Abstract
The newly emerging SARS-CoV-2 variants are potential threat and posing new challenges for medical intervention due to high transmissibility and escaping neutralizing antibody (NAb) responses. Many of these variants have mutations in the receptor binding domain (RBD) of SARS-CoV-2 spike protein that interacts with the host cell receptor. Rapid mutation in the RBD through natural selection to improve affinity for host receptor and antibody pressure from vaccinated or infected individual will greatly impact the presently adopted strategies for developing interventions. Understanding the nature of mutations and how they impact the biophysical, biochemical and immunological properties of the RBD will help immensely to improve the intervention strategies. To understand the impact of mutation on the protease sensitivity, thermal stability, affinity for the receptor and immune response, we prepared several mutants of soluble RBD that belong to the variants of concern (VoCs) and interest (VoIs) and characterize them. Our results show that the mutations do not impact the overall structure of the RBD. However, the mutants showed increase in the thermal melting point, few mutants were more sensitive to protease degradation, most of them have enhanced affinity for ACE2 and some of them induced better immune response compared to the parental RBD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The emergence of SARS-CoV-2 variants of concerns (VoCs) and interests (VoIs) has become a significant risk to human life and economies due to high mortality and implemented lockdowns to curb the spread of infection [1]. SARS-CoV-2 causes lower respiratory tract infection that can progress to severe acute respiratory syndrome and multiple organ failure [2]. As of now, almost every country in the world is reeling under the threat of COVID-19, with more than 6 million casualties globally. In the absence of standard medical intervention for COVID-19, current clinical management comprises mainly of supportive care [3, 4]. Several vaccines with good reported efficacy and antibody therapeutics are now available for use while others are in the different stages of development and clinical trials [5,6,7].
The genome of SARS-CoV-2 is a single positive stranded RNA that encodes four structural proteins, the spike (S), the envelope (E), the matrix (M) and the nucleocapsid (N) proteins [8, 9]. The S protein is a trimeric glycoprotein on the virion surface and is a type I fusion protein. It is composed of two subunits, the S1 subunit is responsible for binding to the host receptor and the S2 subunit facilitates membrane fusion [10]. The receptor binding domain (RBD) is located in the S1 subunit and binds with the angiotensin converting enzyme 2 (ACE2) present on the host cell surface [11]. Being a RNA virus it shows rapid mutation that is driven by the low fidelity RNA-dependent RNA polymerase (RdRp) [12, 13]. The continued existence of the mutants depends on the selective advantages during natural infection, transmission, and host adaptation [14, 15]. Mutations in the spike protein of SARS-CoV and MERS-CoV have played a significant evolutionary role in the transmission of these viruses from animal to human through an improved binding to the human host receptor [16,17,18]. It is also evident that the mutations in the spike protein and RBD in particular, influence the infectivity and transmissibility of the virus [19]. Even for SARS-CoV-2, the mutants’ showed more severe pathogenesis and potential to evade antibody therapy [20,21,22]. Also, the spike protein is the major antigen for the host immune system to augment its defense by inducing both the humoral and cellular arm. Naturally, most of the vaccine development efforts are centered around the S protein of SARS-CoV-2 [23, 24]. However, the emergence of mutations in the S protein and frequently in the RBD can significantly alter receptor recognition, viral attachment, host cell entry and interactions with NAbs as well as may facilitate escape from current antibody therapy and vaccine strategies [19,20,21].
Since the first identification of the paternal strain of SARS-CoV-2 isolate, Wuhan-Hu-1, the virus has undergone rapid mutation in its genome [25,26,27]. Some of these mutants have been identified as the ‘Variant of Interest’ (VoI) or ‘Variant of Concern’ (VoC) based on their infectivity, transmissibility, rapid spread, disease pathogenesis, death rate etc. [28]. The virus is still mutating and it is expected that many more VoIs and VoCs will emerge with time. The RBD of spike protein plays a key role in the virus transmissibility and antigenicity and a major target for vaccine development. Here, we investigated whether the mutations in the RBD have altered their stability, changes in secondary structure and impact on receptor binding affinity. We, purified several RBD mutants along with the parental Wuhan strain, henceforth wild-type (WT), in mammalian expression system to homogeneity and performed their binding studies with human ACE2 and characterization of structural stabilities. Our results suggest that the RBD variants are monomeric, there is no significant impact on the overall secondary structure, some of the variants found to be more susceptible to proteolysis, mostly the single mutation showed increased affinity for ACE2 but interestingly, all the variants exhibited higher stability to melting temperature. The mutants induced robust immune response in mice and the induced sera were able to cross neutralize the VoCs tested.
2 Materials and Methods
2.1 Construction of Mutants
The mutations were introduced in the backbone of the Wuhan strain of RBD (NCBI ACCESSION ID: QHD43416; amino acids 328 to 531) from BEI (Catalog No. NR-52422; BEI Resources; NIH) via site directed mutagenesis PCR using QuikChange Lightning Multi Site-Directed Mutagenesis kit (Agilent Technologies). All of the constructs were designed with an N-terminal signal peptide for efficient extracellular expression in mammalian expression system [29] and a C-terminal poly-His-Tag (8xHis) for Ni-NTA based affinity purification. The details of the mutagenic primers used, the DNA and the amino acid sequence of the mutants are provided in the supplementary information. In brief, mutagenic primers were designed to mutate the targeted residues. The entire plasmid was amplified using the mutagenic primers and the PCR reaction products were transformed into competent bacterial cells and plated onto Luria Broth agar plates for colony selection, followed by plasmid DNA isolation, and sequencing to confirm the insertion of the desired mutation(s).
2.2 Protein Expression and Purification
The RBD mutants and the WT-RBD were expressed in transiently transfected Expi293F cells in suspension culture using Expi293fectin (Invitrogen) following manufactures protocol. Cell culture supernatants were harvested 5–6 days after transfection. For protein purification, the supernatant was passed through a Ni–NTA column. Bound proteins were eluted with 500 mM imidazole and further purified by size exclusion chromatography (SEC) using Superdex 200 increase (10/300) column (GE Biosciences). Eluted proteins were concentrated with an Amicon filter (Millipore; 10 kDa), protein fractions were aliquoted and stored at − 80 °C until further use.
2.3 Circular Dichroism (CD) Spectroscopy
Far-UV CD spectra were acquired on a Jasco-815 spectropolarimeter. The concentration of the protein used in all cases is 5 µM. Cuvette of path length of 0.2 cm was used and spectra were collected from 260 to 190 nm at a rate of 100 nm/min and data pitch of 1 nm for each protein, with averaging of 10 scans for noise reduction. Contributions of the buffer to the spectra were electronically subtracted and for each spectrum, mean residual ellipticity (MRE) was calculated and plotted.
2.4 Protease Sensitivity
The WT-RBD and the variants (50 µg of each) were treated with 1.5 µg of trypsin (T-1426 Type XIII from Bovine Pancreas – Sigma Aldrich) in a final reaction volume of 100 µl. The reaction was incubated at room temperature. At intervals of 2H and 4H, 10 µl of fractions of each protein were collected and subsequently mixed with 4 × loading dye and denatured at 95 °C for 5 min. The trypsinized samples were separated on a 15% Bis–Tris SDS polyacrylamide gel.
2.5 Differential Scanning Calorimetry
Thermal melting of the RBD mutants was analyzed with a Nano-DSC (TA instruments). Samples were dialyzed in PBS buffer and protein concentrations were adjusted between 0.5 and 0.7 mg/ml prior to measurement. Thermal melting was performed at a scanning rate of 1 °C/min under 3.0 atmospheres of pressure. Data collected were analyzed with NanoAnalyze software, 3.6.0 (TA instruments), after buffer correction, normalization, and baseline subtraction [30, 31].
2.6 Bio-layer Interferometry (BLI)
For binding kinetics anti-human Fc sensors (ForteBio Inc.) was used to capture the ACE2-Fc on the sensor and the RBD mutants were used as analyte in concentrations ranging from 200 to 25 nM with 1/2 serial dilution along with a no analyte as background control. The PBS buffer background supplemented with 0.01% Tween 20 and 0.1% BSA. The experiment was performed at room temperature with agitation at 1000 rpm. To capture the ACE2-Fc, the biosensors were immersed in wells containing ACE2-Fc at a concentration of 10 µg/ml for 120 s. Association was recorded for 120 s followed by dissociation for 150 s. After running the assay, the data were open in the Data Analysis Software, 10.0 (Forte-Bio Inc) and the data were loaded for individual run. The sensor that was run without any analyte was designated as reference well and values were electronically subtracted from other sensor with analyte as background. At the beginning of the association curve all the data points were aligned at baseline using the data analysis software programming followed by inter-step correction to minimize signal shifts between the association and dissociation steps. Next, Savitzky-Golay filtering function was used to process the data. The kinetic parameters were calculated using the processed data set for each mutant. Since one molecule of ACE2 interacts with one molecule of RBD, 1:1 global fit model strategy was used to fit the dataset for all of the concentrations of the analyte used and the fit curve option was used for nonlinear regression fitting of the data points. The final fit data were analyzed which include overlay of fit curve along with the raw sensor curve, plots of fitting residuals, a table with determined parameter values (kon, koff, KD) and statistics (standard errors for parameters, Chi-squared, R2). The raw data and the fit data were plotted for each mutant as shown in Fig. 4. The mean values of KD, kon and koff from two independent experiments with standard deviation is tabulated in Table 2 [30,31,32].
2.7 Mice Immunization
For immunization study, 7–8 weeks old female BALB/c mice weighing 18–25 gm and inbred in THSTI small animal facility (SAF) were used after approval from the Institutional Animal Ethics Committee (IAEC). Three mice for each of WT-RBD and the mutants L452R, T478K, Delta and DeltaPlus, were immunized with 25 ug of antigen mixed with AddaVax as adjuvant in 1:1 ration. Immunization was performed via intramuscular route (cranial thigh muscles) thrice (prime, first boost and second boost) at interval of three weeks. Blood sample from each mouse was collected at day 0 (pre immune sera), 14 (sera after priming), 35 (sera after the first boost), and at 56 (sera after the second boost). Serum was separated from the blood, heat inactivated at 56 °C for 1 h, and stored at − 20 °C for future use.
2.8 Enzyme-Linked Immunosorbent Assay (ELISA)
The ELISA was performed to characterize the binding of the immunized sera with the WT and the RBD variants. The WT-RBD and the mutant of RBDs were coated on Maxisorp plates (Nunc) with 100 μl of protein at 2 μg/ml concentration in 1 × carbonate/bicarbonate buffer, pH 9.6 for overnight at 4 °C. Next day, the plate was blocked by 250 μl of PBS containing 5% skimmed milk. Pooled immunized sera from different group of mice at the dilution of 1:2700 folds was added to the wells and incubated for 2 h at RT. The wells were washed and HRP-conjugated anti mouse secondary antibody (Jackson Immuno Research, USA) in 1:2000 dilutions were used for developing ELISA.
2.9 Plaque Reduction Neutralization Test (pRNT) Assay
SARS-CoV-2, WT isolate and VoCs were used to perform pRNT assay at THSTI Infectious Disease Research Facility (Biosafety level 3 facility). 0.4 million of Vero E6 cells in 1 ml of DMEM (Gibco) containing 10% FBS (Gibco) and 100U/ml P/S (Gibco) were seeded in a 12-well plate. Next day, heat inactivated pooled serum sample collected three weeks after 2nd boost was diluted to 40-fold in 100ul of DMEM supplemented with 2% FBS and incubated with 100ul of 50 PFU of SARS-CoV-2 for 1 h. at 37 °C. Following incubation, confluent Vero E6 cells in 12 well plates were infected with the sera-virus mixture for 1 h at 37 °C. After 1 h of adsorption, the wells were washed one time with PBS. One ml of DMEM containing 2% FBS, 1 × P/S and 0.5% carboxy methyl cellulose (CMC) was added to each well and the plate was incubated at 37 °C with 5% CO2. After 48 h., media from each well were removed and the cells were fixed with 4% PFA for 1 h. Cells were then stained with 0.5% crystal violet for half an hour and washed with water. The plates were air dried and the numbers of plaques in respective wells were counted.
3 Results
3.1 Expression and Purification of RBD Variants
The WT-RBD used in this study is of 210 amino acids ranging from residue 326 to 536 and has two N-glycosylation sites at N331 and N343 [33]. The RBD mutants’ characterized and compared with the WT are single mutants; (N440K, L452R, E484Q, N501Y and T478), double mutants; Kappa (L452R/E484Q), Gamma (E484K/N501Y) and Delta (L452R/T478K); and triple mutants; Beta (K417N/E484K/N501Y) and DeltaPlus (K417N/L452R/T478K) (Table 1; Fig. 1A). The mutated residues and their location in perspective to ACE2 binding interface are shown in Fig. 1B. All the RBD proteins were expressed transiently in Expi293F mammalian cell culture and purified to homogeneity using Ni–NTA affinity column chromatography followed by SEC. The purity and homogeneity of the purification was confirmed by SDS-PAGE analysis. The calculated molecular weight of WT-RBD and the mutants is approximately 24 kDa. However the full-length RBD migrates on the SDS-PAGE as about 29 kDa band due to the presence of two N-glycosylation sites at N331 and N343 (Fig. 2A) [34]. Analytical SEC result shows that all the RBD mutants are homogeneous and monomer with elution peak at around 17.0 ml from Superdex 200 increase (10/300) column (Fig. 2B). The result suggests that the mutations did not cause any significant change in the oligomeric status and the hydrodynamic properties of the variants.
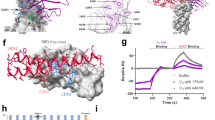
A The different variants of the RBD that were characterized are shown with the location of the residues mutated as green sphere. The mutants include either single, double or triple mutations. B Cartoon representation of the structure of RBD (blue and red) bound to human ACE2 (orange). The red part of the RBD is the receptor binding motif (RBM). The locations of the residues that are mutated are shown in green colour. All the residues mutated (N440, L452, T478, E484 and N501) are part of the RBM of RBD except residue K417 shown in cyan. C A close-up view of the RBD-ACE2 interface with the mutated residues shown as sphere. All the images were prepared using receptor binding domain of Wuhan strain of SARS-CoV-2 (PDB ID: 6M0J) and pymol
A SDS-PAGE analyses of the purified RBD WT and variants showed a well purified band at 29 kDa. B Size exclusion chromatography (SEC) normalized elution profile of the WT and variants of RBD. Elution peak varied from 16.7 ml to 17.5 ml suggesting monomeric form of the RBD and negligible difference in the hydrodynamic volume among the variants. C. Circular dichroism (CD) spectroscopy based analysis of conformational changes in the secondary structure content of the RBD variants compared to the WT. No detectable difference was noticed in any of the variants
3.2 Effect of Mutations on the Secondary Structure of RBD Variants
The RBD protein is mostly rich in β-sheet with little α-helical structure and some random coil conformation lacking structural rigidity, particularly in and around the receptor binding motif (RBM). The mutations that are characterized in this study are located at different secondary structures. Residues K417, N440, N501 are part of α-helical region, residue L452 is located in a β-sheet while T478 and E484 are located in the random coil region. To see whether the single mutation or combination of mutations influences any structural changes in the RBD variants compared to the WT, we performed circular dichroism (CD) spectroscopy analysis to ascertain any changes in the secondary structural signature (Fig. 2C). The CD spectra of the WT-RBD and the mutants were characterized by single peak minima at 208 nm and a maximum at 230 nm, which are characteristic of typical β-sheet rich structures. All the spectra were superimposable suggesting no significant deviation in secondary structure among the mutants compared to the WT. Thus, the mutations did not significantly alter the core structure of the RBD, however, local perturbation of small structural changes cannot be ruled out.
3.3 Sensitivity to Proteolysis of RBD Variants
The intrinsic stability of protein subunit-based vaccine candidates can directly influence the availability of B-cell epitopes to induce conformation dependent neutralizing antibodies. Destabilization of a protein structure can lead to unfolding of the tertiary structure and thus loss of conformational epitopes [35,36,37]. A destabilized unfolded structure with exposed peptide bond is prone to proteolytic degradation by several host proteases, including furin, trypsin, cathepsins, transmembrane protease serine protease-2 (TMPRSS-2) or human airway trypsin-like protease [38, 39] leading to faster clearance from the circulation and unavailability of the antigen for sufficient time to induce antibody response [40, 41]. To determine the comparative sensitivity to degradation of the WT-RBD and the RBD variants, the proteins were treated with trypsin and fractions were collected at different time interval. Since trypsin cleaves peptide bond on the C-terminal side of lysine (K) and arginine (R) amino acids we expected that the N440K, L452R and T478K single mutant and all the RBD variants that bears at least one mutation for R or K will be more sensitive to trypsinization compared to WT-RBD [42]. However, we found that after 2H of digestion only the E484K bearing mutants (Gamma and Beta) were found more sensitive to trypsinization and after 4H all the mutants and WT-RBD were found degraded (Fig. 3A). The, N440 (α-helix) and L452 (β-sheet) residues are part of structured region in the RBD protein while T478 and E484 are located at unstructured random coil (Fig. 1A). The presence of secondary structure at N440 and at L452 is expected to protect exposure of the peptide bonds and proteolytic digestion as seen for L452R (L452R, Kappa, Delta and DeltaPlus) and N440K bearing mutants (Fig. 3A; upper panel). The T478 residue, though part of random coil, is followed by a proline (P) residue [42] and thus not susceptible to trypsinization as seen for T478K, Delta and DeltaPlus (Fig. 3A; upper panel). Thus, only E484K bearing mutants were found more sensitive to trypsinization as observed for Gamma and Beta. After 2 h of trypsin digestion both the Gamma and Beta mutants, bearing E484K mutation, showed one distinct band at ~ 24 kDa. Cleavage at E484K will lead to an N-terminal fragment of 157 amino acids with predicted molecular weight of ~ 18 kDa and a C-terminal fragment of 55 amino acids with predicted molecular weight of ~ 6 kDa. The molecular weights of the fragments were predicted using Expasy ProParam software (https://web.expasy.org/protparam/). However due to the presence of two N-glycosylation sites at N331 and N343 the N-terminal fragment was seen to migrate as 24 kDa instead of the predicted 18 kDa [34]. The C-terminal fragment being very small will migrate out of the SDS-PAGE and no band was visible. We have not characterized the single E484K mutant, but above observation indicate that this mutant will be highly susceptible to trypsin digestion.
A SDS-PAGE profiles of trypsin digestion after 2H (upper panel) and 4H (lower panel) after treatment. B Thermostability of the RBD variants as measured using differential scanning calorimetry (DSC) and the fit normalized curves are plotted. The different Tm of the variants is shown in the adjoining table
3.4 Thermal Stability of RBD Variants
To assess the melting point (Tm) of the mutants we performed differential scanning calorimetry (DSC). The result suggests that the mutants have 3–6 °C higher Tm than the WT (Fig. 3B). The Tm of WT-RBD was recorded at about 48 °C, which was the lowest of all the RBD variants. The highest Tm was recorded for the Gamma variant at 54 °C with the following mutations: E484K and N501Y. All of the variants containing the L452R mutation (Kappa and Delta variants) showed a significantly high Tm of around 53 °C. The increased thermostability of the mutants particularly those with arginine (R) and lysine (K) substitution may be attributed to the said mutations. Arginine (R) and lysine (K) residues are positively charged basic amino acids and mostly surface exposed where the play important roles in protein stability by forming ionic interactions and hydrogen bonds in the proteins as well as by interacting with water molecules [43, 44].
3.5 Affinity to Human ACE2 of RBD Variants
To understand how the mutations in the RBD impacts its affinity to ACE2 may reflect to the cause for the rapid transmissibility and higher infectivity of the VoIs and VoCs. A detailed evaluation of the mutations in the RBD reveals that most of these mutations are clustered in the RBM region, the binding interface of RBD with ACE2, and thus may impact the binding affinity. Additionally, since the host NAbs are mostly directed in and around the RBM, genomic alteration in this region can help the mutants to evade host NAb response [45, 46]. Accordingly, the mutation(s) that enhances the affinity of RBD for ACE2 and also helps to escape the host NAb responses are the best fit to evolve further. Here, we have characterized the binding affinity of different RBD mutants in comparison to the WT (Fig. 4; Table 2). This information will help to disseminate probable role of the key residues that influence the interaction of RBD with ACE2. Most of the variants namely, L452R, E484Q, N501Y, and Kappa, showed improved affinity to ACE2 as compared to the WT. However, mutants, N440K, T478K, Gamma, Beta, Delta and DeltaPlus showed slightly lower binding affinity to ACE2 relative to WT. The highest affinity was recorded for L452R with almost three times to the WT. The RBD variants with increased affinity for ACE2, showed more than two fold increases in affinity which is biologically very significant as an increase in the binding affinity for receptor will hugely impact viral infectivity and transmission (Fig. 4; Table 2). Some other reports have also documented the same observation related to the RBD mutation and increased affinity to ACE2 [47, 48]. Notably, not all the mutations increased the affinity of the RBD to ACE2, mutants such as N440K, T478K, Gamma, Beta, Delta and DeltaPlus showed reduced affinity for ACE2 (Fig. 4; Table 2). The mutants with reduced affinity to ACE2 is probably harbor mutations to escape NAbs present in the naturally infected or vaccinated population as reported for N440K [45]. The Delta variant showed 1.3 fold less affinity, interestingly, the DeltaPlus variant with an extra mutation, K417N, showed even lower affinity (2.3 fold less) with ACE2. The results suggest that the DeltaPlus variant has evolved with K417N mutation in the background of Delta variant to further escape antibody pressure and in the process has further lost its affinity for ACE2. Two major VoC’s, Beta (South Africa) and DeltaPlus (India) have K417N mutation. Mutations of K at 417 position either with neutral amino acids N or T disrupts a salt bridge with ACE2 and has reduced affinity for ACE2 [11, 46]. The DeltaPlus has three mutations K417N, L452R, and T478K. Of these, the K417N and T478K individually has low affinity for ACE2 while L452R enhances the affinity for ACE2 [46]. Taken together there are overall reduction in affinity of DeltaPlus variant for ACE2 compared to the WT. This data suggest that the K417 and T478K mutations are responsible for lower affinity for ACE2 but may be more efficient in evading NAb response [46].
Sensogram of two fold decreasing concentration (200 nM, 100 nM, 50 nM and 25 nM) of RBD variants with immobilized human ACE2-Fc on Fc capture sensor. The fit using 1:1 binding model is shown as black line. A WT, B N440K, C L452R, D E484Q, E N501Y, F T478K, G Kappa, H Gamma, I Beta, J Delta, K DeltaPlus. L The relative binding of different RBD variants compared to WT. Mutants N440K, T478K, Gamma, Beta, Delta and DeltaPlus showed slightly reduced affinity for ACE2 compared to the WT
3.6 Differential Immune Response and Neutralization by RBD-Variants Across VoCs
Mice were immunized with the WT-RBD, variants L452R, T478K, Delta (L452R and T478K) and DeltaPlus (K417N, L452R and T478K) to understand the immunological response by these variants compared to the WT-RBD and their cross binding potential. BALB/c mice were immunized and sera were collected three weeks after 2nd boost (see material and method section for details). Three mice were immunized with each variant and pooled serum for each variant was used for cross-binding study at 2700 × fold dilution. Our result shows that the sera induced by WT-RBD binds more efficiently to the variants tested except L452R and to itself (Fig. 5A). The Delta variant, with L452R and T478K double mutation, showed least immune response while T478K single mutant mounted best immune response among the variants tested. The L452R and DeltaPlus also induced good immune response. This suggests that the mutations present in the RBD have a potential role in inducing differential immune response.
Cross binding titre and cross neutralization potential of immunized sera. A Cross binding titre of immunized sera obtained from mice immunized with the WT-RBD and its variants against self and other variants of RBD. B Cross neutralization potential in percentage of the immunized sera obtained from mice immunized with the WT-RBD and its variants against WT and VoCs using live virus in pRNT assay
We also tested the cross-neutralization potential of the sera induced by the WT-RBD and the variants (L452R, T478K, Delta and DeltaPlus) against some of the VoC’s that resulted in the recent outbreak. The neutralization efficiency was tested for the Alpha (with N501Y mutation in the RBD), Kappa (with L452R and E484Q mutations in the RBD) and Delta (with L452R and T478K mutations in the RBD) VoCs using live viruses in pRNT assay. The WT-RBD sera showed least neutralization potential to itself or the VoC’s tested at the 1:40 sera dilution (Fig. 5B). The neutralization response from the variant sera showed about twofold more potent neutralization while the L452R mutant showed best neutralization across all the VoC’s tested. The result suggests that the variants not only induced better immune response but also showed better neutralization across the VoC’s. To be noted here we also see escape neutralization for the Delta variant (47% neutralization compared to 57% neutralization) when tested with sera of WT-RBD. Neutralization escape by sera from naturally infected or vaccinated individuals are widely reported particularly for Delta virus and the magnitude of neutralization escape is dependent on antibody titer, full vaccine coverage and duration of vaccine does [49]. The marginal escape of neutralization seen for the Delta variants against WT-RBD sera may be attributed to the sera collected only three weeks after 2nd booster does.
4 Discussion
Development of several vaccines and therapeutic antibodies has helped immensely to fight SARS-CoV-2, still the options in terms of medical intervention to treat or prevent COVID-19 is very limited. SARS-CoV-2 has high rate of genomic mutations and their ability to rapidly change their genome correlate with their emergence in novel hosts, enhanced virulence, escape natural and vaccine-induced immunity and evolve to circumvent disease resistance induced by the host [50]. Surveillance of biologically significant SARS-CoV-2 mutants is an important public health measure as the use of antiviral treatments and vaccines are likely to mount selection pressures that could alter viral evolutionary dynamics to favor resistant strains [51, 52]. Genome variability also add to the problem for development of universal vaccine or drugs that acts with equal efficacy on different variants and a challenging problem to the scientific community for the development of universal drugs and vaccines for all variants. Similar challenges have been reported in the case of Influenza and HIV due to their high genomic variability. Biological characterization of the viral mutants can provide precious insights in re-assessing viral drug resistance, immune escape and pathogenesis related mechanisms and are equally crucial for designing new vaccines, antiviral drugs and diagnostic assays.
Mutants of SARS-CoV-2 with high infectivity and the ability to evade NAb pressures are far more likely to evolve further into a dominant variant. Interestingly, some of these mutations arose independently in different geographical regions indicating convergent evolution and selective advantage of the acquired variations [53]. However, not only the binding affinity, but changes in other biophysical properties such as increased stability may also impact the rate of transmission of the virus. Understanding the impact of mutations on the biophysical and biochemical properties of RBD will help in selecting and designing improved RBD based subunit vaccine candidates. The protease digestion experiments showed that E484K bearing mutants are highly sensitive to trypsin as this mutation is located in the random coil region. The L452R bearing mutants (L452R, Kappa, Delta and DeltaPlus) are less sensitive to trypsinization, have high Tm and also induce robust antibody response. The L452 residue is located in the middle of a β-sheet present in the RBM [11]. This β-sheet is the only structured region within the RBM. It might be possible that the L452R mutation further stabilized the RBM to account for both increased affinity to ACE2 and stability.
Natural selection assist mutants with enhanced affinity for ACE2 at the same time antibody pressure from the host may mount additional pressure on viral survival that may lead to antibody escape mutations. The antibody escape mutants may sometime have lowered affinity for ACE2 as seen in case of N440K and T478K bearing mutants (N440K, T478K, Delta and DeltaPlus) [45]. However, during viral infection, actual binding of the virus particles to the host cells are further influenced not only by the biochemical environment but also due to the trimeric nature of the spike protein and the dynamic equilibrium of up and down positions of RBD. The reduced affinity thus may be compensated to some extent by avidity for infectivity and same time evades immunity. It is imperative to see how long the mutant such as N440K, T478K and DeltaPlus with reduced affinity to ACE2 but with potential to escape antibody pressure can survive in the circulation. The variants also induced better immune response and neutralization efficacy compared to the WT-RBD when tested in mice. The SARS-CoV-2 is spreading fast and the same time mutating significantly. Over time it may pose the challenges that we have with Influenza and HIV in terms of medical intervention such as vaccine and therapeutic antibody development [54,55,56]. Possibility of recurring SARS-CoV-2 vaccination, depending on the strain and endemic regions are already under discussion as in the case of seasonal Influenza vaccine. The reason behind such consideration is both antibody escape and faster waning of antibody from the circulating sera of already vaccinated or naturally infected individuals. RBD, particularly, the RBM region is the site of receptor binding and a potent site of immune response. Mutation(s) in this region impacts receptor binding as well as antibody escape. Our data clearly suggests that even single mutation have impact on stability, receptor binding and the induction of immune-response. Screening might be helpful in selection of vaccine candidate in the future when plethoras of mutants are available as in the case of HIV and Influenza.
In summary, our data demonstrate that the mutations have not caused any significant change in the overall structure of RBD mutants. However, variants with L452R, E484Q, N501Y bearing mutations in the RBD of VoIs and VoCs favor improved affinity for ACE2 and the E484K bearing mutants are more sensitive to proteolysis (Gamma and Beta). The L452R bearing mutant showed improved affinity and stability of all the variants characterized, induced better immune response and has better coverage of neutralizing antibodies. This indicates that some mutations are quite favorable for high affinity to ACE2, stability and induces strong immune response and may prove ideal for selection of subunit vaccine candidate.
Data Availability
The data supporting the findings of this study are available within the article.
Abbreviations
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- ACE2:
-
Angiotensin converting enzyme 2
- RBD:
-
Receptor binding domain
- RBM:
-
Receptor binding motif
- NAb:
-
Neutralizing antibody
- HIV:
-
Human immunodeficiency virus
- SEC:
-
Size exclusion chromatography
References
Lasaulce S, Zhang C, Varma V, Morărescu IC (2021) Analysis of the tradeoff between health and economic impacts of the Covid-19 epidemic. Front Public Health 9:620770. https://doi.org/10.3389/fpubh.2021.620770
Shanmugam C, Mohammed AR, Ravuri S et al (2020) COVID-2019 – A comprehensive pathology insight. Pathol Res Pract 216:153222. https://doi.org/10.1016/j.prp.2020.153222
Mølhave M, Agergaard J, Wejse C (2021) Clinical management of COVID-19 patients – An update. Semin Nucl Med. https://doi.org/10.1053/j.semnuclmed.2021.06.004
Twomey JD, Luo S, Dean AQ et al (2020) COVID-19 update: The race to therapeutic development. Drug Resist Updat 53:100733. https://doi.org/10.1016/j.drup.2020.100733
Krammer F (2020) SARS-CoV-2 vaccines in development. Nature 586:516–527. https://doi.org/10.1038/s41586-020-2798-3
Yadav N, Vishwakarma P, Khatri R et al (2021) Comparative immunogenicity analysis of intradermal versus intramuscular administration of SARS-CoV-2 RBD epitope peptide-based immunogen in vivo. Microbes Infect 23:104843. https://doi.org/10.1016/j.micinf.2021.104843
Vishwakarma P, Yadav N, Rizvi ZA et al (2021) Severe acute respiratory syndrome coronavirus 2 spike protein based novel epitopes induce potent immune responses in vivo and inhibit viral replication in vitro. Front Immunol 12:613045. https://doi.org/10.3389/fimmu.2021.613045
Naqvi AAT, Fatima K, Mohammad T et al (2020) Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: structural genomics approach. Biochim Biophys Acta Mol Basis Dis 1866:165878. https://doi.org/10.1016/j.bbadis.2020.165878
Kumar S, Nyodu R, Maurya VK, Saxena SK (2020) Morphology genome organization replication and pathogenesis of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Coronavirus disease 2019 COVID-19. Springer, Singapore. https://doi.org/10.1007/978-981-15-4814-7_3
Huang Y, Yang C, Xu X et al (2020) Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacol Sin 41:1141–1149. https://doi.org/10.1038/s41401-020-0485-4
Lan J, Ge J, Yu J et al (2020) Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581:215–220. https://doi.org/10.1038/s41586-020-2180-5
Callaway E (2020) The coronavirus is mutating — does it matter? Nature 585:174–177. https://doi.org/10.1038/d41586-020-02544-6
Harvey WT, Carabelli AM, Jackson B et al (2021) SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol 19:409–424. https://doi.org/10.1038/s41579-021-00573-0
Wang R, Zhang Q, Ge J et al (2021) Analysis of SARS-CoV-2 variant mutations reveals neutralization escape mechanisms and the ability to use ACE2 receptors from additional species. Immunity 54:1611-1621.e5. https://doi.org/10.1016/j.immuni.2021.06.003
Abdool Karim SS, de Oliveira T (2021) New SARS-CoV-2 Variants - Clinical, public health, and vaccine implications. N Engl J Med 384:1866–1868. https://doi.org/10.1056/NEJMc2100362
Lau SKP, Woo PCY, Li KSM et al (2005) Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci U S A 102:14040–14045. https://doi.org/10.1073/pnas.0506735102
Yang Y, Liu C, Du L et al (2015) Two mutations were critical for bat-to-human transmission of middle east respiratory syndrome coronavirus. J Virol 89:9119–9123. https://doi.org/10.1128/JVI.01279-15
Meyerson NR, Sawyer SL (2011) Two-stepping through time: mammals and viruses. Trends Microbiol 19:286–294. https://doi.org/10.1016/j.tim.2011.03.006
Li Q, Wu J, Nie J et al (2020) The impact of mutations in SARS-CoV-2 spike on viral infectivity and antigenicity. Cell 182:1284-1294.e9. https://doi.org/10.1016/j.cell.2020.07.012
Akkiz H (2021) Implications of the novel mutations in the SARS-CoV-2 genome for transmission, disease severity, and the vaccine development. Front Med. https://doi.org/10.3389/fmed.2021.636532
Gobeil SM-C, Janowska K, McDowell S et al (2021) Effect of natural mutations of SARS-CoV-2 on spike structure, conformation, and antigenicity. Science. https://doi.org/10.1126/science.abi6226
Esper FP, Cheng Y-W, Adhikari TM et al (2021) Genomic epidemiology of SARS-CoV-2 infection during the initial pandemic wave and association with disease severity. JAMA Netw Open 4:e217746–e217746. https://doi.org/10.1001/jamanetworkopen.2021.7746
Dai L, Gao GF (2021) Viral targets for vaccines against COVID-19. Nat Rev Immunol 21:73–82. https://doi.org/10.1038/s41577-020-00480-0
Dong Y, Dai T, Wei Y et al (2020) A systematic review of SARS-CoV-2 vaccine candidates. Signal Transduct Target Ther 5:1–14. https://doi.org/10.1038/s41392-020-00352-y
Neuzil KM (2021) Interplay between emerging SARS-CoV-2 variants and pandemic control. N Engl J Med 384:1952–1954. https://doi.org/10.1056/NEJMe2103931
Tao K, Tzou PL, Nouhin J et al (2021) The biological and clinical significance of emerging SARS-CoV-2 variants. Nat Rev Genet. https://doi.org/10.1038/s41576-021-00408-x
Imai M, Halfmann PJ, Yamayoshi S et al (2021) Characterization of a new SARS-CoV-2 variant that emerged in Brazil. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.2106535118
Tracking SARS-CoV-2 variants. Available at https://www.who.int/emergencies/emergency-health-kits/trauma-emergency-surgery-kit-who-tesk-2019/tracking-SARS-CoV-2-variants. Accessed 16 Aug 2021
Kober L, Zehe C, Bode J (2013) Optimized signal peptides for the development of high expressing CHO cell lines. Biotechnol Bioeng 110:1164–1173. https://doi.org/10.1002/bit.24776
Ahmed S, Shrivastava T, Kumar N et al (2017) Stabilization of a soluble, native-like trimeric form of an efficiently cleaved Indian HIV-1 clade C envelope glycoprotein. J Biol Chem 292:8236–8243. https://doi.org/10.1074/jbc.M117.776419
Ahmed S, Shrivastava T, Kumar R et al (2021) Design and characterization of a germ-line targeting soluble, native-like, trimeric HIV-1 Env lacking key glycans from the V1V2-loop. Biochim Biophys Acta Gen Subj 1865:129733. https://doi.org/10.1016/j.bbagen.2020.129733
Shah NB, Duncan TM (2014) Bio-layer interferometry for measuring kinetics of protein-protein interactions and allosteric ligand effects. J Vis Exp JoVE. https://doi.org/10.3791/51383
Shin Y-J, König-Beihammer J, Vavra U et al (2021) N-Glycosylation of the SARS-CoV-2 receptor binding domain is important for functional expression in plants. Front Plant Sci 12:1154. https://doi.org/10.3389/fpls.2021.689104
Antonopoulos A, Broome S, Sharov V et al (2020) Site-specific characterization of SARS-CoV-2 spike glycoprotein receptor-binding domain. Glycobiology 31:181–187. https://doi.org/10.1093/glycob/cwaa085
Joyce MG, Zhang B, Ou L et al (2016) Iterative structure-based improvement of a fusion-glycoprotein vaccine against RSV. Nat Struct Mol Biol 23:811–820. https://doi.org/10.1038/nsmb.3267
Krarup A, Truan D, Furmanova-Hollenstein P et al (2015) A highly stable prefusion RSV F vaccine derived from structural analysis of the fusion mechanism. Nat Commun 6:8143. https://doi.org/10.1038/ncomms9143
Liang B, Surman S, Amaro-Carambot E et al (2015) Enhanced neutralizing antibody response induced by respiratory syncytial virus prefusion F protein expressed by a vaccine candidate. J Virol 89:9499–9510. https://doi.org/10.1128/JVI.01373-15
Bertram S, Glowacka I, Müller MA et al (2011) Cleavage and activation of the severe acute respiratory syndrome coronavirus spike protein by human airway trypsin-like protease. J Virol 85:13363–13372. https://doi.org/10.1128/JVI.05300-11
Kirchdoerfer RN, Wang N, Pallesen J et al (2018) Stabilized coronavirus spikes are resistant to conformational changes induced by receptor recognition or proteolysis. Sci Rep 8:15701. https://doi.org/10.1038/s41598-018-34171-7
Böttger R, Hoffmann R, Knappe D (2017) Differential stability of therapeutic peptides with different proteolytic cleavage sites in blood, plasma and serum. PLoS ONE 12:e0178943. https://doi.org/10.1371/journal.pone.0178943
Scheiblhofer S, Laimer J, Machado Y et al (2017) Influence of protein fold stability on immunogenicity and its implications for vaccine design. Expert Rev Vaccines 16:479–489. https://doi.org/10.1080/14760584.2017.1306441
Schilling O, Biniossek ML, Mayer B et al (2018) Specificity profiling of human trypsin-isoenzymes. Biol Chem 399:997–1007. https://doi.org/10.1515/hsz-2018-0107
Kumar S, Tsai CJ, Nussinov R (2000) Factors enhancing protein thermostability. Protein Eng 13:179–191. https://doi.org/10.1093/protein/13.3.179
Yokota K, Satou K, Ohki S (2006) Comparative analysis of protein thermostability: differences in amino acid content and substitution at the surfaces and in the core regions of thermophilic and mesophilic proteins. Sci Technol Adv Mater 7:255–262. https://doi.org/10.1016/j.stam.2006.03.003
Barnes CO, Jette CA, Abernathy ME et al (2020) SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature 588:682–687. https://doi.org/10.1038/s41586-020-2852-1
Deshpande A, Harris BD, Martinez-Sobrido L et al (2021) Epitope classification and rbd binding properties of neutralizing antibodies against SARS-CoV-2 variants of concern. Front Immunol 12:691715. https://doi.org/10.3389/fimmu.2021.691715
Laffeber C, de Koning K, Kanaar R, Lebbink JHG (2021) Experimental evidence for enhanced receptor binding by rapidly spreading SARS-CoV-2 variants. J Mol Biol 433:167058. https://doi.org/10.1016/j.jmb.2021.167058
Barton MI, MacGowan SA, Kutuzov MA et al (2021) Effects of common mutations in the SARS-CoV-2 Spike RBD and its ligand, the human ACE2 receptor on binding affinity and kinetics. eLife 10:e70658. https://doi.org/10.7554/eLife.70658
Davis C, Logan N, Tyson G et al (2021) Reduced neutralisation of the Delta (B.1.617.2) SARS-CoV-2 variant of concern following vaccination. PLoS Pathog 17:e1010022. https://doi.org/10.1371/journal.ppat.1010022
Vignuzzi M, Stone JK, Arnold JJ et al (2006) Quasispecies diversity determines pathogenesis through cooperative interactions in a viral population. Nature 439:344–348. https://doi.org/10.1038/nature04388
Domingo E (2000) Viruses at the edge of adaptation. Virology 270:251–253. https://doi.org/10.1006/viro.2000.0320
Domingo E, Holland JJ (1997) RNA virus mutations and fitness for survival. Annu Rev Microbiol 51:151–178. https://doi.org/10.1146/annurev.micro.51.1.151
Singh J, Pandit P, McArthur AG et al (2021) Evolutionary trajectory of SARS-CoV-2 and emerging variants. Virol J 18:166. https://doi.org/10.1186/s12985-021-01633-w
Petrova VN, Russell CA (2018) The evolution of seasonal influenza viruses. Nat Rev Microbiol 16:47–60. https://doi.org/10.1038/nrmicro.2017.118
Cuevas JM, Geller R, Garijo R et al (2015) Extremely high mutation rate of hiv-1 in vivo. PLoS Biol 13:e1002251. https://doi.org/10.1371/journal.pbio.1002251
Bekker L-G, Tatoud R, Dabis F et al (2020) The complex challenges of HIV vaccine development require renewed and expanded global commitment. The Lancet 395:384–388. https://doi.org/10.1016/S0140-6736(19)32682-0
Acknowledgements
We thank Dr. Pramod Garg (Translational Health Science & Technology Institute) for his critical inputs in this study and guidance. We thank Prof. S. Pöhlmann (Infection Biology Unit, Göttingen, Germany) for the kind gift of ACE2-Fc plasmid. The RBD-His construct (Catalog No. NR-52422) deposited by David Veesler, Department of Biochemistry, University of Washington, Seattle, Washington, USA. and obtained through BEI Resources, NIAID, National Institutes of Health: We also thanks Mr. Vijay Kumar of Regional Centre for Biotechnology (RCB), for assisting with DSC and CD experiments. Part of the work was carried out at the Advanced Technology Platform Centre (ATPC) of RCB.
Funding
This work was supported by the Department of Biotechnology and by Translational Health Science & Technology Institute core grant.
Author information
Authors and Affiliations
Contributions
RK, HAP, SS, and SA methodology and investigation; SS, and SA supervision; RK, HAP, GS, SR, AC, RK and VM design and performed experiments; RK, SS, and SA writing, review and editing; SS, and SA conceptualization and project administration.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khatri, R., Parray, H.A., Siddiqui, G. et al. Biophysical and Biochemical Characterization of the Receptor Binding Domain of SARS-CoV-2 Variants. Protein J 41, 457–467 (2022). https://doi.org/10.1007/s10930-022-10073-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-022-10073-6