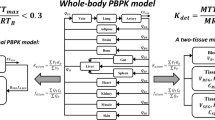
Abstract
In drug discovery and development, classical compartment models and physiologically based pharmacokinetic (PBPK) models are successfully used to analyze and predict the pharmacokinetics of drugs. So far, however, both approaches are used exclusively or in parallel, with little to no cross-fertilization. An approach that directly links classical compartment and PBPK models is highly desirable. We derived a new mechanistic lumping approach for reducing the complexity of PBPK models and establishing a direct link to classical compartment models. The proposed method has several advantages over existing methods: Perfusion and permeability rate limited models can be lumped; the lumped model allows for predicting the original organ concentrations; and the volume of distribution at steady state is preserved by the lumping method. To inform classical compartmental model development, we introduced the concept of a minimal lumped model that allows for prediction of the venous plasma concentration with as few compartments as possible. The minimal lumped parameter values may serve as initial values for any subsequent parameter estimation process. Applying our lumping method to 25 diverse drugs, we identified characteristic features of lumped models for moderate-to-strong bases, weak bases and acids. We observed that for acids with high protein binding, the lumped model comprised only a single compartment. The proposed lumping approach established for the first time a direct derivation of simple compartment models from PBPK models and enables a mechanistic interpretation of classical compartment models.














Similar content being viewed by others
References
Kwon Y (2001) Handbook of essential pharmacokinetics, pharmacodynamics, and drug metabolism for industrial scientists. Springer-Verlag, New York, Inc
Derendorf H, Lesko LJ, Chaikin P, Colburn WA, Lee P, Miller R, Powell R, Rhodes G, Stanski D, Venitz J (2000) Pharmacokinetic/pharmacodynamic modeling in drug research and development. J Clin Pharmacol 40:1399–1418
Schoenwald RD (2002) Pharmacokinetics in drug discovery and development. CRC Press, Boca Raton
Tozer TN, Rowland M (2006) Introduction to pharmacokinetics and pharmacodynamics. Lippincott Williams & Wilkins, Philadelphia
Lüpfert C, Reichel A (2005) Development and application of physiologically based pharmacokinetic modeling toos to support drug discovery. Chem Biodivers 2:1462–1486
Jones HM, Parrott N, Jorga K, Lavé T (2006) A novel strategy for physiologically based predictions of human pharmacokinetics. Clin Pharmacokinet 45:511–542
Theil FP, Guentert TW, Haddad S, Poulin P (2003) Utility of physiologically based pharmacokinetic models to drug development and rational drug discovery candidate selection. Toxicol Lett 138:29–49
Schmitt W, Willmann S (2005) Physiology-based pharmacokinetic modeling: ready to be used. Drug Discov Today 2:125–132
Jones HM, Gardner IB, Watson KJ (2009) Modelling and PBPK simulation in drug discovery. AAPS J 11:155–166
Bourne DWA (1995) Mathematical modeling of pharmacokinetic data. Technomic Publishing Company, Inc., Lancaster
Nestorov IA, Aarons LJ, Arundel PA, Rowland M (1998) Lumping of whole-body physiologcally based pharmacokinetic models. J Pharmacokinet Pharmacodyn 26:21–46
Brochot C, Toth J, Bois FY (2005) Lumping in pharmacokinetics. J Pharmacokinet Pharmacodyn 32:719–736
Gueorguieva I, Nestorov IA, Rowland M (2006) Reducing whole body physiologically based pharmacokinetic models using global sensitivity analysis: diazepam case study. J Pharmacokinet Pharmacodyn 33:1–27
Björkman S (2003) Reduction and lumping of physiologically based pharmacokinetic models: prediction of the disposition of fentanyl and pethidine in human by successively simplified models. J Pharmacokinet Pharmacodyn 30:285–307
Rodgers T, Leahy D, Rowland M (2005) Physiologically based pharmacokinetic modeling 1: predicting the tissue distribution of moderate-to-strong bases. J Pharm Sci 94:1259–1276
Rodgers T, Leahy D, Rowland M (2007) Errata: Physiologically based pharmacokinetic modeling 1: predicting the tissue distribution of moderate-to-strong bases. J Pharm Sci 96:3151–3152
Rodgers T, Rowland M (2005) Physiologically based pharmacokinetic modelling 2: predicting the tissue distribution of acids, very weak bases, neutrals and zwitterions. J Pharm Sci 95:1238–1257
Rodgers T, Rowland M (2007) Errata: physiologically based pharmacokinetic modelling 2: predicting the tissue distribution of acids, very weak bases, neutrals and zwitterions. J Pharm Sci 96:3153–3154
Gerlowski LE, Jain RK (1983) Physiologically based pharmacokinetic modeling: principles and application. J Pharm Sci 72:1103–1127
Nestorov IA (2003) Whole body pharmacokinetic models. Clin Pharmacokinet 42:883–908
Luttringer O, Theil FP, Poulin P, Schmitt-Hoffmann AH, Guentert TW, Lavé T (2003) Physiologically based pharmacokinetic modelling of disposition of epiroprim in humans. J Pharm Sci 92:1990–2007
Willmann S, Höhn K, Edginton A, Sevestre M, Solodenko J, Weiss W, Lippert J, Schmitt W (2007) Development of a physiologically-based whole-body population model for assessing the influence of individual variability on the pharmacokinetics of drugs. J Pharmacokinet Pharmacodyn 34:401–431
International Commission on Radiological Protection (ICRP) (2002) Basic anatomical and physiological data for use in radiological protection: reference values. ICRP Publication 89
Poulin P, Theil FP (2009) Development of a novel method for predicting human volume of distribution at steady-state of basic drugs and comparative assessment with existing methods. J Pharm Sci 9999:1–29
Obach RS (1999) Prediction of human clearance of twenty-nine drugs from hepatic microsomal intrinsic clearance data: an examination of in vitro half-life approach and nonspecific binding to microsomes. Drug Metab Dispos 27:1350–1359
Riley RJ, McGinnity DF, Austin RP (2005) A unified model for predicting human hepatic, metabolic clearance from in vitro intrinsic clearance data in hepatocytes and microsomes. Drug Metab Dispos 33:1304–1311
Rodgers T, Rowland M (2007) Mechanistic approaches to volume of distribution predictions: understanding the process. Pharm Res 24:918–933
Jones HM, Houston JB (2004) Substrate depletion approach for determining in vitro metabolic clearance: time dependencies in hepatocyte and microsomal incubations. Drug Metab Dispos 32:973–982
DrugBank (2010) Diclofenac. http://drugbank.ca/drugs/DB00586
DrugBank (2010) Warfarin. http://www.drugbank.ca/drugs/DB00682
Jung D, Mayersohn M, Perrier D, Calkins J, Saunders R (1982) Thiopental disposition in lean and obese patients undergoing surgery. Anesthesiology 56:269–274
Russo H, Simon N, Duboin MP, Urien S (1997) Population pharmacokinetics of high-dose thiopental in patients with cerebral injuries. Clin Pharmacol Ther 62:15–20
Löscher W (1978) Serum protein binding and pharmacokinetics of valproate in man, dog, rat and mouse. J Pharmacol Exp Ther 204:255–261
Food and Drug Administration (2004) Approval letter: Lidocaine
Blakey GE, Nestorov IA, Arundel PA, Aarons LJ, Rowland M (1997) Quantitative structure pharmacokinetics relationship I: development of a whole-body physiologically based pharmacokinetic model to characterize changes in pharmacokinetics across a homologous series of barbiturates in the rat. J Pharmacokinet Biopharm 25:277–312
von Kleist M, Huisinga W (2007) Physiologically based pharmacokinetic modelling: a sub-compartmentalized model of tissue distribution. J Pharmacokinet Pharmacodyn 34:789–806
Boyes RN, Adams HJ, Duce BR (1970) Oral absorption and disposition kinetics of lidocaine hydrochloride in dogs. J Pharmacol Exp Ther 174:1–8
Adjepon-Yamoah KK, Scott DB, Prescott LF (1974) The effect of atropine on the oral absorption of lidocaine in man. Eur J Clin Pharmacol 7:397–400
Berezhkovskiy LM (2004) Volume of distribution at steady state for a linear pharmacokinetic system with peripheral elimination. J Pharm Sci 93:1628–1640
Yates JWT, Arundel PA (2008) On the volume of distribution at steady state and its relationship with two-compartmental models. J Pharm Sci 97:111–122
Heizmann P, Eckert M, Ziegler WH (1983) Pharmacokinetics and bioavailability of midazolam in man. Br J Clin Pharmacol 16:43–49
Zomorodi K, Donner A, Somma J, Barr J, Sladen R, Ramsay J, Geller E, Shafer SL (1998) Population pharmacodynamics of midazolam administered by target controlled infusion for sedation following coronary artery bypass grafting. Anesthesiology 89:1418–1429
Swart EL, Zuideveld KP, de Jongh J, Danhof M, Thijs LG, van Schijndel RMJS (2003) Comparative population pharmacokinetics of lorazepam and midazolam during long-term continuous infusion in critically ill patients. Br J Clin Pharmacol 57:135–145
Stanski DR, Maitre PO (1990) Population pharmacokinetics and pharmacodynamics of thiopental: the effect of age revisited. Anesthesiology 72:399–402
Gugler R, Schell A, Eichelbaum M, Fröscher W, Schulz HU (1977) Disposition of valproic acid in man. Eur J Clin Pharmacol 12:125–132
Blanco-Serrano B, Otero MJ, Santos-Buelga D, Garcia-Sanchez MJ, Serrano J, Dominguez-Gil A (1999) Population estimation of valproic acid clearance in adult patients using routine clinical pharmacokinetic data. Biopharm Drug Dispos 20:233–240
Rostami-Hodjegan A, Peacey SR, George E, Heller SR, Tucker GT (1998) Population-based modeling to demonstrate extrapancreatic effects of tolbutamide. Am J Physiol Endocrinol Metab 274:758–771
Kwa A, Sprung J, Van Guilder M, Jelliffe RW (2008) A population pharmacokinetic model of epidural lidocaine in geriatric patients: effects of low-dose dopamine. Ther Drug Monit 30:379–389
Price PS, Conolly RB, Chaisson CF, Gross EA, Young JS, Mathis ET, Tedder DR (2003) Modeling interindividual variation in physiological factors used in PBPK models of humans. Crit Rev Toxicol 33:469–503
Acknowledgements
The authors kindly acknowledge comments on the manuscript by Charlotte Kloft (Clinical Pharmacy, Martin-Luther-Universität Halle-Wittenberg/ Germany), Steve Kirkland (Hamilton Institute, NUIM/Ireland), Andreas Reichel (Bayer Schering Pharma) and Olaf Lichtenberger (Abbott). S.P. acknowledges financial support from the Graduate Research Training Program PharMetrX: Pharmacometrics and Computational Disease Modeling, Martin-Luther-Universität Halle-Wittenberg and Freie Universität Berlin, Germany (http://www.pharmacometrics.de).
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix A: Derivation of the relation between the lumped and the original concentrations
The relation between the concentrations of the lumped compartment and the comprised original compartments is given by Eq. 22:
The idea is to establish a link between the concentration C tis of one of the original organs (or tissues) and the concentration C L of the associated lumped compartment. Let us arbitrarily choose one organ (named ‘ref’). Using the lumping criteria, we have C tis/K tis = C ref/(K ref(1 − E ref)) and thus C tis = C ref K tis/(K ref(1 − E ref)) for all tissue/organs lumped into ‘L’. In combination with the above equation for C L this yielded
or equivalently
Using the definition of K L in Eq. 23 and 24 we obtain the desired relation:
Since the tissue/organ ‘ref’ was arbitrarily chosen, the above relation holds for every original compartment ‘tis’ that is part of the lumped compartment ’L’, being eliminating (in which case E ref > 0) or non-eliminating (in which case E ref = 0).
Appendix B: General derivation of lumped ODEs
In the following, the subscript tis refers to all organs of the PBPK model excluding the liver, i.e., adi, bra, bon, gut, hea, kid, lun, mus, ski, spl.
To derive general equations for the rate of change of the lumped concentrations C L it is advantageous to bring the original ODEs of the generic PBPK in a different but equivalent form, where elimination is associated with the venous compartment and all other compartments have the same structural form. In the generic PBPK model (see Section ‘Material and Methods’), the liver is assumed to be the only eliminating organ with ODE (see Eq. 3)
Defining Rhep = CLint K liv/Q liv and noting that 1 + Rhep = 1/(1 − Ehep), where Ehep is the hepatic extraction ratio defined in Eq. 16, we obtain
Now, the inflowing concentration of the vein is given by (see Eq. 7)
which can be rewritten as
Let us define
where E tis denotes the tissue elimination ratio. In our case, it is E liv = Ehep and E tis = 0 otherwise. Moreover, formally define \(\widehat{K}_{\rm ven} = \widehat{K}_{\rm art} =1\). Then, we finally obtain an equivalent formulation of the whole-body PBPK model. For all organs, tissues and other spaces except vein, it is
while for the vein it is
where we exploit CLblood = Q livEhep based on Eq. 67.
We determined the equation for the rate of change of the lumped concentration C L by differentiating Eq. 22, yielding:
The right hand side V tisd/dt C tis is defined in Eqs. 66 and 67. The right hand side of contain the concentrations of the original PBPK model C tis, which can be determined using Eq. 57.
We obtained a very simple form of equations for the mechanistically lumped model, when spleen and gut were lumped together into the same compartment as the liver (hence they were part of the lumped ‘Liv’ compartment), and when lung and artery were lumped together into the same compartment as the vein (hence, they were part of the lumped ‘cen’ compartment). In this case, the influent concentrations will be identical for all compartments different from the central compartment. The rate of change for the concentration of the pre-lumped compartments spl-gut-liv (‘sgl’) and ven-lun-art (‘vla’) are
and
where C in is the concentration flowing into the vein.
Now, we have for any lumped compartment excluding the central compartment:
where the sum is taken over all tissues that are lumped into ‘L’. Our above assumption (resulting in Eq. 69) ensures that the inflowing concentration is the same and identical to C art for all tissue/organs. Exploiting the lumping condition \({C_{\rm tis}}/{\widehat{K}}={C_{\rm L}}/{K_{\rm L}}\) yields
Finally, using Q L = ∑tis Q tis and C art = C cen/K cen results in
For the central compartment, we obtain analogously
where we exploited the fact that \(C_{\rm liv}/\widehat{K}_{\rm liv} = C_{\rm Liv}/K_{\rm Liv}\).
The above equations do not take into account any dosing. Again, as in the whole-body PBPK case, the corresponding ODEs of the lumped compartments comprising vein and liver have to be amended correspondingly. For an i.v. infusion r iv (see Eq. 9 for the definition), it is
while for a p.o. administration \(r_{{\rm po}({\rm F}_{{\rm F}\cdot {\rm G}})}\) (see Eq. 9 for the definition) it is
If ‘cen’ and ‘Liv’ are identical, i.e., the liver is lumped into the central compartment, then it is C cen/K cen = C Liv/K Liv and thus
Note that in this case, the p.o. administration model has to account for the hepatic extraction, i.e., the absorption model with F bio = (1 − Ehep)F F·G (see Eq. 13) is used rather then the model with F F·G (see Eq. 10).
Appendix C: Lumping of permeability rate-limited tissue model
The rates of change of the vascular concentration C vas and the tissue concentration C tis corresponding to a permeability limited tissue model are given by:
The amount of drug that can transfer from the vascular to the tissue part is limited by the maximal amount of drug, entering the tissue, i.e, Q tis C in. This has to be taken into account, when lumping the tissue space together with other compartments. The following idea is similar to the approach to determine the blood clearance from the intrinsic clearance. It is based on a quasi-steady state assumption on C vas (Eq. 81) yielding
or
Thus, in steady state, the vascular concentration is a weighted sum of the influent concentration C in and the concentration leaving the tissue compartment C tis/K tis. For permeability-rate limited organs, we would usually expect
and as a consequence, we lump the vascular compartment with volume V vas together with the blood compartment and approximate C vas = C blood.
Inserting this formula for C vas into Eq. 82 for the tissue concentration yields
and finally
Hence, when lumping the tissue part of a permeability-rate limited tissue model, the term
should take the role of the tissue blood flow Q tis. It is bounded by both Q tis and PStis, as one would expect.
Appendix D: Automated determination of the number of lumped compartments of the mechanistically lumped model, and its composition
To compare the mechanistically lumped PK compartment models for a number of different drugs, we used an automated detection algorithm to determine the number of lumped compartment and its composition. The input were the normalized concentration-time profiles as predicted by the detailed whole-body PBPK model (cf. Eq. 19):
We determined the similarity matrix M = (M ij ) with entries
where 〈·, ·〉 denotes the Euclidian scalar product. In our setting, if C tis is given as a vector at different time points c tis(t 1), ..., c tis(t M ) for some M > 0, then
where we, for simplicity, assume that the time points are equally spaced with distance \(\Updelta t\).
Next, we determined the eigenvector v corresponding to the maximal eigenvalue of M. This eigenvector has an entry corresponding to each organ, tissue or other space of the whole-body PBPK model. We normalized the eigenvector
such that the normalized eigenvector satisfied w(ven) = 1. See Fig. 15 for the eigenvector corresponding to our model compound Lidocaine (for sake of illustration, we ordered the entries in increasing order).
We then considered the smallest entry of w (in the example corresponding to the muscle tissue) and lumped all organs that satisfied:
with \(\Updelta w=0.045\). For our model compound Lidocaine, there was no such organ. Hence, the muscle tissue comprised a single lumped compartment. We proceeded with the next tissue (adipose in our example):
In this case, w(bon) was the only organ to satisfy the above inequality so that adipose and bone were lumped together. We then proceeded with skin etc. The value of \(\Updelta w\) was chosen so that the automated lumping procedure gave the same results as the manually chosen lumping for Lidocaine. The smaller the value of \(\Updelta w\) the more similar the concentration-time profiles have to be for two organs/tissues to be lumped together.
For Caffeine, Diazepam and Amobarbital the predicted mechanistically lumped model was not sufficient to predict all concentration-time profiles of the 13-compartment whole-body PBPK model. In this case, we manually added a lumped compartment, which resolved the problem and increased the number of compartments by 1.
Rights and permissions
About this article
Cite this article
Pilari, S., Huisinga, W. Lumping of physiologically-based pharmacokinetic models and a mechanistic derivation of classical compartmental models. J Pharmacokinet Pharmacodyn 37, 365–405 (2010). https://doi.org/10.1007/s10928-010-9165-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10928-010-9165-1