Abstract
Biallelic null or hypomorphic variants in JAK3 cause SCID and less frequently Omenn syndrome. We investigated homozygous hypomorphic JAK3 mutations in two patients, and expression and function of a novel JAK3R431P variant in Omenn syndrome. Immunophenotyping of PBMC from the patient with the novel JAK3R431P variant was undertaken, by flow cytometry and Phosflow after stimulation with IL-2, IL-7, and IL-15. JAK3 expression was investigated by Western blotting. We report two patients with homozygous hypomorphic JAK3 variants and clinical features of Omenn syndrome. One patient had a previously described JAK3R775H variant, and the second had a novel JAK3R431P variant. One patient with a novel JAK3R431P variant had normal expression of JAK3 in immortalised EBV-LCL cells but reduced phosphorylation of STAT5 after stimulation with IL-2, IL-7, and IL-15 consistent with impaired kinase activity. These results suggest the JAK3R431P variant to be hypomorphic. Both patients are alive and well after allogeneic haematopoietic stem cell transplantation. They have full donor chimerism, restitution of thymopoiesis and development of appropriate antibody responses following vaccination. We expand the phenotype of hypomorphic JAK3 deficiency and demonstrate the importance of functional testing of novel variants in disease-causing genes.
Similar content being viewed by others
Introduction
Severe combined immunodeficiency (SCID) usually results from recessive loss of function variants in genes involved in T-lymphocyte development, and is characterised immunologically by absent or low and dysfunctional T-cells, and variably affected B- or NK-cell number and function, depending on the pathway affected [1, 2]. Patients with typical SCID present in the first year of life with failure to thrive, diarrhea, and recurrent infections, including opportunistic infections such as Pneumocystis jirovecii pneumonia [2]. Absence of T-cells may also allow the transplacental engraftment of maternally-derived T-cells, which are typically oligoclonal and alloreactive and manifest as graft-versus-host disease (GvHD) with lichenified erythroderma, alopecia, diarrhea, and lymphadenopathy [3]. These clinical features are shared with Omenn syndrome (OS), which results from an incomplete block of T-cell development. Low numbers of autologous T-cells emerge from the thymus and show oligoclonal expansion with end-organ infiltration and damage. Since Omenn’s first description in an Irish family in 1965 [4], the syndrome has expanded to include T-lymphocytosis with an activated phenotype and oligoclonal expansion of TCRVβ families, raised serum IgE, peripheral eosinophilia, and absent or reduced T-cell proliferative response to antigens [5]. OS is predominantly associated with hypomorphic mutations in recombination-activating genes 1 and 2 (RAG1/2) [6], but has also been described in patients with mutations in DCLRE1C [7] and, less commonly, IL7RA [8]. All of these genes play critical roles during thymocyte development but are more or less redundant for mature T-cell effector function. Other genes implicated in SCID include those affecting expression of proteins involved in transducing TCR and cytokine signals, such as IL2RG and JAK3. Omenn syndrome as a manifestation of JAK3 SCID has only been reported in one previous patient [9].
Null or hypomorphic variants in JAK3 are associated with SCID of T-B + NK- phenotype [10], similar to X-linked SCID caused by loss of function of IL2RG, encoding the common gamma chain of type I cytokine receptors [11]. As a member of the Janus kinase family of proteins, JAK3 transduces signals from interleukins IL-2, IL-4, IL-7, IL-9, IL-15, and IL-21, critical for regulating lymphocyte expansion and differentiation [12,13,14].
We report the presentation and clinical course of two unrelated patients with OS, each of whom bore different homozygous mutations in JAK3. We demonstrate that the novel JAK3R431P variant protein is normally expressed, however its signalling function in response to interleukin stimulation is diminished. Our findings emphasise the genetic heterogeneity of OS, the phenotypic spectrum associated with hypomorphic JAK3 deficiency and the importance of functional analysis to clarify the significance of novel missense variants in known disease genes.
Methods
Clinical data were collected retrospectively from medical records. Both patients’ families gave written consent for approved research including data collection, analysis, and publication. (REC reference 20/NE/0044).
Sample Collection and Generation of Patient’s Primary EBV-LCL Cell Line
Peripheral blood mononuclear cells (PBMC) were isolated from EDTA blood samples using Lymphoprep (StemCell Technologies, 07851) density gradient centrifugation as per manufacturer’s instructions.
EBV-LCL cell lines were generated by immortalization of isolated PBMC by infecting the cells with EBV supernatant produced by the B95-8 marmoset cell, and cyclosporine A (1ug/ml). Cells were subcultured in RPMI-1640 culture medium (Sigma Aldrich, R0883) supplemented with 10% (v/v) foetal calf serum (FCS, Gibco, 10270–106), 1% (v/v) Penicillin/Streptomycin (100 U/mL and 100 μg/mL respectively; Sigma Aldrich, P0781) and 1% (v/v) L-Glutamine (2 mM; Sigma Aldrich, G7513), referred as RPMI10.
Immunophenotyping by Flow Cytometry
PBMC were thawed at 37°C, transferred into pre-warmed complete RPMI10 culture medium, pelleted by centrifugation, resuspended in RPMI10, and left to rest at 37˚C for 1.5 h. Cells were stained with a cocktail of cell surface antibodies in FACS buffer (PBS + 2% FCS) for 30min at room temperature (RT) in the dark. Cells were washed and resuspended in FACS buffer, 7AAD viability dye (Biolegend) was added, and cells were acquired on a BD FACSymphony A5 flow cytometer (BD Biosciences). Data were analysed by FlowJo V10 (BD Biosciences). The following anti-human flow cytometry antibodies were used: TCR-gd-FITC (B1, Biolegend), CD19-PerCP-Cy5.5 (HIB19, Biolegend), CD14-PE (M5E2, Biolegend), CD56-PE-CF594 (NCAM16.2, BD Biosciences), CCR7-PE-Cy7 (G043H7, Biolegend), PD-1-AF700 (EH12.2H7, Biolegend), CD45RO-APC-Cy7 (UCHL1, BD Biosciences), CD127-BV421 (A019D5, Biolegend), CD28-BV480 (CD28.2, BD Biosciences), CD20-BV605 (L27, BD Biosciences), CD16-BV650 (3G8, BD Biosciences), HLA-DR-BV711 (L243, Biolegend), CD4-BV786 (SK3, Biolegend), CD3-BUV395 (UCHT1, BD Biosciences), CD8-BUV496 (RPA-T8, BD Biosciences), CD25-BUV737 (2A3, BD Biosciences).
Phosphoflow Cytometry
PBMC and EBV-LCL cells were rested in serum free RPMI culture media for 2hr or overnight in incubator at 37°C, respectively. Cells were stained with fixable viability dye Zombie yellow (Biolegend), CD16-BV421 (3G8, BD Biosciences), CD20-BV510 (2H7, Biolegend) and CD8-BUV496 (RPA-T8, BD Biosciences) for 30 minutes. Following the pre-staining, cells were divided into the appropriate number of FACS tubes and stimulated separately with IL-2, IL-7, IL-15 (all 100ng/ml, ThermoFisher Scientific), and IL-21 (50ng/ml, Miltenyi Biotec) for 15min at 37˚C. Cells were fixed 1:1 using Cytofix buffer (BD Biosciences) for 20min at RT. Permeabilization was achieved by adding ice-cold PermIII buffer (BD Biosciences), and incubation on ice for 20 min. After repeated washing steps with FACS buffer, cells were stained with antibodies: pSTAT1-AF488 (pY701, 4a, BD Biosciences), CD19-PerCP-Cy5.5 (HIB19, Biolegend), pSTAT3-PE (pY705, 4/P-STAT3, BD Biosciences), FOXP3-PE-Dazzle594 (206D, Biolegend), pSTAT5-AF647 (pY694, 47/Stat5(pY694), BD Biosciences), CD56-BV711 (5.1H11, Biolegend), CD4-BV786 (SK3, Biolegend), CD3-BUV395 (UCHT1, BD Biosciences) and CD25-BUV737 (2A3, BD Biosciences) for 60 min at RT in the dark. The staining panel for EBV-LCL cells was reduced to pSTAT antibodies, Zombie Yellow, and B cell markers CD19 and CD20. Samples were acquired on a BD FACSymphony A5 flow cytometer (BD Biosciences) and analyzed using FlowJo software. For data analysis, mean fluorescence intensity of unstimulated cells was subtracted from values after interleukin stimulation.
JAK3 Protein Expression by Western Blotting
EBV-LCL cells were washed in PBS and lysed in lysis buffer for 15min [50 mM Tris–HCl (pH 7.5), 150 mM NaCl, 1% Nonidet P-40, 0.1% SDS, 0.5% Na-Deoxycholate] containing 100 mM dithiothreitol (Merck), 1 × complete protease inhibitor cocktail (Roche), 1 × PhosSTOP phosphatase inhibitors (Roche), and 1 × NuPAGE Loading Buffer (Thermo Fisher Scientific). Lysates were denatured at 95°C for 10 min before being subjected to 4–12% tris–glycine polyacrylamide gel (Thermo Fisher Scientific) electrophoresis in 1 × SDS NuPAGE MOPS Running Buffer (Thermo Fisher Scientific). Proteins were transferred to 0.45-mm Immobilon®-P polyvinyl difluoride membranes (Thermo Fisher Scientific) in 1 × NuPAGE Tris–Glycine Transfer Buffer supplemented with 20% Methanol. Membranes were blocked for 60 min in 5% bovine serum albumin in tris-buffered saline with 0.1% Tween (TBS-T) buffer before overnight immunostaining at 4°C with mouse anti-human JAK3 (Cell Signaling, 5481S, 1:1000) and rabbit anti-human GAPDH antibodies (Cell Signaling, 8884S, 1:5000). Membranes were washed in TBS-T and stained with secondary antibodies anti-mouse IgG HRP-linked (Cell Signaling, 7076S, 1:3000) and anti-rabbit IgG HRP-linked (Cell Signaling, 7074S, 1:5000) for 1h at RT. Membranes were developed with Immobilon ECL Ultra Western Substrate solution (Merck), imaged on a LI-COR Odyssey Fc (LI-COR) and Image Studio software was used for analysis.
Statistical Analysis
For experiments with multiple repeats, data are expressed as the mean ± standard deviation. Where appropriate, statistical comparisons between Controls and Patient P2 data were calculated using unpaired t-test carried out through GraphPad Prism software with probability values (P) of P<0.05 designated significant.
Results
Omenn Syndrome-Like Patients with JAK3 Mutation
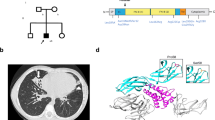
Patient 1 (P1), a male infant, was born at 40 weeks gestation following a normal pregnancy to consanguineous parents. There was no family history of immunodeficiency. Aged 8 weeks, he developed oral candidiasis, chronic diarrhoea, and recurrent chest infections, leading to faltering growth. He received immunisations as per schedule including Bacille Calmette-Guérin (BCG) vaccination. He subsequently developed alopecia with an erythematous, lichenified rash and axillary lymphadenopathy. Initial immunological investigations revealed a total lymphocytosis (11,060 cells/microlitre) with low CD3 + CD8 + T-cells (296 cells/microlitre) and absent naïve T-cells but preserved NK cells numbers (Table 1). There was no maternofoetal engraftment on cytogenetic testing. Flow phenotyping for TCRVβ usage showed expansion of certain subsets and absence of others, consistent with oligoclonality, and his lymphocytes had impaired proliferation following incubation with phytohemagglutinin. Immunoglobulin concentrations were low (IgA: 0.05g/L, IgM: 0.27g/L, IgG < 0.3g/L) except IgE, which was elevated (640kU/L). Virological testing showed rhinovirus and parainfluenza 4 in nasopharyngeal secretions, and sapovirus in his faeces. Due to the dermatological features of OS, he was commenced on topical tacrolimus and clobetasone 0.05%, and oral rifampicin, isoniazid and ethambutol were given for BCGosis.
At 6 months of age, P1 underwent a bone marrow transplant from an HLA-identical sibling, following conditioning with treosulfan (36g/m2), fludarabine (150mg/m2) and serotherapy with alemtuzumab (1mg/kg). He received ciclosporin A and mycophenolate mofetil as GvHD prophylaxis. Peri-transplant, his course was complicated by engraftment pneumonitis which required oxygen, methylprednisolone, and a dose of infliximab prior to resolution, along with immune reconstitution against BCG presenting as axillary lymphadenitis requiring antimycobacterial therapy and incision and drainage. He received pre-emptive treatment with ganciclovir between days + 39 and + 70 post-HSCT for cytomegalovirus (CMV) viraemia. Two years post-transplant, whole exome sequencing of the patient’s germline (fibroblast) DNA revealed a homozygous pathogenic variant (c.2324G > A, p.R775H) in JAK3. This variant results in substitution of arginine for histidine at position 775 in the JH2 pseudokinase domain of the JAK3 protein, and has been reported in two other patients [9, 15]. No other pathogenic variants were identified in known IEI genes. Now 10 years post-transplant, P1 is well, off immunoglobulin and infection-free with normal lymphocyte subsets and stable mixed donor chimerism (CD3 + : 95%, CD19 + : 27%, CD15 + : 9%). At latest clinical review, P1’s alopecia has resolved and he has normal skin with no features of OS, and has had antimicrobial prophylaxis discontinued.
Patient 2 (P2), a female infant, was born at 40 weeks gestation as the first child to consanguineous parents of Syrian origin, with no family history of immunodeficiency. She did not receive BCG vaccination at birth. She presented aged 3 weeks with a persistent maculopapular rash which developed into erythroderma unresponsive to emollients, and initial investigations identified eosinophilia and total lymphocytosis (9,037 cells/microlitre) despite low CD3 + CD8 + T-cells (69 cells/microlitre) and absent naïve T-cells (Table 1). NK and B cell numbers were within normal limits. T-cell receptor (TCR) staining identified expansion of the TCRVβ-5.1, -14, and -20 subsets with several absent Vβ subsets. Maternofoetal engraftment studies were negative. Prior to HSCT, she required treatment with ganciclovir for CMV and human herpesvirus-6 (HHV6) viraemia. She had persistent fever despite broad-spectrum antibiotics and antifungal treatment, which resolved following intravenous methylprednisolone and serotherapy with anti-thymocyte globulin. Aged 5 months, P2 underwent HSCT using her 12/12 HLA-matched maternal grandfather as donor. She received a peripheral blood stem cell graft, following conditioning with treosulfan (30g/m2), fludarabine (150mg/m2), and alemtuzumab (1mg/kg), and GvHD prophylaxis with ciclosporin A. Four weeks post-infusion, she developed a left hemiparesis; investigations revealed CSF lymphocytosis and DNA polymerase chain reaction was positive for CMV DNA. Magnetic resonance imaging showed evidence of small vessel vasculopathy, consistent with CMV meningoencephalitis. She was treated with foscarnet followed by ganciclovir with neurological recovery and was discharged home. Her post-transplant course was further complicated by the development of autoimmune haemolytic anaemia, requiring treatment with intravenous immunoglobulin, prednisolone, and sirolimus. Twenty-six months post-HSCT, she has discontinued immunosuppression. She has high mixed donor chimerism (CD3 + : 90%, CD19 + : 32%, CD15 + : 25%), has stopped immunoglobulin replacement, and has made good responses to primary vaccination. At latest follow-up, her skin is normal with no features of OS, and she remains on azithromycin prophylaxis only. Neurologically, she displays delayed development of expressive language with evolving spastic diplegia.
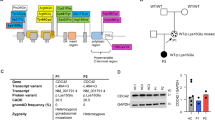
Her latest lymphocyte subset analysis is summarised in Table 1. Whole exome sequencing identified a homozygous variant of unknown significance in JAK3 (NM_0002153) c.1292G > C, p.Arg431Pro in the SH2 domain. This variant has a CADD score of 22.1 and is absent from the gnomAD population database (Supplementary Figure S1). Comparison of VARITY score for the R431P variant against homozygous JAK3 variants present in gnomAD v4 shows it has the highest predicted pathogenicity for variants in its protein domain (Supplementary Figure S2). The allele balance for this variant was 1.0, excluding somatic reversion mosaicism. No other pathogenic variants in known IEI genes were identified on whole exome sequencing.
JAK3R431P Mutation is Associated with Diminished T-Cells and Raised B-Cells in Peripheral Blood
Immunophenotyping of two healthy controls and patient P2’s peripheral blood cells by flow cytometry indicated significantly reduced T- and elevated B-cells in P2. Among her T-cells, CD8 + cells and circulating naïve CD4 + cells (CD45RO-CCR7 +) were absent while the frequency of memory CD4 + T-cells was increased. Moreover, we observed a high percentage of circulating activated CD4 + cells in the patient, expressing HLA-DR and/or PD-1. We also noted slightly decreased frequencies of TCRγδ + cells and Treg (CD4 + CD127lowCD25 +). Analysis of NK-cells revealed skewing of CD56lowCD16 + NK-cells toward CD56-CD16 + phenotype, which can be associated with high viral load (Fig. 1A, B) [16,17,18].
Immunophenotyping of healthy control and patient P2’s peripheral blood cells by flow cytometry. A Contour flow plots showing relative abundance of B-cells and reduced T-cells, absence of CD8 + cells and naïve CD4 + cells, hyper activation of CD4 + cells, and skewing of NK-cells towards CD56-CD16 + phenotype in patient P2 with the JAK3R431P variant. B Quantification of flow cytometry data in controls and patient P2. Data presented as mean ± SD
Preserved Expression of Mutated JAK3R431P Protein
To further assess the impact of the variant, we measured JAK3 protein expression in P2-derived EBV-LCL cell line by immunoblotting. Expression of the variant JAK3 was equivalent to wild-type JAK3 expressed in EBV-LCL cells derived from healthy control donor (Fig. 2A).
Impaired pSTAT5 response to interleukin IL-2, IL-7, and IL-15 stimulation in patient P2’s PBMC subsets. A Unaffected expression of JAK3R431P protein in EBV-LCL cell lines by immunoblotting. B Histograms of pSTAT5 in patient P2’s and healthy controls’ (C1, C2) peripheral CD4 cells in response to IL-2, IL-7, and IL-15 stimulation. Dashed area: IL-stimulation, empty line: unstimulated. C Impaired pSTAT5 response to interleukin IL-2, IL-7, and IL-15 stimulation in patient P2’s cell subsets. C: Controls C1 and C2, P: patient P2, pSTAT5: phosphorylated STAT5. Data presented as mean ± SD
Hypomorphic Effect of JAK3R431P Variant on its Kinase Activity and JAK-STAT Signalling after Interleukin Stimulation
To further assess any potential hypomorphic effect of this mutation on JAK3 kinase activity, we stimulated PBMC isolated from patient P2 and unaffected individuals with interleukins IL-2, IL-7, and IL-15, and measured their response by downstream STAT5 phosphorylation. The phosphorylation of STAT5 was reduced in patient P2 lymphocytes and NK-cells after stimulation with all three interleukins, IL-2, IL-7, and IL-15, suggesting an impaired kinase activity of normally expressed mutated JAK3R431P protein (Fig. 2B, C; Supplementary Figure S3).
The observation of impaired kinase activity of mutated JAK3 protein was also shown in EBV-LCL cell lines derived from patient P2 and healthy controls. The magnitude of response (MFI) measured by STAT5 phosphorylation after IL-2 and IL-7 stimulation in all EBV-LCL cell lines was too low to draw clear conclusions. However, stimulation with IL-15 suggested a trend towards impaired signalling in patient P2’s EBV-LCL cell line, although in lower magnitude than in data obtained from PBMC (Supplementary Figure S4).
Discussion
We report two patients with hypomorphic JAK3 deficiency presenting with an OS phenotype and preserved NK-cell production. The JAK3 variant in P1 (R775H) has been previously reported in two patients. The first report describes a female from Iran with failure to thrive, chronic diarrhoea, and respiratory distress [9]. This patient’s presentation was notable for diffuse erythroderma affecting > 50% of body surface area with acute-on-chronic inflammation extending from the upper to the deep dermis, as well as alopecia. Immunophenotyping demonstrated normal total lymphocyte count, reduced T cell receptor excision circle numbers, and impaired T-lymphocyte proliferation to phytohemagglutinin. However, populations of naïve CD4 + and CD8 + T cells were reportedly normal. Maternal lymphocytes were identified in the patient’s blood at a frequency of 17%, indicative of maternofetal engraftment and thus SCID. The authors demonstrate that the identified R775H variant in JAK3 is likely to be deleterious, with a low population allele frequency and high predictive scores for pathogenicity using in silico models.
Maternofetal engraftment denotes SCID due to an absence of functional neonatal T cells to identify and reject allogeneic cells. Clinically, maternofetal engraftment and OS are indistinguishable, reflecting their shared pathology of “self”-reactive T cells causing organ damage. Analysis of short tandem repeats from patient lymphocytes is necessary to identify the presence or absence of maternal DNA, and diagnose or exclude maternofetal engraftment. In our patient with the R775H variant, no maternofetal engraftment was identified.
A further case of SCID with this variant was reported by Firtina et al., though it is unclear if this patient had an OS phenotype [15]. The R775H variant is in the pseudokinase domain of JAK3. This domain regulates catalytic activity of the kinase domain and therefore modulates its signal transduction activity; mutations in the pseudokinase domain result in normal JAK3 protein expression but a failure to transduce cytokine-dependent signals, possibly due to increased inhibition of kinase activity [19].
Analysis of the novel R431P JAK3 variant in P2 demonstrated normal expression of the resulting protein. Flow cytometry data indicated that the patient’s JAK3-STAT3 signalling is impaired but not absent in response to IL-2, IL-7, and IL-15 stimulation. This is consistent with the variant’s position in the SH2 domain of JAK3, which is implicated in cytokine receptor binding [20]. Feasibly, differential defects in JAK-STAT signalling downstream of alternative receptors could be contributing to the immunological phenotype, as has been documented for certain hypomorphic TYK2 variants [21]. For example, IL-15 is not only linked to NK cell development [22], but also CD8 + memory T-cell homeostasis [23]. Its role in maintaining a CD8 + T-cell population has been shown as crucial, potentially contributing to the patient’s CD8 negative status [24,25,26]. Impaired IL-7 signalling has been linked to atypical JAK3-SCID with preserved NK cell production and impaired T cell function [12, 27].
Defects in signal transduction downstream of IL-7 and IL-15 are understood to lead to the T- and NK- immunophenotype seen in both JAK3- and IL2RG-SCID due to their role in thymic development, differentiation, and expansion of T and NK lymphocytes [12, 22, 25, 28]. Both patients presented with normal total T-lymphocyte counts but extremely elevated CD4:CD8 ratio due to CD8 + lymphopenia, which has normalised following HSCT. Unlike in complete JAK3 deficiency, both patients demonstrated normal NK cell numbers, suggesting the partial kinase activity conferred by their hypomorphic proteins may be sufficient to allow NK cell development.
Rarely, reversion mutations of pathogenic genes back to wild-type may mitigate the deleterious impact of a germline mutation within a somatic clone of that cell line, leading to reversion mosaicism [29]. This may occur through a true “back” mutation, or via a compensatory second-site mutation, and is described in several inborn errors of immunity including SCID caused by pathogenic variants in ADA, IL2RG, RAG1, and CD3Z, leading to preserved T-cell numbers and a milder phenotype [29]. Reversion mosaicism has been described in two siblings with hypomorphic JAK3 SCID, one showing reversion mosaicism in both CD4 + and CD8 + T-cells, and one with revertant CD8 + T-cells only [30]. Somatic reversion was excluded in our patient P2 by identification of an allelic balance of 1.0 for the R431P variant.
OS is classically associated with mutations in RAG1 and RAG2, which impair V(D)J recombination and result in expansion of an oligoclonal population of T-cells [31]. However, OS has also been described in other genetic aetiologies that impair lymphocyte development, including IL7RA, IL2RG, ADA, DCLRE1C, RMRP and LIG4 deficiencies, and CHARGE syndrome [7, 8, 32,33,34,35,36,37]. It remains rare and is estimated to represent 5% of cases of SCID, and is typically diagnosed later than classical SCID [38]. The presence of OS was historically associated with a poorer outcome at HSCT, though more recently, a report by the Primary Immune Deficiency Treatment Consortium stratifying SCID patients by ‘typical’ or ‘atypical’ (including OS) features found no impact on survival, rates of GvHD, or immune reconstitution post-HSCT [39, 40]. Historic outcomes for OS patients may relate to pre-HSCT immunosuppression or a tendency for more myeloablative conditioning regimens compared to ‘typical’ SCID: in a previous Primary Immune Deficiency Treatment Consortium study, 46% of patients with ‘atypical’ SCID received myeloablative chemotherapy compared to 19% with ‘typical’ SCID [39]. Today, this added myeloablation typically comes from addition of thiotepa to treosulfan/fludarabine, or use of pharmacokinetically-dosed busulfan [41]. Patients may also require pre-HSCT ciclosporin A or alemtuzumab to reduce organ infiltration by autoreactive T-cells and reduce the risk of GvHD arising from an inflamed milieu at the point of graft infusion.
In conclusion, we report two homozygous hypomorphic JAK3 variants causing SCID with OS and demonstrate impaired signal transduction by the JAK3R431P variant despite normal protein expression. We suggest that JAK3 defects that disrupt but do not abrogate lymphocyte development may lead to OS, similarly to other genetic aetiologies of OS that do not directly impact V(D)J recombination.
Data Availability
The datasets generated during and analysed during the current study are not available for sharing, but the corresponding author may be contacted with queries.
Abbreviations
- BCG:
-
Bacille Calmette-Guérin
- C:
-
Control
- CMV:
-
Cytomegalovirus
- EBV-LCL:
-
Epstein-Barr Virus-bearing lymphoblastoid cell line
- FCS:
-
Foetal calf serum
- GvHD:
-
Graft-versus-host disease
- HHV6:
-
Human herpesvirus-6
- Ig:
-
Immunoglobulin
- IL:
-
Interleukin
- JAK:
-
Janus kinase
- MFI:
-
Mean fluorescence intensity
- OS:
-
Omenn syndrome
- P:
-
Patient
- PBMC:
-
Peripheral blood mononuclear cells
- RAG:
-
Recombination-activating gene
- RPMI medium:
-
Roswell Park Memorial Institute medium
- SCID:
-
Severe combined immunodeficiency
- TCR:
-
T-cell receptor
References
Tangye SG, Al-Herz W, Bousfiha A, Chatila T, Cunningham-Rundles C, Etzioni A, et al. Human Inborn Errors of Immunity: 2019 Update of the IUIS Phenotypical Classification. J Clin Immunol. 2020;40(1):66–81.
Chinn IK, Shearer WT. Severe Combined Immunodeficiency Disorders. Immunol Allergy Clin North Am. 2015;35(4):671–94 (Available from: https://doi.org/10.1016/j.iac.2015.07.002).
Müller SM, Ege M, Pottharst A, Schulz AS, Schwarz K, Friedrich W. Transplacentally acquired maternal T lymphocytes in severe combined immunodeficiency: A study of 121 patients. Blood. 2001;98(6):1847–51.
Omenn GS. Familial Reticuloendotheliosis with Eosinophilia. N Engl J Med. 1965;273(8):427–32 (Available from: http://www.ncbi.nlm.nih.gov/pubmed/14328107).
Shearer WT, Dunn E, Notarangelo LD, Dvorak CC, Puck JM, Logan BR, et al. Establishing diagnostic criteria for severe combined immunodeficiency disease (SCID), leaky SCID, and Omenn syndrome: The Primary Immune Deficiency Treatment Consortium experience. J Allergy Clin Immunol. 2014;133(4):1092–8 (Available from: https://linkinghub.elsevier.com/retrieve/pii/S0091674913014954).
Villa A, Sobacchi C, Notarangelo LD, Bozzi F, Abinun M, Abrahamsen TG, et al. V(D)J recombination defects in lymphocytes due to RAG mutations: severe immunodeficiency with a spectrum of clinical presentations. Blood. 2001;97(1):81–8.
Ege M, Ma Y, Manfras B, Kalwak K, Lu H, Lieber MR, et al. Omenn syndrome due to ARTEMIS mutations. Blood. 2005;105(11):4179–86.
Giliani S, Bonfim C, De Saint BG, Lanzi G, Brousse N, Koliski A, et al. Omenn syndrome in an infant with IL7RA gene mutation. J Pediatr. 2006;148(2):272–4.
Shahbazi Z, Parvaneh N, Shahbazi S, Rahimi H, Hamid M, Shahbazi D, et al. Graft versus host disease and microchimerism in a JAK3 deficient patient. Allergy, Asthma Clin Immunol. 2019;15(1):1–9 (Available from: https://doi.org/10.1186/s13223-019-0361-2).
Notarangelo LD, Mella P, Jones A, de Saint BG, Savoldi G, Cranston T, et al. Mutations in severe combined immune deficiency (SCID) due to JAK3 deficiency. Hum Mutat. 2001;18(4):255–63.
Goldberg L, Simon AJ, Lev A, Barel O, Stauber T, Kunik V, et al. Atypical immune phenotype in severe combined immunodeficiency patients with novel mutations in IL2RG and JAK3. Genes Immun. 2020;21(5):326–34.
Li J, Nara H, Rahman M, Juliana FM, Araki A, Asao H. Impaired IL-7 signaling may explain a case of atypical JAK3-SCID. Cytokine. 2010;49(2):221–8.
Brugnoni D, Notarangelo LD, Sottini A, Airò P, Pennacchio M, Mazzolari E, et al. Development of autologous, oligoclonal, poorly functioning T lymphocytes in a patient with autosomal recessive severe combined immunodeficiency caused by defects of the Jak3 tyrosine kinase. Blood. 1998;91(3):949–55.
Candotti F, Oakes SA, Johnston JA, Giliani S, Schumacher RF, Mella P, et al. Structural and functional basis for JAK3-deficient severe combined immunodeficiency. Blood. 1997;90(10):3996–4003.
Firtina S, Yin Ng Y, Hatirnaz Ng O, Kiykim A, Aydiner E, Nepesov S, et al. Mutational landscape of severe combined immunodeficiency patients from Turkey. Int J Immunogenet. 2020;47(6):529–38.
Milush JM, López-Vergès S, York VA, Deeks SG, Martin JN, Hecht FM, et al. CD56negCD16+ NK cells are activated mature NK cells with impaired effector function during HIV-1 infection. Retrovirology. 2013;10:158.
Caduff N, McHugh D, Rieble L, Forconi CS, Ong’echa JM, Oluoch PO, et al. KSHV infection drives poorly cytotoxic CD56-negative natural killer cell differentiation in vivo upon KSHV/EBV dual infection. Cell Rep. 2021;35(5):109056.
Farnault L, Chambost H, Michel G, Thuret I, de Saint BG, Fischer A, et al. Persistence of natural killer cells with expansion of a hypofunctional CD56-CD16+KIR+NKG2C+ subset in a patient with atypical Janus kinase 3-deficient severe combined immunodeficiency. J Allergy Clin Immunol. 2013;131:1230–3 (United States).
Chen M, Cheng A, Candotti F, Zhou YJ, Hymel A, Fasth A, et al. Complex effects of naturally occurring mutations in the JAK3 pseudokinase domain: evidence for interactions between the kinase and pseudokinase domains. Mol Cell Biol. 2000;20(3):947–56.
Ferrao R, Lupardus PJ. The Janus Kinase (JAK) FERM and SH2 Domains: Bringing Specificity to JAK-Receptor Interactions. Front Endocrinol (Lausanne). 2017;8:71.
Boisson-Dupuis S, Ramirez-Alejo N, Li Z, Patin E, Rao G, Kerner G, et al. Tuberculosis and impaired IL-23-dependent IFN-γ immunity in humans homozygous for a common TYK2 missense variant. Sci Immunol. 2018;3(30):eaau8714.
Kennedy MK, Glaccum M, Brown SN, Butz EA, Viney JL, Embers M, et al. Reversible defects in natural killer and memory CD8 T cell lineages in interleukin 15-deficient mice. J Exp Med. 2000;191(5):771–80.
Zeng R, Spolski R, Finkelstein SE, Oh S, Kovanen PE, Hinrichs CS, et al. Synergy of IL-21 and IL-15 in regulating CD8+ T cell expansion and function. J Exp Med. 2005;201(1):139–48.
Johnston JA, Bacon CM, Finbloom DS, Rees RC, Kaplan D, Shibuya K, et al. Tyrosine phosphorylation and activation of STAT5, STAT3, and Janus kinases by interleukins 2 and 15. Proc Natl Acad Sci U S A. 1995;92(19):8705–9.
Yajima T, Yoshihara K, Nakazato K, Kumabe S, Koyasu S, Sad S, et al. IL-15 regulates CD8+ T cell contraction during primary infection. J Immunol. 2006;176(1):507–15.
Tripathi P, Kurtulus S, Wojciechowski S, Sholl A, Hoebe K, Morris SC, et al. STAT5 is critical to maintain effector CD8+ T cell responses. J Immunol. 2010;185(4):2116–24.
Khanolkar A, Wilks JD, Liu G, Simpson BM, Caparelli EA, Kirschmann DA, et al. A case of aberrant CD8 T cell-restricted IL-7 signaling with a Janus kinase 3 defect-associated atypical severe combined immunodeficiency. Immunol Res. 2020;68(1):13–27.
O’Shea JJ, Husa M, Li D, Hofmann SR, Watford W, Roberts JL, et al. Jak3 and the pathogenesis of severe combined immunodeficiency. Mol Immunol. 2004;41(6):727–37 (Available from: https://www.sciencedirect.com/science/article/pii/S0161589004001312).
Miyazawa H, Wada T. Reversion Mosaicism in Primary Immunodeficiency Diseases. Front Immunol. 2021;12:783022.
Ban SA, Salzer E, Eibl MM, Linder A, Geier CB, Santos-Valente E, et al. Combined immunodeficiency evolving into predominant CD4+ lymphopenia caused by somatic chimerism in JAK3. J Clin Immunol. 2014;34(8):941–53.
Marrella V, Maina V, Villa A. Omenn syndrome does not live by V(D)J recombination alone. Curr Opin Allergy Clin Immunol. 2011;11(6):525–31.
Roifman CM, Gu Y, Cohen A. Mutations in the RNA component of RNase mitochondrial RNA processing might cause Omenn syndrome. J Allergy Clin Immunol. 2006;117(4):897–903.
Ibusuki A, Nishikawa T, Hiraki T, Okano T, Imai K, Kanegane H, et al. Prominent dermal Langerhans cells in an Omenn syndrome patient with a novel mutation in the IL2RG gene. J Dermatol. 2019;46(11):1019–23.
Joshi AY, Ham EK, Shah NB, Dong X, Khan SP, Abraham RS. Atypical Omenn Syndrome due to Adenosine Deaminase Deficiency. Case reports Immunol. 2012;2012:919241.
Gennery AR, Slatter MA, Rice J, Hoefsloot LH, Barge D, McLean-Tooke A, et al. Mutations in CHD7 in patients with CHARGE syndrome cause T-B + natural killer cell + severe combined immune deficiency and may cause Omenn-like syndrome. Clin Exp Immunol. 2008;153(1):75–80.
Gennery A. Recent advances in understanding RAG deficiencies. F1000Research. 2019;8:148.
Grunebaum E, Bates A, Roifman CM. Omenn syndrome is associated with mutations in DNA ligase IV. J Allergy Clin Immunol. 2008;122:1219–20 (United States).
Dvorak CC, Haddad E, Heimall J, Dunn E, Cowan MJ, Pai SY, et al. The diagnosis of severe combined immunodeficiency: Implementation of the PIDTC 2022 Definitions. J Allergy Clin Immunol. 2023;151(2):547-555.e5 (Available from: https://doi.org/10.1016/j.jaci.2022.10.021).
Haddad E, Logan BR, Griffith LM, Buckley RH, Parrott RE, Prockop SE, et al. SCID genotype and 6-month posttransplant CD4 count predict survival and immune recovery. Blood. 2018;132(17):1737–49.
Gennery AR, Slatter MA, Grandin L, Taupin P, Cant AJ, Veys P, et al. Transplantation of hematopoietic stem cells and long-term survival for primary immunodeficiencies in Europe: Entering a new century, do we do better? J Allergy Clin Immunol. 2010;126(3):602–10.
Lankester AC, Albert MH, Booth C, Gennery AR, Güngör T, Hönig M, et al. EBMT/ESID inborn errors working party guidelines for hematopoietic stem cell transplantation for inborn errors of immunity. Bone Marrow Transplant. 2021;56(9):2052–62 (Available from: https://doi.org/10.1038/s41409-021-01378-8).
Acknowledgements
We acknowledge all study participants, patients and their families, healthy control donors, and clinical team looking after patients. We thank Newcastle University Flow Cytometry and Genomics Core Facilities for their assistance.
Funding
This work was supported by the Wellcome Trust (grant number207556_Z_17_Z, to Professor Sophie Hambleton).
Author information
Authors and Affiliations
Contributions
Christo Tsilifis: cared for the patients, provided clinical data, and wrote the manuscript.
Jarmila Stremenova Spegarova: designed and performed experiments, analysed data, and wrote the manuscript.
Ross Good: performed experiments and analysed data.
Helen Griffin: performed bioinformatics analyses.
Karin Engelhardt: helped with bioinformatics analyses.
Sophie Graham: cared for the patients and provided clinical data.
Stephen Hughes: cared for the patients, provided clinical data.
Peter Arkwright: cared for the patients, provided clinical data.
Sophie Hambleton: cared for the patients, provided clinical data, designed experiments, and revised the manuscript.
Andrew R. Gennery: cared for the patients, provided clinical data, designed experiments, and revised the manuscript.
All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
SH declares research funding from Pharming and Miltenyi and speaker honoraria/consultancy fees from Pharming, Takeda, CSL Behring and Videregen. AG declares research funding from Mallinckrodt and JAZZ Pharmaceuticals and speaker honoraria/consultancy fees from Pharming and Miltenyi Biotec.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Christo Tsilifis and Jarmila Stremenova Spegarova are Joint first authors.
Sophie Hambleton and Andrew R. Gennery are Joint senior authors.
Supplementary Information
Below is the link to the electronic supplementary material.
10875_2024_1699_MOESM1_ESM.pdf
Supplementary file1 Supplemental Figure S1. Correlation of Combined Annotated Dependent Depletion (CADD) scores with minor allele frequencies (MAF) for the JAK3 variants identified in this report and other homozygous JAK3 variants in gnomAD v4 (PDF 6 KB)
10875_2024_1699_MOESM2_ESM.pdf
Supplementary file2 Supplemental Figure S2. High predicted pathogenicity for the JAKR431P variant compared to other JAK3 homozygous variants in its protein domain using the VARITY score (PDF 7 KB)
10875_2024_1699_MOESM3_ESM.pdf
Supplementary file3 Supplemental Figure S3. Impaired pSTAT5 response to interleukin IL-2, IL-7, and IL-15 stimulation in patient P2’s cell subsets. C: Controls C1 and C2, P: patient P2, pSTAT5: phosphorylated STAT5. Data presented as mean ± SD (PDF 66 KB)
10875_2024_1699_MOESM4_ESM.pdf
Supplementary file4 Supplemental Figure S4. Phospho-STATs signalling after stimulation with IL-2, IL-7, IL-15, and IL-21 in EBV-LCL cell lines showing impaired response of patient P2’s cells. A) Representative flow cytometry histograms of pSTAT5 stimulation in patient P2 and controls (C1, C2). Dashed area: IL-stimulation, empty line: unstimulated. B) Quantification of three independent flow cytometry experiments. Data presented as mean ± SD, ns: non-significant, unpaired t-test (PDF 87 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsilifis, C., Spegarova, J.S., Good, R. et al. Omenn Syndrome in Two Infants with Different Hypomorphic Variants in Janus Kinase 3. J Clin Immunol 44, 98 (2024). https://doi.org/10.1007/s10875-024-01699-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10875-024-01699-5