane 5, L679D/M889K; lane 6, L679S/M889K. The numbers provided on the left side of the image are the molecular weight of marker proteins in kDa.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Moradi, K., Bayani, Z., Jafarian, V. et al. Enzyme Kinetics Features of the Representative Engineered Recombinants of Chondroitinase ABC I. Protein J 42, 55–63 (2023). https://doi.org/10.1007/s10930-023-10093-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-023-10093-w