Abstract
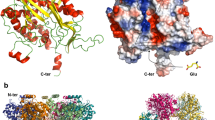
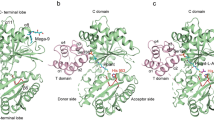
Glutamyl-queuosine-tRNAAsp synthetase (Glu-Q-RS) is a paralog of glutamyl-tRNA synthetase (GluRS) and is found in more than forty species of proteobacteria, cyanobacteria, and actinobacteria. Glu-Q-RS shows striking structural similarity with N-terminal catalytic domain of GluRS (NGluRS) but it lacks the C-terminal anticodon binding domain (CGluRS). In spite of structural similarities, Glu-Q-RS and NGluRS differ in their functional properties. Glu-Q-RS glutamylates the Q34 nucleotide of the anticodon of tRNAAsp whereas NGluRS constitutes the catalytic domain of GluRS catalyzing the transfer of Glu on the acceptor end of tRNAGlu. Since NGluRS is able to catalyze aminoacylation of only tRNAGlu the glutamylation capacity of tRNAAsp by Glu-Q-RS is surprising. To understand the substrate specificity of Glu-Q-RS we undertook a systemic approach by investigating the biophysical and biochemical properties of the NGluRS (1–301), CGluRS (314–471) and Glu-Q-RS-CGluRS, (1–298 of Glu-Q-RS fused to 314–471 from GluRS). Circular dichroism, fluorescence spectroscopy and differential scanning calorimetry analyses revealed absence of N-terminal domain (1–298 of Glu-Q-RS) and C-terminal domain (314–471 from GluRS) communication in chimera, in contrast to the native full length GluRS. The chimeric Glu-Q-RS is still able to aminoacylate tRNAAsp but has also the capacity to bind tRNAGlu. However the chimeric protein is unable to aminoacylate tRNAGlu probably as a consequence of the lack of domain–domain communication.







Similar content being viewed by others
Abbreviations
- Glu-Q-RS:
-
Glutamyl-queuosine-tRNAAsp Synthetase
- GluRS:
-
Glutamyl-tRNA synthetase
- NGluRS:
-
N-terminal catalytic domain of GluRS
- CGluRS:
-
C-terminal anticodon binding domain
- aaRS:
-
Aminoacyl tRNA synthetase
- Ec :
-
Escherichia coli
- Tth :
-
Thermus thermophilus
- CP:
-
Connective peptide
- l-Glu:
-
l-Glutamic acid
- IPTG:
-
Isopropyl β-D-1-thiogalactopyranoside
- PMSF:
-
Phenylmethanesulfonyl fluoride
- CD:
-
Circular dichroism
- UV:
-
Ultra violet
- MRE:
-
Mean residue ellipticity
- θ :
-
Observed ellipticity
- GdnHCl:
-
Guanidine chloride
- f D :
-
Fraction unfolded
- ∆G:
-
Free energy change
- m :
-
Slope of the transition
- DSC:
-
Differential scanning calorimetry
- Tm :
-
Midpoint of transition
- Kd :
-
Dissociation constant
- GoA:
-
Glutamol-AMP
References
Ibba M, Söll D (2000) Aminoacyl-tRNA synthesis. Annu Rev Biochem 69:617–650
Eriani G, Delarue M, Poch O, Gangloff J, Moras D (1990) Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature 347:203–206
Pham Y, Li L, Kim A, Erdogan O, Weinreb V, Butterfoss GL, Kuhlman B, Carter CW Jr (2007) A minimal TrpRS catalytic domain supports sense/antisense ancestry of class I and II aminoacyl-tRNA synthetases. Mol Cell 25:851–862
Saha R, Dasgupta S, Basu G, Roy S (2009) A chimaeric glutamyl: glutaminyl-tRNA synthetase: implications for evolution. Biochem J 417:449–455
Alexander RW, Schimmel P (2001) Domain-domain communication in aminoacyl-tRNA synthetases. Prog Nucleic Acids Res Mol Biol 69:317–349
Schimmel P, Ribas de Pouplana L (2000) Footprints of aminoacyl-tRNA synthetases are everywhere. Trends Biochem Sci 25:207–209
Olmedo-Verd E, Santamaría-Gómez J, Ochoa de Alda JA, Ribas de Pouplana L, Luque I (2011) Membrane anchoring of aminoacyl-tRNA synthetases by convergent acquisition of a novel protein domain. J Biol Chem 286:41057–41068
Ibba M, Söll D (2004) Aminoacyl-tRNAs: setting the limits of the genetic code. Genes Dev 18:731–738
Sampath P, Mazumder B, Seshadri V, Gerber CA, Chavatte L, Kinter M, Ting SM, Dignam JD, Kim S, Driscoll DM, Fox PL (2004) Noncanonical function of glutamyl-prolyl-tRNA synthetase: gene-specific silencing of translation. Cell 119:195–208
Ibba M, Francklyn C (2004) Turning tRNA upside down: when aminoacylation is not a prerequisite to protein synthesis. Proc Natl Acad Sci USA 101:7493–7494
Salazar JC, Ambrogelly A, Crain PF, McCloskey JA, Söll D (2004) A truncated aminoacyl–tRNA synthetase modifies RNA. Proc Natl Acad Sci USA 101:7536–7541
Dubois DY, Blaise M, Becker HD, Campanacci V, Keith G, Giegé R, Cambillau C, Lapointe J, Kern D (2004) An aminoacyl-tRNA synthetase-like protein encoded by the Escherichia coli yadB gene glutamylates specifically tRNAAsp. Proc Natl Acad Sci USA 101:7530–7535
Fujita N, Mori H, Yura T, Ishihama A (1994) Systematic sequencing of the Escherichia coli genome: analysis of the 2.4–4.1 min (110,917-193,643 bp) region. Nucleic Acids Res 22:1637–1639
Gagnon Y, Lacoste L, Champagne N, Lapointe J (1996) Widespread use of the glu-tRNAGln transamidation pathway among bacteria. A member of the alpha purplebacteria lacks glutaminyl-trna synthetase. J Biol Chem 271:14856–14863
Campanacci V, Dubois DY, Becker HD, Kern D, Spinelli S, Valencia C, Pagot F, Salomoni A, Grisel S, Vincentelli R, Bignon C, Lapointe J, Giegé R, Cambillau C (2004) The Escherichia coli YadB gene product reveals a novel aminoacyl-tRNA synthetase like activity. J Mol Biol 337:273–283
Blaise M, Becker HD, Lapointe J, Cambillau C, Giegé R, Kern D (2005) Glu-Q-tRNA(Asp) synthetase coded by the yadB gene, a new paralog of aminoacyl-tRNA synthetase that glutamylates tRNA(Asp) anticodon. Biochimie 87:847–861
Blaise M, Becker HD, Keith G, Cambillau C, Lapointe J, Giegé R, Kern D (2004) A minimalist glutamyl-tRNA synthetase dedicated to aminoacylation of the tRNAAsp QUC anticodon. Nucleic Acids Res 32:2768–2775
Kern D, Lapointe J (1979) Glutamyl transfer ribonucleic acid synthetase of Escherichia coli. Study of the interactions with its substrates. Biochemistry 25:5809–5818
Dubois DY, Blais SP, Huot JL, Lapointe J (2009) A C-truncated glutamyl-tRNA Synthetase specific for tRNA (Glu) is stimulated by its free complementary distal domain: mechanistic and evolutionary implications. Biochemistry 48:6012–6021
Saha R, Dasgupta S, Banerjee R, Mitra-Bhattacharyya A, Söll D, Basu G, Roy S (2012) A functional loop spanning distant domains of glutaminyl-tRNA synthetase also stabilizes a molten globule state. Biochemistry 51:4429–4437
Dasgupta S, Saha R, Dey C, Banerjee R, Roy S, Basu G (2009) The role of the catalytic domain of E. coli GluRS in tRNAGln discrimination. FEBS Lett 583:2114–2120
Madore E, Florentz C, Giegé R, Sekine S, Yokoyama S, Lapointe J (1999) Effect of modified nucleotides on Escherichia coli tRNAGlu structure and on its aminoacylation by glutamyl-tRNA synthetase. Predominant and distinct roles of the mnm5 and s2 modifications of U34. Eur J Biochem 266:1128–1135
Chen YH, Yang JT, Martinez HM (1972) Determination of the secondary structures of proteins by circular dichroism and optical rotatory dispersion. Biochemistry 11:4120–4131
Gull N, Sen P, Khan RH, Kabir-ud-Din (2009) Spectroscopic studies on the comparative interaction of cationic single-chain and gemini surfactants with human serum albumin. J Biochem 145:67–77
Greene RF, Pace CN (1974) Urea and guanidine hydrochloride denaturation of ribonuclease, lysozyme, alpha-chymotrypsin, and beta-lactoglobulin. J Biol Chem 249:5388–5393
Saito Y, Wada A (1983) Comparative study of GuHCl denaturation of globular proteins I. Spectroscopic and chromatographic analysis of the denaturation curves of ribonuclease A, cytochrome c, and pepsinogen. Biopolymers 22:2105–2122
Pace CN (1986) Determination and analysis of urea and guanidine hydrochloride denaturation curves. Method Enzymol 131:266–280
Scholtz JM (1995) Conformational stability of HPr: the histidine-containing phosphocarrier protein from Bacillus subtilis. Protein Sci 4:35–43
Matthews CR, Crisanti MM (1981) Urea-induced unfolding of the alpha subunit of tryptophan synthease: evidence for a multistate process. Biochemistry 20:784–792
Scholtz JM, Grimsley GR, Pace CN (2009) Solvent denaturation of proteins and interpretations of the m value. Method Enzymol 466:549–565
Shaw KL, Scholtz JM, Pace CN, Grimsley GR (2009) Determining the conformational stability of a protein using urea denaturation curves. Method Mol Biol 490:41–55
Mandal AK, Samaddar S, Banerjee R, Lahiri S, Bhattacharyya A, Roy S (2003) Glutamate counteracts the denaturing effect of urea through its effect on the denatured state. J Biol Chem 278:36077–36084
Banerjee R, Mandal AK, Saha R, Guha S, Samaddar S, Bhattacharyya A, Roy S (2003) Solvation change and ion release during aminoacylation by aminoacyl-tRNA synthetases. Nucleic Acid Res 31:6035–6042
Lakowicz JR (2006) Principles of fluorescence spectroscopy. Springer, New York
Roy S (2004) Fluorescence quenching methods to study protein–nucleic acid interactions. Method Enzymol 379:175–187
Bruylants G, Wouters J, Michaux WC (2005) Differential scanning calorimetry in life science: thermodynamics, stability, molecular recognition and application in drug design. Curr Med Chem 12:2011–2020
Maity H, O’Dell C, Srivastava A, Goldstein J (2009) Effects of arginine on photostability and thermal stability of IgG1 monoclonal antibodies. Curr Pharm Biotechnol 10:761–766
Paulsson M, Hegg PO, Castberg HB (1985) Thermal stability of whey proteins studied by differential scanning calorimetry. Thermochim Acta 95:435–440
Lapointe J, Delcuve G (1975) Thermosensitive mutants of Escherichia coli K-12 altered in the catalytic subunit and in a regulatory factor of the glutamy-transfer ribonucleic acid synthetase. J Bacteriol 122:352–358
Kiefer F, Arnold K, Künzli M, Bordoli L, Schwede T (2009) The swiss-model repository and associated resources. Nucleic Acid Res 37:D387–D392
Sekine S, Nureki O, Dubois DY, Bernier S, Chênevert R, Lapointe J, Vassylyev DG, Yokoyama S (2003) ATP binding by glutamyl-tRNA synthetase is switched to the productive mode by tRNA binding. EMBO J 22:676–688
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Features and development of Coot. Acta Crystallogr D Biol Crystallogr 66:486–501
Adams PD, Afonine PV, Bunkóczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH (2010) PHENIX: a comprehensive python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr 66:213–221
Chen VB, Arendall WB 3rd, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, Murray LW, Richardson JS, Richardson DC (2010) MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr 66:12–21
Kelly SM, Price NC (2000) The use of circular dichroism in the investigation of protein structure and function. Curr Protein Pept Sci 1:349–384
Carra JH, Privalov PL (1995) Energetics of denaturation and m values of staphylococcal nuclease mutants. Biochemistry 34:2034–2041
Agashe VR, Udgaonkar JB (1995) Thermodynamics of denaturation of barstar: evidence for cold denaturation and evaluation of the interaction with guanidine hydrochloride. Biochemistry 34:3286–3299
Callis PR, Liu T (2004) Quantitative prediction of fluorescence quantum yields for tryptophan in proteins. J Phys Chem B 108:4248–4259
Hammamieh R, Yang DC (2001) Magnesium ion-mediated binding to tRNA by an amino-terminal peptide of a class II tRNA synthetase. J Biol Chem 276:428–433
Blaise M, Olieric V, Sauter C, Lorber B, Roy B, Karmakar S, Banerjee R, Becker HD, Kern D (2008) Crystal structure of glutamyl-Queuosine tRNAAsp synthetase complexed with l-glutamate: structural elements mediating tRNA-independent activation of glutamate and glutamylation of tRNAAsp anticodon. J Mol Biol 381:1224–1237
Patthy L (2008) Evolution of paralogous proteins. Protein evolution. John Wiley and sons Blackwell Publishing Ltd, Chichester
Siatecka M, Rozek M, Barciszewski J, Mirande M (1998) Modular evolution of the Glx-tRNA synthetase family–rooting of the evolutionary tree between the bacteria and archaea/eukarya branches. Eur J Biochem 256:80–87
Sekine S, Nureki O, Tateno M, Yokoyama S (1999) The identity determinants required for the discrimination between tRNAGlu and tRNAAsp by glutamyl-tRNA synthetase from Escherichia coli. Eur J Biochem 261:354–360
Rogers MJ, Adachi T, Inokuchi H, Söll D (1994) Functional communication in the recognition of tRNA by Escherichia coli glutaminyl-tRNA synthetase. Proc Natl Acad Sci USA 91:291–295
Sherman JM, Thomann HU, Söll D (1996) Functional connectivity between tRNA binding domains in glutaminyl-tRNA synthetase. J Mol Biol 256:818–828
Uter NT, Perona JJ (2004) Long-range intramolecular signaling in a tRNA synthetase complex revealed by pre-steady-state kinetics. Proc Natl Acad Sci USA 101:14396–14401
Moulinier L, Eiler S, Eriani G, Gangloff J, Thierry JC, Gabriel K, McClain WH, Moras D (2001) The structure of an AspRS-tRNA(Asp) complex reveals a tRNA-dependent control mechanism. EMBO J 20:5290–5301
Sauter C, Lorber B, Cavarelli J, Moras D, Giegé R (2000) The free yeast aspartyl-tRNA synthetase differs from the tRNA (Asp)-complexed enzyme by structural changes in the catalytic site, hinge region, and anticodon-binding domain. J Mol Biol 299:1313–1324
Tzenq SR, Kalodimos CG (2012) Protein activity regulation by conformational entropy. Nature 488:236–240
Acknowledgments
We are grateful to Professor Gautam Basu, Department of Biophysics, Bose Institute for allowing us to use the CD instrument. Dr. Saumya Dasgupta is acknowledged for his assistance during CD measurements. Ec His-tagged GluRS overproducing strain, pKR15 plasmid expressing Ec tRNAGlu and JP1449 (DE3) were kindly gifted by Professor Jacques Lapointe (Department of Biochemistry and Microbiology, Université Laval, Québec, Canada). We also like to thank Professor Gabor Igloi, University of Freiburg, Germany for critical reading of the manuscript and for providing in vitro transcribed jack bean tRNAArg. Molecular Mechanism of Disease and Drug Action (MMDDA) project (Grant 11-R&D-SIN-5.04) Department of Atomic Energy, Govt. of India is kindly acknowledged for providing the fund for purchasing differential scanning calorimetry (DSC) instrument. S.R. thanks to University of Calcutta for providing fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ray, S., Blaise, M., Roy, B. et al. Fusion with Anticodon Binding Domain of GluRS is Not Sufficient to Alter the Substrate Specificity of a Chimeric Glu-Q-RS. Protein J 33, 48–60 (2014). https://doi.org/10.1007/s10930-013-9537-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-013-9537-7