Abstract
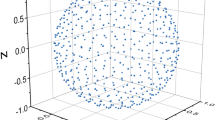
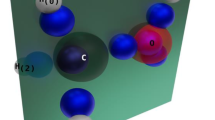
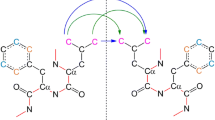
We propose an improved algorithm for calculating Avnir’s continuous symmetry and chirality measures of molecules. These measures evaluate the deviation of a given structure from symmetry by calculating the distance between the structure and its nearest symmetric counterpart. Our new algorithm utilizes structural properties of the given molecule to increase the accuracy of the calculation and dramatically reduce the running time by up to tens orders of magnitude. Consequently, a wide variety of molecules of medium size with ca. 100 atoms and even more can be analyzed within seconds. Numerical evidence of the algorithm’s efficiency is presented for several families of molecules such as helicenes, porphyrins, dendrimers building blocks, fullerene and more. The ease and efficiency of the calculation make the continuous symmetry and chirality measures promising descriptors for integration in quantitative structure–activity relationship tools, as well as chemical databases and molecular visualization software.









Similar content being viewed by others
References
M. Petitjean, Entropy 5, 271–312 (2003)
P.G. Mezey, Mol. Phys. 104, 723–729 (2006)
P.G. Mezey, J. Math. Chem. 45, 544–549 (2009)
P.G. Mezey, K. Fukui, S. Arimoto, K. Taylor, Int. J. Quantum Chem. 66, 99–105 (1998)
G.M. Crippen, Curr. Comput. Aided Drug Des. 4, 259–264 (2008)
H. Zabrodsky, S. Peleg, D. Avnir, J. Am. Chem. Soc. 114, 7843–7851 (1992)
M. Pinsky, C. Dryzun, D. Casanova, P. Alemany, D. Avnir, J. Comput. Chem. 29, 2712–2721 (2008)
H. Zabrodsky, D. Avnir, J. Am. Chem. Soc. 117, 462–473 (1995)
S. Alvarez, P. Alemany, D. Avnir, Chem. Soc. Rev. 34, 313–326 (2005)
M. Pinsky, D. Avnir, Inorg. Chem. 37, 5575–5582 (1998)
S. Alvarez, Dalton Trans. 13, 2209–2233 (2005)
S. Alvarez, P. Alemany, D. Casanova, J. Cirera, M. Llunell, D. Avnir, Coord. Chem. Rev. 249, 1693–1708 (2005)
D. Yogev-Einot, D. Avnir, Tetrahedron Asymmetry 17, 2723–2725 (2006)
C. Dryzun, Y. Mastai, A. Shvalb, D. Avnir, J. Mater. Chem. 19, 2062–2069 (2009)
I. Tuvi-Arad, D. Avnir, J. Math. Chem. 47, 1274–1286 (2010)
I. Tuvi-Arad, D. Avnir, J. Org. Chem. 76, 4973–4979 (2011)
I. Tuvi-Arad, D. Avnir, Chem. Eur. J. 18, 10014–10020 (2012)
I. Tuvi-Arad, T. Rozgonyi, A. Stirling, J. Phys. Chem. A 117, 12726–12733 (2013)
I. Tuvi-Arad, A. Stirling, Isr. J. Chem. 56, 1067–1075 (2016)
M. Bonjack-Shterengartz, D. Avnir, Proteins Struct. Funct. Bioinf. 83, 722–734 (2015)
S. Keinan, D. Avnir, J. Am. Chem. Soc. 122, 4378–4384 (2000)
M.H. Jamroz, J.E. Rode, S. Ostrowski, P.F.J. Lipinski, J.C. Dobrowolski, J. Chem. Inf. Model. 52, 1462–1479 (2012)
D. Milner, S. Raz, H. Hel-Or, D. Keren, E. Nevo, Pattern Recogn. 40, 2237–2250 (2007)
I. Saragusti, I. Sharon, O. Katzenelson, D. Avnir, J. Archaeol. Sci. 25, 817–825 (1998)
R. Iovita, I. Tuvi-Arad, M.H. Moncel, J. Despriee, P. Voinchet, J.J. Bahain, Plos One 12, e0177063 (2017)
C. Dryzun, A. Zait, D. Avnir, J. Comput. Chem. 32, 2526–2538 (2011)
O. Katzenelson, J. Edelstein, D. Avnir, Tetrahedron Asymmetry 11, 2695–2704 (2000)
Molecular Operating Environment (MOE), Chemical Computing Group Inc, 1010 Sherbooke St. West, Suite #910, Montreal, QC, Canada, H3A 2R7 (2016)
C.R. Groom, I.J. Bruno, M.P. Lightfoot, S.C. Ward, Acta Cryst. B72, 171–179 (2016)
J.-J. Zhang, J. Glaser, S.A. Gamboa, A. Lachgar, J. Chem. Crystallogr. 39, 1–8 (2009)
X.-L. Wang, D.-N. Liu, H.-Y. Lin, N. Han, G.-C. Liu, J. Inorg. Organomet. Polym Mater. 25, 671–679 (2015)
Z. Yi, X. Yu, W. Xia, L. Zhao, C. Yang, Q. Chen, X. Wang, X. Xu, X. Zhang, CrystEngComm 12, 242–249 (2010)
C. du Peloux, A. Dolbecq, P. Mialane, J. Marrot, F. Secheresse, Dalton Trans. 1259–1263 (2004). doi:10.1039/B401250J
H.-Y. Ma, L.-Z. Wu, H.-J. Pang, X. Meng, J. Peng, J. Mol. Struct. 967, 15–19 (2010)
M.P. Byrn, C.J. Curtis, Y. Hsiou, S.I. Khan, P.A. Sawin, S.K. Tendick, A. Terzis, C.E. Strouse, J. Am. Chem. Soc. 115, 9480–9497 (1993)
V.E. de Oliveira, C.C. Corrêa, C.B. Pinheiro, R. Diniz, L.F.C. de Oliveira, J. Mol. Struct. 995, 125–129 (2011)
P. Dastidar, I. Goldberg, Acta Crystallogr. Sect. C 52, 1976–1980 (1996)
S. McGill, V.N. Nesterov, S.L. Gould, Acta Crystallogr. Sect. E 69, m471 (2013)
L. Abbassi, Y.M. Chabre, N. Kottari, A.A. Arnold, S. Andre, J. Josserand, H.-J. Gabius, R. Roy, Polym. Chem. 6, 7666–7683 (2015)
W. C. Marsh, J. Trotter, J. Chem. Soc. A, 161–173 (1971). doi:10.1039/J19710000169
H.A. Alidağı, Ö.M. Gırgıç, Y. Zorlu, F. Hacıvelioğlu, S.Ü. Çelik, A. Bozkurt, A. Kılıç, S. Yeşilot, Polymer 54, 2250–2256 (2013)
H. R. Allcock, S. Al-Shali, D. C. Ngo, K. B. Visscher, M. Parvez, J. Chem. Soc. Dalton Trans. 3549–3559 (1996). doi:10.1039/DT9960003549
Y. Tümer, H. Bati, N. Çalişkan, Çd Yüksektepe, O. Büyükgüngör, Zeitschrift für anorganische und allgemeine Chemie 634, 597–599 (2008)
Acknowledgements
Supported by the Israel Science Foundation (Grant 411/15). We are sincerely grateful for fruitful discussions with Prof. David Avnir (The Hebrew University of Jerusalem). The programming of the new code was done by Itay Zandbank and Devora Witty (The scientific software company, Israel). We are thankful to Sagiv Barhoom (The Open University) for his help in programming and Yaffa Shalit (The Open University) for her help in testing the code. Researchers interested in using the CSM code are welcome to contact Dr. Tuvi-Arad.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Funding
This study was funded by the Israel Science Foundation (Grant Number 411/15).
Appendix: Proof of formulas (3) and (4)
Appendix: Proof of formulas (3) and (4)
For the sake of completeness, we present a rigorous proof of formulas (3) and (4). Given a permutation \(\pi \) of \(\{1,2,\ldots ,N\}\) satisfying \(\pi ^{n}=id\), we look at the minimum of \(\mathop {\sum }\limits _{k=1}^N {\left| {\mathbf{Q}_k -\mathbf{P}_k } \right| ^{2}} \) over all the \(\pi \)-symmetric structures \(\left\{ {\mathbf{P}_k } \right\} \). Let \(C_1 ,C_2 ,\ldots ,C_f \) be the cycles of \(\pi \), as in Eq. (8). Given the symmetry operation T, we have
where the minimum is over all the structures \(\left\{ {\mathbf{P}_k } \right\} \) satisfying Eq. (2).
For each cycle \(C_j \) and \(k_0 \in C_j \), we have
where l is a divisor of n. We therefore have (as T is an isometry)
Therefore, the minimum in (13) can be found by minimizing the expression in (15) separately for each cycle, and summing over j.
It is a well known and easy to prove fact that given vectors \(\mathbf{x}_1 ,\ldots ,\mathbf{x}_m \), the vector \(\mathbf{x}\) which minimizes the sum \(\sum _{k=1}^m {\left| {\mathbf{x}_k -\mathbf{x}} \right| ^{2}} \) is the average, \(\mathbf{x}=\frac{1}{m}\mathop {\sum }\limits _{k=1}^m {\mathbf{x}_k } \). The minimum value is
Therefore, the minimum of (15) is attained when
Since \(\pi ^{l}(k_0 )=k_0 \), and by (2), the summand in Eq. (17) is periodic in u with period l. Since l divides n, we conclude that
This proves Eq. (4).
By Eq. (16), the minimum value of Eq. (15) is
Summing over j, we get (3).
Rights and permissions
About this article
Cite this article
Alon, G., Tuvi-Arad, I. Improved algorithms for symmetry analysis: structure preserving permutations. J Math Chem 56, 193–212 (2018). https://doi.org/10.1007/s10910-017-0788-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10910-017-0788-y