Abstract
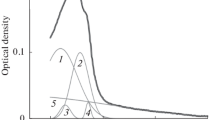
Fluorescence spectroscopy was used to differentiate between different states of acrylodan-labeled cAMP-dependent protein kinase catalytic subunits in urea, guanidine hydrochloride and 3-(N-morpholino)propanesulfonic acid solutions, by measuring changes in the emission spectrum of the protein-coupled dye, which is very sensitive to its microenvironment. Decomposition of the observed fluorescence spectra by a parameterized log-normal distribution function allowed the resolution of overlapping spectral bands and revealed the formation of three distinct protein states, denominated as native, denatured and unfolded structures. At low denaturant concentrations the formation of the denatured form from the native protein was observed, and this process was characterized by a blue-shift of the fluorescence spectrum of acrylodan, indicating that the dye was transferred into some water-deficit hydrophobic environment inside the protein molecule. Therefore, formation of a “dry molten globule” structure could be suggested in state. At high denaturant concentrations a red-shift of the emission spectrum of the protein-coupled probe was observed indicating significant extrusion of the dye molecule into water environment as a result of the unfolding of the protein structure.







Similar content being viewed by others
Abbreviations
- Acrylodan:
-
6-Acryloyl-2-dimethylaminonaphthalene
- ATP:
-
Adenosine triphosphate
- cAMP:
-
3′,5′-Cyclic adenosine monophosphate
- kemptide:
-
Peptide substrate for PKAc with the sequence LRRASLG
- MOPS:
-
3-(N-Morpholino)propanesulfonic acid
- PKAc:
-
3′5′-Cyclic adenosine monophosphate dependent protein kinase catalytic subunit
- TRIS:
-
tris(Hydroxymethyl)aminomethane
- GdnHCl:
-
Guanidine hydrochloride
References
Prendergast F, Meyer M, Carlson GL, Iida S, Potter JD (1983) Synthesis, spectral properties and use of 6-acryloyl-2-dimethylaminonaphtalene (Acrylodan). J Biol Chem 258:7541–7544
Lew J, Coruh N, Tsigelny I, Garrod S, Taylor SS (1997) Synergistic binding of nucleotides and inhibitors to cAMP-dependent protein kinase examined by acrylodan fluorescence spectroscopy. J Biol Chem 272:1507–1513
Kivi R, Loog M, Jemth P, Järv J (2013) Kinetics of acrylodan-labeled cAMP-dependent protein kinase catalytic subunit denaturation. Protein J 32:519–525
Siano DB, Metzler DE (1969) Band shapes of the electronic spectra of complex molecules. J Chem Phys 51:1856–1861
Emel’ianenko VI, YaK Reshetniak, Andreev OA, Burshteĭn EA (2000) Analysis of the log–normal components of the fluorescence spectra of protein-bound prodan and acrylodan. Biophysics (Biofizika) 45:202–214
Bacalum M, Zorilă B, Radu M (2013) Fluorescence spectra decomposition by asymmetric functions: Laurdan spectrum revisited. Anal Biochem 440:123–129
Dempsey CE, Piggot TJ, Mason PE (2005) Dissecting contributions to the denaturant sensitivities of proteins. Biochemistry 44:775–781
Shaknovich EI, Finkelstein AV (1989) Theory of cooperative transition in protein folding. I. Why denaturation of globular protein is a first-order phase transition. Biopolymers 28:1667–1680
Santucci R, Sinibaldi F, Fiorucci L (2008) Protein folding, unfolding and misfolding: role played by intermediate states. Mini Rev Med Chem 8:57–62
Kuznetsov A, Kivi R, Järv J (2015) Computational modeling of acrylodan-labeled cAMP dependent protein kinase catalytic subunit unfolding. Comput Biol Chem 61:197–201
Acknowledgments
This work was financially supported by Estonian Ministry of Education and Research Grant IUT14-20. The authors are thankful to Prof Per Jemth at Uppsala University for thorough discussion of the work, and Dr. Meeme Utt at Tartu University for help with FPLC analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Kivi, R., Järv, J. Different States of Acrylodan-Labeled 3′5′-Cyclic Adenosine Monophosphate Dependent Protein Kinase Catalytic Subunits in Denaturant Solutions. Protein J 35, 331–339 (2016). https://doi.org/10.1007/s10930-016-9676-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-016-9676-8