Abstract
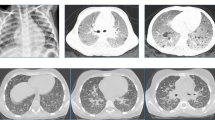
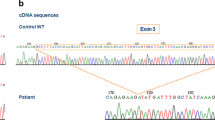
Chronic granulomatosis disease (CGD) is a rare inborn error of immunity, characterized by phagocytic respiratory outbreak dysfunction. Mutations causing CGD occur in CYBB on the X chromosome and in the autosomal genes CYBA, NCF1, NCF2, NCF4, RAC2, and CYBC1. Nevertheless, some patients are clinically diagnosed with CGD, due to abnormal respiratory outbursts, while the pathogenic gene mutation is unidentified. Here, we report a patient with CGD who first presented with Bacillus Calmette-Guérin disease and had recurrent pneumonia. He was diagnosed with CGD by nitro blue tetrazolium and respiratory burst tests. Detailed assessment of neutrophil activity revealed that patient neutrophils were almost entirely nonfunctional. Sanger sequencing detected a 6-kb insertion of a LINE-1 transposable element in the third intron of CYBB, leading to abnormal splicing and pseudoexon insertion, as well as introduction of a premature termination codon, resulting in predicted protein truncation. Clonal analysis demonstrated that the patient had somatic mosaicism, and the phagocytes were almost all variant CYBB, while the mosaicism rate of PBMC was about 65%. Finally, deep RNA sequencing and gp91phox expression analysis confirmed the pathogenicity of the mutation. In conclusion, we demonstrate that insertion of a LINE-1 transposon in a CYBB intron was responsible for CGD in our patient. Intron LINE-1 transposon element insertion should be examined in CGD patients without any known disease-causing gene mutation, in addition to identification of new genes.





Similar content being viewed by others
Data Availability
The dataset described in this study is available from the corresponding authors upon reasonable request.
Code Availability
Not applicable.
Change history
20 January 2023
A Correction to this paper has been published: https://doi.org/10.1007/s10875-023-01435-5
References
Yu HH, Yang YH, Chiang BL. Chronic granulomatous disease: a comprehensive review. Clin Rev Allergy Immunol. 2021;61(2):101–13. https://doi.org/10.1007/s12016-020-08800-x.
Roos D, van Leeuwen K, Hsu AP, Priel DL, Begtrup A, Brandon R, et al. Hematologically important mutations: X-linked chronic granulomatous disease (fourth update). Blood Cells Mol Dis. 2021;90: 102587. https://doi.org/10.1016/j.bcmd.2021.102587.
Rosenzweig SD. Inflammatory manifestations in chronic granulomatous disease (CGD). J Clin Immunol. 2008;28(Suppl 1):S67-72. https://doi.org/10.1007/s10875-007-9160-5.
Holland SM. Chronic granulomatous disease. Clin Rev Allergy Immunol. 2010;38(1):3–10. https://doi.org/10.1007/s12016-009-8136-z.
Arnadottir GA, Norddahl GL, Gudmundsdottir S, Agustsdottir AB, Sigurdsson S, Jensson BO, et al. A homozygous loss-of-function mutation leading to CYBC1 deficiency causes chronic granulomatous disease. Nat Commun. 2018;9(1):4447. https://doi.org/10.1038/s41467-018-06964-x.
Thomas DC, Charbonnier LM, Schejtman A, Aldhekri H, Coomber EL, Dufficy ER, et al. EROS/CYBC1 mutations: decreased NADPH oxidase function and chronic granulomatous disease. J Allergy Clin Immunol. 2019;143(2):782–5 e1. https://doi.org/10.1016/j.jaci.2018.09.019
Neehus A, Moriya K, Nieto-Patlán A, Le Voyer T, Lévy R, Özen A, et al. Impaired respiratory burst contributes to infections in PKCδ-deficient patients. The Journal of experimental medicine 2021;218(9). https://doi.org/10.1084/jem.20210501
Neehus A, Tuano K, Le Voyer T, Nandiwada S, Murthy K, Puel A, et al. Chronic granulomatous disease-like presentation of a child with autosomal recessive PKCδ deficiency. J Clin Immunol. 2022. https://doi.org/10.1007/s10875-022-01268-8.
Wang S, Wang T, Xiang Q, Xiao M, Cao Y, Xu H, et al. Clinical and molecular features of chronic granulomatous disease in mainland China and a XL-CGD female infant patient after prenatal diagnosis. J Clin Immunol. 2019;39(8):762–75. https://doi.org/10.1007/s10875-019-00680-x.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921. https://doi.org/10.1038/35057062.
Symer D, Connelly C, Szak S, Caputo E, Cost G, Parmigiani G, et al. Human l1 retrotransposition is associated with genetic instability in vivo. Cell. 2002;110(3):327–38. https://doi.org/10.1016/s0092-8674(02)00839-5.
Cordaux R, Batzer MA. The impact of retrotransposons on human genome evolution. Nat Rev Genet. 2009;10(10):691–703. https://doi.org/10.1038/nrg2640.
Payer LM, Burns KH. Transposable elements in human genetic disease. Nat Rev Genet. 2019;20(12):760–72. https://doi.org/10.1038/s41576-019-0165-8.
Roos D, de Boer M. Retrotransposable genetic elements causing neutrophil defects. Eur J Clin Invest. 2018;48(Suppl 2): e12953. https://doi.org/10.1111/eci.12953.
Luo X, Liu Q, Jiang J, Tang W, Ding Y, Zhou L, et al. Characterization of a cohort of patients with LIG4 deficiency reveals the founder effect of p.R278L, unique to the Chinese population. Front Immunol. 2021;12:695993. https://doi.org/10.3389/fimmu.2021.695993
Sottini A, Serana F, Bertoli D, Chiarini M, Valotti M, Vaglio Tessitore M, et al. Simultaneous quantification of T-cell receptor excision circles (TRECs) and K-deleting recombination excision circles (KRECs) by real-time PCR. J Vis Exp. 2014(94). https://doi.org/10.3791/52184
Bylund J, Bjornsdottir H, Sundqvist M, Karlsson A, Dahlgren C. Measurement of respiratory burst products, released or retained, during activation of professional phagocytes. Methods Mol Biol. 2014;1124:321–38. https://doi.org/10.1007/978-1-62703-845-4_21.
Khandagale A, Lazzaretto B, Carlsson G, Sundin M, Shafeeq S, Römling U, et al. JAGN1 is required for fungal killing in neutrophil extracellular traps: implications for severe congenital neutropenia. J Leukoc Biol. 2018;104(6):1199–213. https://doi.org/10.1002/jlb.4a0118-030rr.
Tangye SG, Gray PE, Pillay BA, Yap JY, Figgett WA, Reeves J, et al. Hyper-IgE syndrome due to an elusive novel intronic homozygous variant in DOCK8. J Clin Immunol. 2022;42(1):119–29. https://doi.org/10.1007/s10875-021-01152-x.
Miyano K, Sumimoto H. Assessment of the role for Rho family GTPases in NADPH oxidase activation. Methods Mol Biol. 2012;827:195–212. https://doi.org/10.1007/978-1-61779-442-1_14.
Russell SM, Tayebi N, Nakajima H, Riedy MC, Roberts JL, Aman MJ, et al. Mutation of Jak3 in a patient with SCID: essential role of Jak3 in lymphoid development. Science. 1995;270(5237):797–800. https://doi.org/10.1126/science.270.5237.797.
Ding Y, Zhou L, Xia Y, Wang W, Wang Y, Li L, et al. Reference values for peripheral blood lymphocyte subsets of healthy children in China. J Allergy Clin Immunol. 2018;142(3):970–3 e8. https://doi.org/10.1016/j.jaci.2018.04.022
Kwok JSY, Cheung SKF, Ho JCY, Tang IWH, Chu PWK, Leung EYS, et al. Establishing simultaneous T cell receptor excision circles (TREC) and K-deleting recombination excision circles (KREC) quantification assays and laboratory reference intervals in healthy individuals of different age groups in Hong Kong. Front Immunol. 2020;11:1411. https://doi.org/10.3389/fimmu.2020.01411.
Zhang Z, Xin D, Wang P, Zhou L, Hu L, Kong X, et al. Noisy splicing, more than expression regulation, explains why some exons are subject to nonsense-mediated mRNA decay. BMC Biol. 2009;7:23. https://doi.org/10.1186/1741-7007-7-23.
Bousfiha A, Jeddane L, Picard C, Al-Herz W, Ailal F, Chatila T, et al. Human Inborn errors of immunity: 2019 update of the IUIS phenotypical classification. J Clin Immunol. 2020;40(1):66–81. https://doi.org/10.1007/s10875-020-00758-x.
Ishikawa T, Okai M, Mochizuki E, Uchiyama T, Onodera M, Kawai T. Bacillus Calmette-Guérin (BCG) infections at high frequency in both AR-CGD and X-CGD patients following BCG vaccination. Clin Infect Dis. 2021;73(9):e2538–44. https://doi.org/10.1093/cid/ciaa1049.
Richardson SR, Doucet AJ, Kopera HC, Moldovan JB, Garcia-Perez JL, Moran JV. The influence of LINE-1 and SINE retrotransposons on mammalian genomes. Microbiol Spectr. 2015;3(2):MDNA3–0061–2014. https://doi.org/10.1128/microbiolspec.MDNA3-0061-2014
Wimmer K, Callens T, Wernstedt A, Messiaen LJPg. The NF1 gene contains hotspots for L1 endonuclease-dependent de novo insertion. 2011;7(11):e1002371. https://doi.org/10.1371/journal.pgen.1002371
Hancks DC, Kazazian HH Jr. Roles for retrotransposon insertions in human disease. Mob DNA. 2016;7:9. https://doi.org/10.1186/s13100-016-0065-9.
Kazazian HH Jr, Moran JV. Mobile DNA in health and disease. N Engl J Med. 2017;377(4):361–70. https://doi.org/10.1056/NEJMra1510092.
Beck CR, Collier P, Macfarlane C, Malig M, Kidd JM, Eichler EE, et al. LINE-1 retrotransposition activity in human genomes. Cell. 2010;141(7):1159–70. https://doi.org/10.1016/j.cell.2010.05.021.
Kumatori A, Faizunnessa N, Suzuki S, Moriuchi T, Kurozumi H, Nakamura M. Nonhomologous recombination between the cytochrome b558 heavy chain gene (CYBB) and LINE-1 causes an X-linked chronic granulomatous disease. Genomics. 1998;53(2):123–8. https://doi.org/10.1006/geno.1998.5510.
Meischl C, Boer M, Ahlin A, Roos D. A new exon created by intronic insertion of a rearranged LINE-1 element as the cause of chronic granulomatous disease. Eur J Hum Genet. 2000;8(9):697–703. https://doi.org/10.1038/sj.ejhg.5200523.
Brouha B, Meischl C, Ostertag E, de Boer M, Zhang Y, Neijens H, et al. Evidence consistent with human L1 retrotransposition in maternal meiosis I. Am J Hum Genet. 2002;71(2):327–36. https://doi.org/10.1086/341722.
de Boer M, van Leeuwen K, Geissler J, Weemaes CM, van den Berg TK, Kuijpers TW, et al. Primary immunodeficiency caused by an exonized retroposed gene copy inserted in the CYBB gene. Hum Mutat. 2014;35(4):486–96. https://doi.org/10.1002/humu.22519.
Gentsch M, Kaczmarczyk A, van Leeuwen K, de Boer M, Kaus-Drobek M, Dagher M, et al. Alu-repeat-induced deletions within the NCF2 gene causing p67-phox-deficient chronic granulomatous disease (CGD). 2010;31(2):151–8. https://doi.org/10.1002/humu.21156
Yamada M, Okura Y, Suzuki Y, Fukumura S, Miyazaki T, Ikeda H, et al. Somatic mosaicism in two unrelated patients with X-linked chronic granulomatous disease characterized by the presence of a small population of normal cells. Gene. 2012;497(1):110–5. https://doi.org/10.1016/j.gene.2012.01.019.
Muotri AR, Chu VT, Marchetto MC, Deng W, Moran JV, Gage FH. Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature. 2005;435(7044):903–10. https://doi.org/10.1038/nature03663.
Kano H, Godoy I, Courtney C, Vetter M, Gerton G, Ostertag E, et al. L1 retrotransposition occurs mainly in embryogenesis and creates somatic mosaicism. Genes Dev. 2009;23(11):1303–12. https://doi.org/10.1101/gad.1803909.
Richardson S, Gerdes P, Gerhardt D, Sanchez-Luque F, Bodea G, Muñoz-Lopez M, et al. Heritable L1 retrotransposition in the mouse primordial germline and early embryo. J Genome research. 2017;27(8):1395–405. https://doi.org/10.1101/gr.219022.116.
Aluri J, Cooper M. Genetic mosaicism as a cause of inborn errors of immunity. J Journal of clinical immunology. 2021;41(4):718–28. https://doi.org/10.1007/s10875-021-01037-z.
Mensa-Vilaró A, Bravo García-Morato M, de la Calle-Martin O, Franco-Jarava C, Martínez-Saavedra M, González-Granado L, et al. Unexpected relevant role of gene mosaicism in patients with primary immunodeficiency diseases. J The Journal of allergy clinical immunology. 2019;143(1):359–68. https://doi.org/10.1016/j.jaci.2018.09.009.
Al-Mousa H, Abouelhoda M, Monies DM, Al-Tassan N, Al-Ghonaium A, Al-Saud B, et al. Unbiased targeted next-generation sequencing molecular approach for primary immunodeficiency diseases. J Allergy Clin Immunol. 2016;137(6):1780–7. https://doi.org/10.1016/j.jaci.2015.12.1310.
Postel M, Culver J, Ricker C, Craig D. Transcriptome analysis provides critical answers to the “variants of uncertain significance” conundrum. Hum Mutat. 2022. https://doi.org/10.1002/humu.24394.
Acknowledgements
We thank the patient and his family for participating in this study.
Funding
This work was supported by the Graduate Mentor Team of Chongqing Municipal Education Commission (Grant number 2019–09-66), the National Natural Science Foundation of China (Grant number 82070135), and the CQMU Program for Youth Innovation in Future Medicine (W0100).
Author information
Authors and Affiliations
Contributions
XD.Z and YF.A designed this study, and reviewed and revised the manuscript. L.Yu collected clinical data, followed the patient, performed experiments, and drafted the manuscript. WH.L., G.L., G.S., L.Yang, JJ.C., and LN.Z. performed some experiments. Y.D., ZY.Z., and XM.T. provided essential help for clinical management and follow-up of the patient. All authors contributed to the final version of the manuscript and approved submission of the final version.
Corresponding authors
Ethics declarations
Ethics Approval
The study was performed following the Declaration of Helsinki and approved by the ethics committee of Children’s Hospital of Chongqing Medical University (Chongqing, China). Written informed consent for participation in this study was obtained from patient’s parents.
Consent to Participate
Written informed consent was obtained from patient’s parents.
Consent for Publication
Written informed consent for publication of the study was obtained from patient’s parents. All the authors approved the final version of the manuscript.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Yunfei An was not captured as the co-corresponding author.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yu, L., Li, W., Lv, G. et al. De Novo Somatic Mosaicism of CYBB Caused by Intronic LINE-1 Element Insertion Resulting in Chronic Granulomatous Disease. J Clin Immunol 43, 88–100 (2023). https://doi.org/10.1007/s10875-022-01347-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-022-01347-w