Abstract
Diversification and adaptive radiations are tied to evolvability, which in turn is linked to morphological integration. Tightly integrated structures typically evolve in unison, whereas loosely integrated structures evolve separately. Highly integrated structures are therefore thought to constrain evolutionary change by limiting morphological disparity. Mounting evidence suggests that high integration may facilitate evolutionary change along a single trajectory. We used geometric morphometrics to compare cranial disparity and integration among phyllostomid bats—which exhibit the greatest dietary diversity of any mammalian family— and their sister taxa within the superfamily Noctilionoidea. Our results reveal that phyllostomids are more tightly integrated and have less disparity in cranial shape than their outgroups, despite exhibiting tenfold higher species richness and significantly increased rates of speciation. Phyllostomid cranial morphology appears to have diverged from that of other noctilionoids by evolving along a single axis of morphological variation that describes the relative length of the rostrum. We propose that phyllostomids were able to evolve to occupy a wide range of dietary niches by varying rostrum length, possibly along a line of least evolutionary resistance. This study provides a compelling empirical example of how increased integration can lead to adaptation, implying that both high and low integration can underlie diverse phenotypes in adaptive radiation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Adaptive radiations are characterized by the rapid diversification of species and divergence of new forms, often as a result of an evolutionary innovation (Simpson 1953; Schluter 2000; Yoder et al. 2010). As lineages move into a variety of previously unoccupied niches, they can experience high rates of morphological change, increasing clade-wide morphological disparity and functional disparity. The term ‘evolvability’ refers to the potential for diversification and describes the capacity of a population to evolve in the direction of selection by generating adaptive genetic diversity (Kirschner and Gerhart 1998; Hansen and Houle 2008). Highly evolvable populations are predicted to have the potential to speciate more rapidly in response to ecological opportunity (Young et al. 2010; Goswami et al. 2014). Evolvability can be measured quantitatively from morphological data through tests of morphological integration, which examine whether components of an organism, such as the cranium, are composed of one or more parts (Olson and Miller 1958; Mitteroecker and Bookstein 2007; Klingenberg 2008, 2013). While a tightly integrated cranium is composed of a single unit (e.g., the cranium), a less integrated cranium may be composed of multiple components, termed modules (e.g., rostrum, basicranium, cranial vault), which are individually tightly integrated, but only loosely integrated with other modules (Klingenberg 2009).
Integration is thought to affect phenotypic divergence in two different ways, one that requires loosely organized modules and another that operates within highly integrated modules. Theoretically, when structures are not tightly integrated and are composed of two or more modules, constituent modules are able to evolve independently of one another, increasing a taxon’s ability to adapt rapidly to environmental changes, which in turn drives an increase in species diversity and morphological shape disparity. A number of studies demonstrate this association empirically (Drake and Klingenberg 2008; Marroig et al. 2009; Young et al. 2010; Collar et al. 2014; Arlegi et al. 2018). For example, Young and Hallgrímsson (2005) found lower magnitudes of integration of forelimb and hind limb shape in bipedal mammals (bats and primates) in comparison with quadrupedal mammals. In this case, the authors hypothesized that decreased integration allowed limb proportions to escape phylogenetic and developmental constraints, explore new regions of morphospace (e.g., bipedalism), develop novel morphologies, and thereby increase overall morphological disparity.
In contrast, other studies identify clades with high rates of speciation and morphological evolution that are characterized by high levels of integration (Hu et al. 2016; Sherratt et al. 2017), possibly as a result of selection occurring along a single trajectory that may represent a line of least evolutionary resistance (Schluter 1996; Marroig and Cheverud 2005; Felice et al. 2018). For example, Hu et al. (2016) found high rates of speciation and morphological evolution coupled with high integration and neutral levels of disparity in icefish crania. They hypothesized that high integration constrained icefish crania to a single evolutionary trajectory, which decreased disparity but allowed for increased speciation relative to less integrated sister clades because that trajectory allowed increased differentiation in craniofacial height and length.
New World leaf-nosed bats (Family Phyllostomidae) are a well-known example of an adaptive radiation that was spurred, or at least accompanied by, morphological innovations (e.g., Freeman 2000; Jones et al. 2005; Dumont et al. 2011, 2014; Shi and Rabosky 2015; Hedrick and Dumont 2018). Phyllostomids are among the largest bat families and have a broader range of dietary ecologies than any other mammalian family (insectivory, folivory, frugivory, sanguinivory, nectarivory, and omnivory). With the exception of Old World fruit bats (Family Pteropodidae), all other bats are primarily insectivorous. Recently, Rossoni et al. (2019) used a quantitative genetic approach to assess the evolution of morphological integration within the crania of phyllostomids. They found overarching commonalities in patterns of integration but striking differences among species with very different morphologies. They also found that the magnitudes of integration decreased substantially during three periods of increased species diversification, suggesting a link between low integration and speciation. While this quantitative genetic approach is very different from the morphometric analyses typically used to investigate integration, it recovered the same signal of strong selection for divergent morphologies associated with dietary specialization found in previous studies (e.g., Freeman 1981; Dumont et al. 2011, 2014; Rossoni et al. 2017).
Rossoni et al. (2019) provided an informative assessment of integration and morphological disparity within and among phyllostomids, but the study did not address whether and how phyllostomids differ from their close relatives. Here we extend the analysis of integration and disparity in the shape of the cranium to all families within the Superfamily Noctilionoidea: Noctilionidae, Mormoopidae, Mystacinidae, Furipteridae, and Thyropteridae. We use three-dimensional geometric morphometrics to investigate three questions: (1) Do insectivorous non-phyllostomid noctilionoids and primarily insectivorous generalist phyllostomids evolve along separate evolutionary trajectories? Given the similarities in their diets, fundamental differences in shape between generalist phyllostomids and non-phyllostomid noctilionoids would suggest that factors beyond diet-driven functional divergence influenced morphological change within Noctilionoidea. We then evaluate disparity and integration in the Noctilionoidea and identify patterns to suggest how these factors have shaped noctilionoid evolution. (2) Is the shape of the cranium in phyllostomids more disparate than the shape of the cranium in other noctilionoids? Phyllostomids are often assumed to be more disparate because of their tremendous ecological and apparent morphological diversity. Nevertheless, few studies have compared them to other families of bats, especially with respect to noctilionoids as a whole (Freeman 1981, 2000; Hedrick and Dumont 2018). (3) What is the relationship between disparity in cranial shape and integration within phyllostomids and within other noctilionoids? We predict that high disparity will be coupled with low integration, and that this relationship will be expressed most strongly by phyllostomids.
Materials and Methods
Sampling and Data Collection
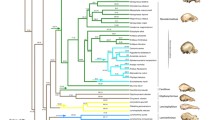
We selected bat species for study from across the New World noctilionoid tree that represent all constituent families of the superfamily Noctilionoidea (Phyllostomidae, Noctilionidae, Mormoopidae, Mystacinidae, Furipteridae, and Thyropteridae), as well as all major diet categories within the Phyllostomidae. These data included 141 individual bats from 65 species (Fig. 1). To account for individual variation, we included at least two specimens for most species. Several more common species were better sampled (e.g., for Artibeus jamaicensis, n = 5). Prior to all analyses, the mean shape of within-taxon individuals was taken and analyses were run on species rather than individuals, as in previous studies (e.g., Sherratt et al. 2017; Hedrick and Dumont 2018) due to low within-species cranial shape variation in noctilionoids (Dumont et al. 2014; Hedrick and Dumont 2018).
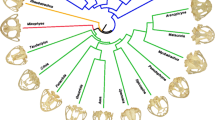
Pruned phylogeny of taxa included in the analyses from Shi and Rabosky (2015). Node colors refer to clade (as in Fig. 3a). Colored nodes indicate families (dark orange = Mystacinidae, green = Furipteridae, purple = Noctilionidae, yellow = Thyropteridae, pink = Mormoopidae, dark green = Phyllostomidae). Example cranium silhouettes shown to demonstrate high variability in cranium shape within the Noctilionoidea. Scale = 50 mm. From top to bottom (Noctilio albiventris, Mormoops sp., Macrotus waterhousii, Glossophaga soricina, Phyllostomus sp., Carollia sp., Centurio senex, Artibeus lituratus)
Bat crania were scanned using a Nikon Metrology (X-Tek) HMXST225 MicroCT system at the Center for Nanoscale Systems at Harvard University. Image stacks were generated using proprietary software associated with the X-Tek scanner (CTPro, Nikon Metrology Inc., Japan), and 3D reconstructions created in VGStudio Max 3.0 (Volume Graphics Inc., Germany), and then segmented in Mimics v. 16.0 (Materialise, Leuven, Belgium). Crania were exported as PLY files and imported into Landmark Editor (Wiley et al., 2005) to generate three-dimensional landmark configurations (PLY files to be uploaded to Morphosource). Nineteen individual landmarks and two semi-landmark curves composed of 12 and 16 semi-landmarks, respectively, were used to characterize cranium shape (Fig. 2, see ESM 1 for a list of landmarks definitions and their biological relevance).
Three-dimensional scan of Artibeus jamaicensis with landmarks overlain in (a) lateral, (b) dorsal, (c) ventral, (d) rostral, and (e) caudal views (see ESM 1 for landmark definitions). Semi-landmark curves in red. All landmarks were placed on left side, but are visualized on the right in some cases here to more effectively show the landmark position
Exploratory Analyses and Evolutionary Modeling
Landmark data were imported into RStudio v. 0.99.902 (R Core Team 2017) and opened in the geomorph package (Adams and Otárola-Castillo 2013). Data were subjected to Generalized Procrustes Analysis (GPA) with semi-landmarks slid according to the bending energy criterion (Perez et al. 2006). GPA translates, rescales, and rotates all specimens into a common shape space, minimizing all non-shape information (Zelditch et al. 2012). Following GPA, we took species means of all specimens using custom code and the mshape function in geomorph. An exploratory principal component analysis (PCA) was used to visualize how species and dietary regimes occupy morphospace. We ran an additional PCA with ancestral states and the phylogenetic tree mapped over species observations. We compared the cranium shape of insectivorous phyllostomids, non-phyllostomid insectivores, the most recent common ancestor (MRCA) of the Phyllostomidae, and the MRCA of the clade including phyllostomids and their sister family, the mormoopids.
Prior to examining the disparity and integration structure within the Noctilionoidea, taxa were generally placed into the dietary guilds following Dumont et al. (2014) to evaluate the effect of diet on cranium shape and specifically, whether non-phyllostomid noctilionoid insectivores and generalist phyllostomid insectivores grouped together. Dumont et al. (2014) split the Phyllostomidae into four adaptive regimes: nectarivores, generalists (primarily insectivorous), frugivores (stenodermatines excluding short-faced bats), and short-faced bats. Those analyses focused on adaptive regimes for mechanical advantage, particularly on the basis for the exceptionally high bite force of short-faced bats (Dumont et al. 2009). However, as we are more interested in the relationship between phyllostomids and their outgroups, we combined frugivores and short-faced bats into a fig-eating specialist category (Stenodermatinae). We also added: sanguivorous bats and non-phyllostomid insectivores. Therefore, we define five groups in total: [1] non-phyllostomid insectivores (k species = 12), [2] phyllostomid generalists (k = 23), [3] nectarivores (k = 12), [4] fig-eating specialists (k = 17), and [5] sanguivores (k = 1).
We examined the relationship between phyllostomids and their outgroups using three methods prior to evaluating integration and disparity to determine whether or not phyllostomids (particularly generalist phyllostomids) and non-phyllostomid noctilionoids evolved along different trajectories. First, a phylogenetic generalized least squares (PGLS) regression tested if phyllostomids and their outgroups had significantly different cranium shapes. Second, an exploratory canonical variates analysis (CVA) was conducted using the hypothesized adaptive regimes as groups in the R package Morpho (Schlager 2017) to establish whether the proposed groups separated based on cranium shape. We calculated the Mahalanobis distances between groups and assessed significant differences in groupings through permutation. To visualize differences between groups, canonical variate 1 (CV1) was plotted against canonical variate 2 (CV2) and a dendrogram based on Mahalanobis distances was generated. Classification success was estimated using a discriminant function analysis. Finally, evolutionary modeling analyses were then used to assess the effects of diet on cranium shape evolution.
To evaluate how noctilionoid cranium shape evolved and the nature of the relationship between the hypothesized adaptive regimes and cranium shape, we examined seven different evolutionary models (ESM 2). Sanguivores were only represented by a single taxon in this dataset, Desmodus rotundus, and hence were not included in evolutionary modeling analyses. The first model was Brownian motion (BM), which assumes that evolution follows a random walk and is not drawn to any particular optimum. This represents the null hypothesis of no directed evolutionary change. Multiple Ornstein-Uhlenbeck (OU) models were also examined. OU models describe evolutionary change in a continuous trait, in this case, principal component scores of cranium shape, as a deterministic component describing the strength of selection to one or more optima (Hansen and Houle 2008; King and Butler 2009). This optimum is characterized by θ, with α representing the strength of the attraction toward that optimum, and σ representing the intensity of random walk fluctuations. A single-peak OU model represents stabilizing selection, with change drawn to a single optimum, whereas multi-peak OU models hypothesize evolution towards different adaptive optima.
OU peaks were chosen in accordance with the four hypothesized adaptive regimes: phyllostomid generalists θG, fig-eating specialists θF, nectarivores θN, and non-phyllostomid generalists θNP. Three two-peak OU models were examined. OU2 (Fig-eating bats) split fig-eating specialists from all other groups to test whether these bats – which have been shown to have an accelerated rate of speciation relative to other bats (Dumont et al. 2011; Shi and Rabosky 2015) – represent an optimum within the Noctilionoidea while other groups converge on a single optimum (θF ≠ θN = θG = θNP). We ran a similar model for nectarivores (OU2 (Nectarivores), θN ≠ θF = θG = θNP). Nectarivorous bats have developed some of the most derived bat skull shapes and demonstrate the only instance of convergent evolution to an adaptive regime within the dataset. Finally, we examined non-phyllostomid noctilionoid insectivores (OU2 (Outgroups), θNP ≠ θN = θG = θF). Non-phyllostomid noctilionoid insectivores use oral instead of nasal echolocation, and phyllostomids may represent an adaptive radiation, unlike non-phyllostomid noctilionoid outgroups (Freeman 2000). Therefore, it is of interest to determine if phyllostomids and other noctilionoids have separate optima within Noctilionoidea. Finally, we tested our hypothesis that diet is the primary driver of shape directly with a model in which fig-eating specialists and nectarivores represent separate optima while phyllostomid generalists and non-phyllostomid insectivores converge on the same optimum (OU3, θF ≠ θN ≠ θG = θNP). This would suggest that basal phyllostomids and generalists have cranium shapes similar to other noctilionoids. We then tested if all four groups diverge to separate optima (OU4, θF ≠ θN ≠ θG ≠ θNP).
Evolutionary modeling was carried out in the R package ouch (King and Butler 2009). These models were run for PC1 and PC2 separately. Models were fit using the brown (BM) and hansen (OU) functions, and Akaike Information Criteria modified for sample size (AICc) were calculated to assess model fit.
Disparity Analyses
Disparity was calculated for the phyllostomids and their outgroups separately both by estimating the Procrustes variance of each group without phylogenetic corrections and by calculating the Procrustes distance for each taxon from the mean shape of the data using phylogenetic corrections. Significance between group disparities for Procrustes variance was assessed with 999 permutations (Zelditch et al. 2012). More permutations did not alter the significance of the test. A phylogenetic ANOVA in the package phytools was used to compare Procrustes distances among groups (Revell 2012).
Modularity and Integration Analyses
The mammalian cranium has been hypothesized to be composed of two primary modules across the mammalian tree, a rostral and a caudal module (Porto et al. 2009). This is the case even in bats that have developed cranial elaboration for nasal echolocation (Santana and Lofgren 2013). These studies suggest that rather than differences in the number of cranial modules, differences in magnitudes of integration between the two modules drive mammalian cranial variation (Porto et al. 2009). Landmarks 1, 3, 4, 5, 8, 10, 12, 16, 17, and 20–37 were included in the rostral module and landmarks 2, 6, 7, 9, 13, 14, 15, 18, 19, and 38–47 were included in the caudal module (ESM 3). Additionally, a zero-module hypothesis and two-module hypothesis were tested using EMMLi (Goswami and Finarelli 2016). EMMLi uses a maximum likelihood approach to compare module patterns using Akaike Information Criteria (AICs) rather than testing a single hypothesis of modularity a priori. We note that several other studies have found that mammal crania are made up of as many as six modules (Cheverud 1995; Goswami 2006; Goswami and Finarelli 2016). Our landmark coverage was designed to test two modules and therefore higher-level modularity hypotheses were unfortunately not possible with our dataset.
After assessing modularity and prior to comparing levels of between-module integration among groups, we ran phylogenetic partial least squares (PLS) analysis (Adams and Felice 2014) for all taxa, and for the phyllostomids and their outgroups separately. Data for the rostral block were permuted 500 times across the tips of the phylogeny and the resulting distribution was compared to the observed covariation between the rostral and caudal cranium blocks to assess significance. These data were also compared with the results from the EMMLi analysis, which assesses within- and between-module integration through the rho statistic. Following this, we compared the levels of between-module integration between the phyllostomids and their outgroups by calculating the effect sizes of the two PLS analyses using the compare.pls function in geomorph (Adams and Collyer 2016). We additionally compared the levels of between-module integration within the Phyllostomidae by comparing stenodermatines to non-stenodermatine phyllostomids. Rather than using the PLS correlation coefficient, this analysis does two-sample z-tests, which are robust to differences in sample size and number of landmarks.
The datasets generated and/or analyzed during the current study (including PLY files) are available in the Morphosource repository.
Results
Exploratory Analyses and Evolutionary Modeling
In principal component (PC) morphospace, PC 1 accounts for 43.2% of total variance and PC 2 accounts for 33.8% of total variance. PC 3 and subsequent PCs account for 5% of total variance or less and are not considered further in this study (ESMs 4, 5). PC 1 primarily divides non-phyllostomid taxa from phyllostomids (Fig. 3a) while PC 2 distinguishes different dietary modes within phyllostomids (Fig. 3b). Taxa with strongly positive PC 1 scores (e.g., Mormoops spp.) have an upturned rostrum, while taxa at the negative end of PC 1 have flattened rostra similar to that of many phyllostomid generalists (e.g., Micronycteris spp.). Thyroptera, Mystacina, Furipterus, and Noctilio all plot towards the center of morphospace between the more exaggerated rostra of mormoopids and the flattened rostra of phyllostomids (Fig. 3c). Taxa with strongly negative PC 2 scores have extremely shortened rostra as exemplified by Centurio and Sphaeronycteris. Nectarivorous taxa with elongate rostra characterize the positive end of PC 2. Non-phyllostomids all have intermediate PC 2 values with non-phyllostomids showing neither the extreme rostral elongation nor the rostral shortening found in phyllostomids. The morphospace thus represents two qualitatively different trajectories, one leading from the center of morphospace to derived mormoopids (along PC 1) and another along PC 2 dividing ecological groups within the Phyllostomidae. Finally, a PGLS of cranium shape showed that phyllostomids and their outgroups had significantly different cranium shapes (p = 0.002), but group membership explained a relatively small amount of total shape variation (R2 = 0.054; Table 1).
a Principal component analysis (PC1 x PC2) of all species (n = 65). Each point represents a single species. Species for which multiple specimens were scanned are represented by the mean of the specimens. Colors reflect node colors in Fig. 1. Note that the basal insectivorous phyllostomids (dark green) plot away from all insectivorous non-phyllostomids. Wireframes show major changes along the first principal component axis. b As Fig. 3a, but species are colored based on hypothesized dietary regimes (see branch colors in Fig. 1). c Cartoon morphospace showing overall trends in cranium shape using sample taxa. d Canonical variates analysis of major dietary regimes. All groups are fully separated. Inset shows dendrogram of Mahalanobis distances
The CVA fully distinguishes all five dietary groups in morphospace (Fig. 3d). Canonical variate (CV) 1 summarizes 41.3% of total variance and distinguishes non-phyllostomid insectivores from phyllostomid groups (fig-eating specialists, nectarivores, phyllostomid generalists, Desmodus). CV2 summarizes an additional 31.6% of total variance and distinguishes nectarivores and fig-eating specialists from Desmodus. Based on Mahalanobis distances, Desmodus is farthest away from other groups. Phyllostomid generalists, fig-eating specialists, and nectarivores are more similar to one another than to non-phyllostomid insectivores (Fig. 3d). Furthermore, a discriminant function analysis shows all five groups significantly differ from one another (ESM 6).
Seven adaptive regimes were modeled and AICcs were used to compare between models (Table 2; ESM 2). For PC1, the best-supported model is the OU4 model. The OU2 (Outgroups) model has a ΔAICc of 3.67 suggesting it was relatively well supported, but less so than the OU4 model. For PC2, the BM, OU1, OU2 (fig-eating specialists and others), and the OU2 (phyllostomids and outgroups) models are all equally well supported with OU4 being poorly supported (Table 2). The PCA, CVA, and evolutionary modeling analyses, combined, strongly suggest that while the outgroup taxa and generalist phyllostomid taxa predominantly eat insects, they have diverged in cranium shape.
Disparity Analyses
Analyses compared levels of disparity in phyllostomids and in their outgroups. Despite having fewer taxa represented (53 phyllostomids versus 12 outgroup taxa), the outgroups have significantly higher shape disparity, as measured by Procrustes variance, than phyllostomids. Indeed, shape disparity is more than three times higher among outgroup taxa (Procrustes variance = 0.0381) than phyllostomids (Procrustes variance = 0.0112, p = 0.004). In contrast, phylogenetically corrected Procrustes distance suggested that the two groups did not have significantly different levels of disparity (p = 0.139). However, Procrustes distance between the two groups is significantly different without phylogenetic corrections (p < 0.001) suggesting phylogeny plays a strong role in structuring disparity in noctilionoids. Further, phyllostomids have a mean Procrustes distance of 0.094 while outgroups have a mean Procrustes distance of 0.178 mirroring the same pattern suggested by Procrustes variance.
Modularity and Integration Analyses
The EMMLi analyses (Goswami and Finarelli 2016) suggest the crania are modular with the two-module hypothesis more strongly supported than no modularity (Table 3). EMMLi further shows between-modules integration is high (rho = 0.4), but lower than within-module integration (module 1: rho = 0.55; module 2: rho = 0.63). The rostral and caudal aspects of the cranium are significantly correlated (PLS correlation coefficient = 0.862, p < 0.001) for all species (Fig. 4a). Block one (rostral component) changes from an elongated rostrum (e.g., nectarivorous phyllostomid) to a foreshortened rostrum (e.g., short-faced bat). Block two (caudal component) displays minimal changes, but those present relate to the shortening of the cranium roof relative to the ventral aspect of the basicranium. The rostral and caudal aspects of the cranium are also significantly correlated when the outgroups are evaluated on their own (PLS corr. = 0.964, p < 0.001). In the outgroups-only analyses, changes in both the rostral and caudal aspect of the cranium relate to how upturned the rostrum is with taxa such as Mormoops spp. having more negative PLS scores for both blocks, and taxa with less upturned crania having more positive PLS scores for both blocks (Fig. 4b). Results of analysis of the phyllostomid-only dataset mirror those of the all-species between-module integration analysis in the changes that are displayed along blocks one and two (Fig. 4c). As in the other analyses, block one and two are also significantly correlated for the phyllostomids-only dataset (PLS corr. = 0.873, p < 0.001).
Phylogenetic partial least squares analysis comparing degree of integration of the rostral and caudal portions of the cranium for (a) all specimens, (b) only non-phyllostomid noctilionoids, and (c) only phyllostomids. Warp grids show gradation in shape from positive to negative values for each block. PLS correlation coefficients and p values are inset
Although the PLS correlation coefficient is higher in outgroup taxa than phyllostomids in the within-group PLS analyses, between module integration effect size is nearly twice as high in phyllostomids than in the outgroup taxa (Table 3). This difference is significant (p = 0.024). Within the Phyllostomidae, non-stenodermatine phyllostomids have a higher integration effect size (7.038) than stenodermatines (3.563). However, this difference is not significant (p = 0.27).
Discussion
Although a number of studies have examined the relationship between morphological disparity, species richness, and the magnitude of integration with theoretical modeling (Klingenberg 2005; Marroig et al. 2009; Goswami et al. 2014; Randau and Goswami 2017; Felice et al. 2018), the relationships among these factors have rarely been investigated with empirical data (but see Hu et al. 2016; Sherratt et al. 2017; Felice et al. 2018). To empirically test hypotheses regarding these relationships, we generated a 3D cranium dataset for the Phyllostomidae and outgroups within the superfamily Noctilionoidea. Phyllostomidae is one of the most species-rich families within Chiroptera (Shi and Rabosky 2015), and, with an unparalleled number of dietary types of any mammalian family, is often cited as a textbook example of an adaptive radiation (Jacobs 2016). This group therefore provides an excellent opportunity for investigating how integration, or the lack thereof, can drive such radiations. Theoretical studies have posited that increased integration leads to evolution along a single trajectory into a more limited morphospace, whereas decreased integration leads to greater morphospace occupation and thus higher disparity (Goswami et al. 2014; Felice et al. 2018). Likewise, several empirical studies have suggested high integration leads to evolutionary constraints (Young and Hallgrímsson 2005; Marroig et al. 2009; Young et al. 2010; Drake and Klingenberg 2008; Collar et al. 2014; Arlegi et al. 2018), while others have found that high levels of integration can lead to niche expansion and increases in morphological disparity (e.g., Hu et al. 2016) via evolution along a single trajectory, potentially representing a line of least evolutionary resistance. In this study, we found decreased morphological disparity and increased integration in the crania of phyllostomids relative to those of their outgroups in spite of the relatively high rates of speciation, high species richness, and greater functional breadth that characterize the Phyllostomidae (Dumont et al. 2014; Shi and Rabosky 2015).
Ecological shifts have almost certainly influenced phyllostomid evolution, as the magnitude of selection in phyllostomids has been shown to be strongest on branches leading to novel feeding habits, followed by a decrease in the magnitude of selection after novel dietary niches have been occupied (Rossoni et al. 2017). Therefore, before evaluating relative levels of integration and disparity among phyllostomids and their outgroups, we examined whether primarily insectivorous, generalist phyllostomids differed from their obligatory insectivorous sister groups in cranium shape. If this were the case, then it would suggest that factors beyond functional divergence (such as differing magnitudes of integration) played a role in the generation of the morphological differences within Noctilionoidea. We identified two qualitatively different trajectories in morphospace separating phyllostomids from their outgroups. While the crania of insectivorous phyllostomids are closest in shape to those of outgroups, they remain distinct (Fig. 3b, d), a result supported by PGLS. Furthermore, the insectivorous MRCA of the Phyllostomidae plots away from the outgroup morphospace and within the phyllostomid morphospace (ESM 7) suggesting a shift in cranium shape likely occurred as the ancestor of extant phyllostomids evolved. We also found evidence that phyllostomid generalists and their insectivorous outgroups evolved to occupy separate adaptive peaks (Table 2). Although the morphospace and adaptive peak analyses together suggest that morphological change in the Noctilionoidea has responded to other factors besides selection for dietary function, they do not elucidate what those factors may have been. To gain further insights into these factors, we analyzed the levels of integration and disparity in their crania.
Despite higher rates of speciation and greater functional breadth among phyllostomids (Dumont et al. 2014; Shi and Rabosky 2015), their crania have lower morphological disparity and greater integration than those of their outgroups (Fig. 4, Table 2). These results seem, on the surface, contrary to the observation that the Phyllostomidae represents one of the most functionally diverse chiropteran families (Freeman 2000), and that stenodermatine phyllostomids are the only clade within Chiroptera to show increased rates of speciation. We had predicted that phyllostomids would exhibit increased disparity relative to their less species-rich outgroups (although previous studies on the phyllostomid palate found high levels of integration––Sorenson et al. 2014). Other studies have found a reduction of between-limb integration and integration of cervical vertebrae in humans relative to great apes, which may have contributed to the emergence of bipedalism (Young et al. 2010; Arlegi et al. 2018), decreased limb integration in some non-quadrupedal mammals compared to some quadrupedal mammals (Young and Hallgrímsson 2005), and a decrease in magnitudes of integration in some biting eels allowing for more varied morphologies as a result of a relaxation of constraints (Collar et al. 2014).
While some authors have found evidence that reduced integration may not correlate with functional specializations (Botton-Divet et al. 2018), others have also recently found that high levels of integration can be coupled with high levels of speciation and morphological evolution (Hu et al. 2016; Sherratt et al. 2017), suggesting that increased evolutionary potential may be achieved via a single axis of evolutionary change (Schluter 1996; Marroig and Cheverud 2005; Goswami et al. 2014; Felice et al. 2018). Still others have found that high levels of integration do not necessarily limit or increase morphological evolution over long time scales (Goswami and Polly 2010). These empirical findings have been supported by some theoretical modeling studies as well. For example, Goswami et al. (2014) noted that high magnitudes of integration evoke large responses to selection, albeit along paths determined by trait covariances such that perfectly integrated structures evolve along a single axis. These authors further showed that strong integration may limit morphological disparity and can lead to the evolution of both extreme morphologies and convergence in morphology. These findings raise the question of whether a similar case might apply to phyllostomids whereby phyllostomids have reduced overall disparity in comparison with their outgroups, but have evolved extreme morphologies and multiple convergent shapes.
Cranial morphology primarily differs among phyllostomids along a single axis based on the relative length of the rostrum. Taxa with highly positive PC2 values have elongate rostra (e.g., Choeroniscus minor) and taxa with highly negative PC2 values have foreshortened rostra (e.g., Centurio senex) (Fig. 3). Morphological variance among phyllostomids along PC1 is extremely low. Instead, phyllostomids have evolved extreme elongation and extreme shortening of the rostrum relative to other bats, allowing them to access novel diets within Chiroptera. Finally, although there are not many examples of convergence within the Phyllostomidae, nectarivorous bats do converge in morphology (Datzmann et al. 2010; Rojas et al. 2016). Hence the Phyllostomidae appear to follow quite closely the predictions made by Goswami et al. (2014).
Although we now have a substantial amount of data concerning phyllostomid cranial morphology (Nogueira et al. 2009; Rossoni et al. 2017; Hedrick and Dumont 2018), the genetic and developmental mechanisms at the origin of their diversity remain poorly understood. Many candidate genes are linked to variation in face length based on mouse mutants (reviewed in Usui and Tokitam 2018) including BMP2, BMP4, and Msx1 (Bonilla-Claudio et al. 2012). In addition, some genes such as Wnt5a have been shown to directly regulate crucial stages in face formation such as the migration of the ectomesenchyme (Kaucka et al. 2016). These genes are suspected to be potential candidates for the origin of the short face for shorted-faced bats (Bonilla-Claudio et al. 2012). Recently, variation in amino-acid composition of the transcription factor Runx2 (Runt-related transcription factor 2) implicated in bone differentiation, has been correlated to variation in face-length in carnivorans, primates, and phyllostomids (Sears et al. 2007; Ritzman et al. 2017; Ferraz et al. 2018). However, this correlation remains indirect and Runx2 likely acts in concert with other factors. It is possible that changes in some of these genetic and developmental mechanisms have led to the dietary diversity and species richness in phyllostomids without decoupling the rostrum from the back of the cranium. By identifying distinct groups in terms of face-length and morphology, our study could serve as a starting point to investigate which changes in the developmental program drive face-length variation.
In addition to diet, another major difference between phyllostomids and other noctilionoids is their mode of echolocation. Phyllostomids are primarily nasal echolocators, albeit with some plasticity (Griffin and Novick 1955; Schmidt 1988), whereas non-phyllostomid noctilionoids are oral echolocators. This difference, therefore, provides a potential confounding variable in our study (Pedersen 1998). However, it appears that facial measurements do not strongly correlate with echolocation parameters when comparing nasal and oral echolocators (Goudy-Trainor and Freeman 2002). Expanding empirical data for integration magnitudes and disparity levels beyond the Noctilionoidea will be a fruitful area of future research.
This study builds upon the assessment of modularity and integration in the mammalian cranium providing support for a two-module system in comparison with a fully integrated cranium. Although the two-module hypothesis is often supported (e.g., Porto et al. 2009; Santana and Lofgren 2013), other studies have found stronger support for a six-module mammalian cranium (e.g., Cheverud 1995; Goswami 2006; Goswami and Finarelli 2016). Although we did not test hypotheses of more than two modules because there were few landmark locations, it remains possible that the internal structure of the cranium contains additional internal modules not captured in our analyses. The highly integrated external form may allow for high bite force, while decreased levels of integration of internal structures such as sensory structures (e.g., cochlea, eye globe) lead to functional divergence (Goswami and Finarelli 2016). Future work should analyze individual sensory structures in the head using a combination of both internal and external landmarks to assess whether finer-scale modules exist within the phyllostomid cranium.
In conclusion, we propose that the tightly integrated crania of phyllostomids were capable of evolving into unique dietary ecologies along a single trajectory in morphospace, which possibly represents a line of least evolutionary resistance. Although phyllostomid cranial morphology only expanded along a single axis and did not spread into as large a region of morphospace as their immediate outgroups, evolving along a single axis allowed phyllostomids to rapidly evolve exaggerated morphologies and convergent shapes capable of making use of unique diets. This study builds on many previous theoretical and empirical studies examining modularity in the mammalian cranium (Cheverud 1995; Goswami 2006; Drake and Klingenberg 2008; Porto et al. 2009; Marroig et al. 2009) and provides one of the best empirical examples of Goswami et al. (2014) and Felice et al. (2018)‘s predictions of the relationship between morphological disparity, the magnitude of integration, and evolvability, expanding our understanding of how integration, and not just function, can lead to unique morphologies as well as potentially spur adaptive radiations.
References
Adams DC, Collyer ML (2016) On the comparison of the strength of morphological integration across morphometric datasets. Evolution 70:2623–2631
Adams DC, Felice RN (2014) Assessing trait covariation and morphological integration on phylogenies using evolutionary covariance matricies. PLoS One 9(4): e94335
Adams DC, Otárola-Castillo E (2013) Geomorph: an R package for the collection and analysis of geometric morphometric shape data. Meth Ecol Evol 4:393–399.
Arlegi M, Gómez-Robles A, Gómez-Olivencia A (2018) Morphological integration in the gorilla, chimpanzee, and human neck. Am J Phys Anthropol 166:408–416
Bonilla-Claudio M, Wang J, Bai Y, Klysik E, Selever J, Martin JF (2012) Bmp signaling regulates a dose-dependent transcriptional program to control facial skeletal development. Development 139:709–719. doi:https://doi.org/10.1242/dev.073197
Botton-Divet L, Houssaye A, Herrel A, Fabre A, Cornette R (2018) Swimmers, diggers, climbers and more, a study of integration across the mustelids’ locomotor apparatus (Carnivora: Mustelidae). Evol Biol 45:182–195
Cheverud JM (1995) Morphological integration in the saddle-back tamarin cranium. Am Nat 145:63–89
Collar DC, Wainwright PC, Alfaro ME, Revell LJ, Mehta RS (2014) Biting disrupts integration to spur skull evolution in eels. Nature Comm 5:5505
Datzmann T, von Helversen O, Mayer F (2010) Evolution of nectarivory in phyllostomid bats (Phyllostomidae Gray, 1825, Chiroptera: Mammalia). BMC Evol Biol 10:165
Drake AG, Klingenberg CP (2008) The pace of morphological change: historical transformation of skull shape in St Bernard dogs. Proc R Soc B 275:71–76
Dumont ER, Davalos, LM, Goldberg, A, Santana SE, Rex K, Voigt CC (2011) Morphological innovation, diversification, and invasion of a new adaptive zone. Proc R Soc B 279:1797–1805.
Dumont ER, Herrel A, Medellin RA, Vargas-Contreras JA, Santana SE (2009) Built to bite: cranial design and function in the wrinkle-faced bat. J Zool 279:329–337
Dumont ER, Samadevam K, Grosse I, Warsi OM, Baird B, Davalos LM (2014) Selection for mechanical advantage underlies multiple cranial optima in New World leaf-nosed bats. Evolution 68:1436–1449
Felice RN, Randau M, Goswami A (2018) A fly in a tube: macroevolutionary expectations for integrated phenotypes. Evolution 72:2580–2594
Ferraz T, Rossoni DM, Althoff SL, Pissinatti A, Paixäo-Cortês VR, Cátira Bortolini M, González-José R, Marroig G, Salzano FM, Gonçalves GL, Hünemeier T (2018) Contrasting patterns of RUNX 2 repeat variations are associated with palate shape in phyllostomid bats and New World primates. Sci Rep 8:7867. https://doi.org/10.1038/s41598-018-26225-7
Freeman PW (1981) Correspondence of food habits and morphology in insectivorous bats. J Mammal 62:166–173
Freeman PW (2000) Macroevolution in Microchiroptera: recoupling morphology and ecology with phylogeny. Evol Ecol Res 2:317–335
Goswami A (2006) Cranial modularity shifts during mammalian evolution. Am Nat 168:270–280
Goswami A, Finarelli JA (2016) EMMLi: a maximum likelihood approach to the analysis of modularity. Evolution 70:1622–1637
Goswami A, Polly PD (2010) The influence of modularity on cranial morphological disparity in Carnivora and Primates (Mammalia). PLoS One 5(3):e9517
Goswami A, Smaers JB, Soligo C, Polly PD (2014) The macroevolutionary consequences of phenotypic integration: from development to deep time. Proc R Soc B 369:20130254
Goudy-Trainor A, Freeman PW (2002) Call parameters and facial features in bats: a surprising failure of form following function. Acta Chiropterol 4(1):1–16
Griffin DR, Novick A (1955) Acoustic orientation of Neotropical bats. J Exp Zool 130:251–297
Hansen TF, Houle D (2008) Measuring and comparing evolvability and constraint in multivariate characters. J Evol Biol 21(5):1201–1219
Hedrick BP, Dumont ER (2018) Putting the leaf-nosed bats in context: a geometric morphometric analysis of three of the largest families of bats. J Mammal 99(5):1042–1054
Hu Y, Ghigliotti L, Vacchi M, Pisano E, Detrich HW III, Albertson RC (2016) Evolution in an extreme environment: developmental biases and phenotypic integration in the adaptive radiation of antarctic notothenioids. BMC Evol Biol 16:142
Jacobs DS (2016) Evolution's Chimera: Bats and the Marvel of Evolutionary Adaptation. University of Cape Town Press, Cape Town
Jones KE, Bininda-Emonds ORP, Gittleman JL (2005) Bats, clocks, and rocks: diversification patterns in Chiroptera. Evolution 59:2243–2255
Kaucka M, Ivashkin E, Gyllborg D, Zikmund T, Tesarova M, Kaiser J, Xie M, Petersen J, Pachnis V, Nicolis SK, Yu T (2016) Analysis of neural crest–derived clones reveals novel aspects of facial development. Sci Adv 2. https://doi.org/10.1126/sciadv.1600060
King AA, Butler MA (2009) Ouch: Ornstein-Uhlenbeck models for phylogenetic comparative hypotheses (R package)
Kirschner M, Gerhart J (1998) Evolvability. Proc Natl Acad Sci USA 95:8420–8427
Klingenberg CP (2005) Developmental constraints, modules and evolvability. In: Hallgrímsson B, Hall BK (eds) Variation: A Central Concept in Biology. Elsevier Academic Press, San Diego, pp 219–247
Klingenberg CP (2008) Morphological integration and developmental modularity. Annu Rev Ecol Evol Syst 39:115–132
Klingenberg CP (2009) Morphometric integration and modularity in configurations of landmarks: tools for evaluating a priori hypotheses. Evol Dev 11:405–421
Klingenberg CP (2013) Cranial integration and modularity: insights into evolution and development from morphometric data. Hystrix Ital J Mammal 24:43–58
Marroig G, Cheverud JM (2005) Did natural selection or genetic drift produce the cranial diversification of Neotropical monkeys? Am Nat 163:417–428
Marroig G, Shirai LT, Porto A, de Oliveira FB, de Conto V (2009) The evolution of modularity in the mammalian skull II: evolutionary consequences. Evol Biol 36:136–148
Mitteroecker P, Bookstein F (2007) The conceptual and statistical relationship between modularity and morphological integration. Syst Biol 56:818–836
Nogueira MR, Peracchi AL, Monteiro LR (2009) Morphological correlates of bite force and diet in the skull and mandible of phyllostomid bats. Funct Ecol 23:715–723
Olson EC, Miller RL (1958) Morphological Integration. University of Chicago Press, Chicago
Pedersen SC (1998) Morphometric analysis of the chiropteran skull with regard to mode of echolocation. J Mammal 79:91–103
Perez SI, Bernal V, Gonzalez PN (2006) Differences between sliding semi-landmark methods in geometric morphometrics, with an application to human craniofacial and dental variation. J Anat 208:769–784
Porto A, de Oliveira FB, Shirai LT, de Conto V, Marroig G (2009) The evolution of modularity in the mammalian skull I: morphological integration patterns and magnitudes. Evol Biol 36:118–135
R Core Development Team (2017) Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna
Randau M, Goswami A (2017) Unraveling intravertebral integration, modularity and disparity in the Felidae (Mammalia). Evol Dev 19:85–95
Revell L (2012) Phytools: an R package for phylogenetic comparative biology (and other things). Meth Ecol Evol 3:217–223
Ritzman TB, Banovich N, Buss KP, Guida J, Rubel MA, Pinney J, Khang B, Ravosa MJ, Stone AC (2017) Facing the facts: the Runx2 gene is associated with variation in facial morphology in primates. J Hum Evol 111:139–151. https://doi.org/10.1016/j.jhevol.2017.06.014
Rojas D, Warsi OM, Dávalos LM (2016) Bats (Chiroptera: Noctilionoidea) challenge a recent origin of extant Neotropical diversity. Syst Biol 65:432–448
Rossoni DM, Assis APA, Giannini NP, Marroig G (2017) Intense natural selection preceded the invasion of new adaptive zones during the radiation of New World leaf-nosed bats. Sci Rep 7:11076
Rossoni DM, Costa BMA, Giannini NP, Marroig G (2019) A multiple peak adaptive landscape based on feeding strategies and roosting ecology shaped the evolution of cranial covariance structure and morphological differentiation in phyllostomid bats. Evolution 73:961–981
Santana SE, Lofgren SE (2013) Does nasal echolocation influence the modularity of the mammal skull? J Evol Biol 26:2520–2526
Schlager S (2017) Morpho and Rvcg–shape analysis in R: R-packages for geometric morphometrics, shape analysis and surface manipulations. In: Zheng G, Li S, Székely GJ (eds) Statistical Shape and Deformation Analysis. Academic Press, London, pp 217–256
Schluter D (1996) Adaptive radiations along genetic lines of least resistance. Evolution 50(5):1766–1774
Schluter D (2000) The Ecology of Adaptive Radiation. Oxford University Press, New York
Schmidt U (1988) Orientation and sensory functions in Desmodus rotundus. In: Greenhall AM, Schmidt U (eds) Natural History of Vampire Bats. CRC Press, Boca Raton, pp 143–166
Sears KE, Goswami A, Flynn JJ, Niswander LA (2007) The correlated evolution of Runx2 tandem repeats, transcriptional activity, and facial length in Carnivora. Evol Dev 9(6):555–565
Sherratt E, Serb JM, Adams DC (2017) Rates of morphological evolution, asymmetry and morphological integration of shell shape in scallops. BMC Evol Biol 17:248
Shi JJ, Rabosky DL (2015) Speciation dynamics during the global radiation of extant bats. Evolution 69:1528–1545
Simpson GG (1953) The Major Features of Evolution. Simon and Schuster, New York
Sorenson DW, Butkus C, Cooper LN, Cretekos CJ, Rasweiler IV JJ, Sears KE (2014) Palate variation and evolution in New World leaf-nosed and Old World fruit bats (order Chiroptera). Evol Biol 41:595–605
Usui K, Tokitam M (2018) Creating diversity in mammalian facial morphology: a review of potential developmental mechanisms. EvoDevo 9. https://doi.org/10.1186/s13227-018-0103-4
Wiley DF, Amenta N, Alcantara D A, Ghosh D, Yong JK, Delson E, Harcourt-Smith W, Rohlf FJ, St John K, Hamann B (2005) Evolutionary morphing. In VIS 05. IEEE Visualization 2005:431-438
Yoder JB, Clancey E, Des Roches S, Eastman JM, Gentry L, Godsoe W, Hagey TJ, Jochimsen D, Oswald BP, Robertson J, Sarver BAJ, Schenk JJ, Spear SF, Harmon LJ (2010) Ecological opportunity and the origin of adaptive radiations. J Evol Biol 23:1581–1596
Young NM, Hallgrímsson B (2005) Serial homology and the evolution of mammalian limb covariation structure. Evolution 59:2691–2704
Young NM, Wagner GP, Hallgrímsson B (2010) Development and the evolvability of human limbs. Proc Natl Acad Sci USA 107:3400–3405
Zelditch ML, Swiderski DL, Sheets HD, Fink WL (2012) Geometric Morphometrics for Biologists: A Primer, 2nd edn. Elsevier Academic Press, London
Acknowledgements
We would like to thank Greg Lin and Jim Reynolds for access to the μCT scanner used in this study and for technical support. This work was performed in part at the Center for Nanoscale Systems (CNS), a member of the National Nanotechnology Coordinated Infrastructure Network, which is supported by the National Science Foundation under award number ECS 1541959. We also thank Daniel Pulaski for logistical support and Abby Vander Linden, Andy Conith, Sam Cordero, the Pierce lab for discussions. Further, we thank the editor (John Wible) and two anonymous reviewers for their comments improving the quality of this manuscript. B.P.H. was supported in part by the National Science Foundation PRFB-1612211, L.M.D. and S.J.R. by DEB-1442142, K.E.S. by DEB-1442314, and E.R.D. by DEB-1442278.
Author information
Authors and Affiliations
Contributions
BPH carried out CT scanning. BPH, GLM, and VDM segmented and landmarked 3D models. BPH carried out statistical analyses. BPH, GLM, VDM, AS, KTJD, SJR, KES, LMD, and ED participated in design of the study and helped draft the manuscript. BPH, ED, and LMD conceived of the study, designed the study, and coordinated the study. All authors gave final approval for publication.
Corresponding author
Ethics declarations
Conflict of Interest
No authors declare a conflict of interest.
Additional information
Surface scans used in this manuscript will be archived on Morphosource. All other data are given in the manuscript and its supplementary material.
Electronic supplementary material
ESM 1
Landmark numbers, definitions, and reasons each landmark was selected (refer to Fig. 2). (XLSX 9 kb)
ESM 3
Visualization of two-module hypothesis showing which landmarks were included in the rostral module (red) and which landmarks were included in the caudal module (blue). (PNG 1119 kb)
ESM 4
Scree plot demonstrating dominance of PC1 and PC2. (PNG 32 kb)
ESM 5
Table summarizing each taxon’s centroid size, family, hypothesized dietary regime, as well as the first four principal component scores, the CAC, and the first four residual shape component scores. (XLSX 21 kb)
ESM 6
CVA classification matrix results. Mahalanobis distances between each hypothesized dietary regime (above the diagonal) and significance (below the diagonal). Classification matrix showing the percent of each group that were successfully classified. Note that the blood group could not be included here given that it was based on a single taxon (Desmodus rotundus). Also note that two species (one nectarivore and one phyllostomid generalist) were outliers and not classified into any group. (XLSX 9 kb)
ESM 7
Principal components 1 and 2 with phylogeny and ancestral states plotted on top of morphospace highlighting the positions of mormoopids (dark blue) and basal insectivorous phyllostomids (dark green). Note that the phyllostomid MRCA plots within the phyllostomid morphospace and away from the non-phyllostomid morphospace. (PNG 196 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Hedrick, B.P., Mutumi, G.L., Munteanu, V.D. et al. Morphological Diversification under High Integration in a Hyper Diverse Mammal Clade. J Mammal Evol 27, 563–575 (2020). https://doi.org/10.1007/s10914-019-09472-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10914-019-09472-x