Abstract
We report the updated classification of Inborn Errors of Immunity/Primary Immunodeficiencies, compiled by the International Union of Immunological Societies Expert Committee. This report documents the key clinical and laboratory features of 430 inborn errors of immunity, including 64 gene defects that have either been discovered in the past 2 years since the previous update (published January 2018) or were characterized earlier but have since been confirmed or expanded upon in subsequent studies. The application of next-generation sequencing continues to expedite the rapid identification of novel gene defects, rare or common; broaden the immunological and clinical phenotypes of conditions arising from known gene defects and even known variants; and implement gene-specific therapies. These advances are contributing to greater understanding of the molecular, cellular, and immunological mechanisms of disease, thereby enhancing immunological knowledge while improving the management of patients and their families. This report serves as a valuable resource for the molecular diagnosis of individuals with heritable immunological disorders and also for the scientific dissection of cellular and molecular mechanisms underlying inborn errors of immunity and related human diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Inborn errors of immunity, also referred to as primary immunodeficiencies, manifest as increased susceptibility to infectious diseases, autoimmunity, autoinflammatory diseases, allergy, and/or malignancy. These conditions are caused by monogenic germline mutations that result in loss of expression, loss-of-function (LOF; amorphic/hypomorphic), or gain-of-function (GOF; hypermorphic) of the encoded protein [1, 2]. Heterozygous lesions may underlie autosomal dominant traits by GOF, haploinsufficiency, or negative dominance. Biallelic lesions typically cause autosomal recessive traits by LOF of the encoded protein (rarely GOF), while X-linked recessive traits arise from LOF of genes on the X chromosome, either in the hemizygous state in males or in the homozygous state in females. Rare X-linked dominant traits can also arise from LOF or GOF variants. This results in aberrant immunity due to the critical roles of these proteins in the development, maintenance and function of cells of the immune system, or cells other than leukocytes that contribute to immunity, during homeostasis and in response to external (e.g., infectious agents or environmental antigens) and internal (e.g., cytokines, self-antigens and cancer cells) stimuli [3,4,5]. Inborn errors of immunity were traditionally considered to be rare diseases, affecting ~ 1 in 10,000 to 1 in 50,000 births. However, with ongoing discovery of novel inborn errors of immunity (Fig. 1a) and improved definition of clinical phenotypes [6,7,8], the collective prevalence of these conditions is more likely to be at least 1/1000–1/5000 [9]. Indeed, more common inborn errors have recently been described [10]. Regardless of their exact incidence and prevalence, inborn errors of immunity represent an unprecedented model to link defined monogenic defects with clinical phenotypes of immune dysregulation, in a broad sense of the term. As a committee, we are aware that human immunity involves cells other than circulating or tissue leukocytes and that it can be scaled up from the immune system to the whole organism. Inborn errors of immunity have unequivocally revealed non-redundant roles of single genes and their products in immune function [3, 4, 6,7,8], formed the basis of improved mechanism-based therapies for the immunopathology underlying many diseases [8, 11], established immunological paradigms representing the foundations of basic, clinical and translational immunology [3,4,5, 9, 12,13,14], and provided insights into the molecular pathogenesis of more common diseases [9, 15]. Clear examples of these include:
-
The initial description by Bruton of X-linked agammaglobulinemia (XLA) and the ability to treat this condition with antibody replacement therapy (the mainstay treatment for antibody deficiency diseases such as CVID) [16]
-
The discovery of mutations in BTK [12] and the subsequent development of BTK-inhibitors such as ibrutinib for the treatment of B cell malignancies [14]
-
Progressive CD4 T cell deficiency explains opportunistic infections secondary to HIV infection [9].
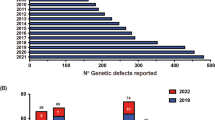
Rate of discovery of novel inborn errors of immunity: 1983–2019. a The number of genetic defects underlying monogenic immune disorders as reported by the IUIS/WHO committee in the indicated year. b The number of pathogenic gene variants listed in each table by the IUIS committee. Report published in 2017, and the number of new genes for each table contained in this report (red bars). The numbers in each column correspond to the number of genes reported in the 2017 IUIS update (blue bars) [1, 2], the number of new genes for each table contained in this report (red bars), and the total number of genes for each table. Note: only data for Tables 1, 2, 3, 4, 5, 6, 7, and 8 are shown, because Table 9 (bone marrow failure) is a new addition to the current report.
Thus, the study of inborn errors of immunity has provided profound advances in the practice of precision molecular medicine.
Since the early 1950s, when XLA was one of the first primary immune deficiencies to be described [16], clinical immunology has leveraged advances in the development of new methods to expedite the identification of defects of the immune system and the cellular, molecular, and genetic aberrations underlying these conditions. Indeed, the completion of the Human Genome Project in the early 2000s, coupled with rapid developments in next generation DNA sequencing (NGS) technologies, enabled the application of cost-effective and time-efficient sequencing of targeted gene panels, whole exomes, or whole genomes to cohorts of patients suspected of having a monogenic explanation for their disease. These platforms have led to a quantum leap in the identification and diagnosis of previously undefined genetically determined defects of the immune system (Fig. 1a, b; [6,7,8]).
The International Union of Immunological Societies Expert Committee of Inborn Errors of Immunity comprises pediatric and adult clinical immunologists, clinician/scientists and researchers in basic immunology from across the globe (https://iuis.org/committees/iei/). A major objective and responsibility of the committee is to provide the clinical and research communities with an update of genetic causes of immune deficiency and dysregulation. The committee has existed since 1970 and has published an updated report approximately every 2 years to inform the field of these advances (Fig. 1a). In March 2019, the committee met in New York to discuss and debate the inclusion of genetic variants published over the preceding 2 years (since June 2017) [1, 2], as well as gene mutations that had appeared in the literature earlier but, based on newly available evidence, were now substantiated (Fig. 1b).
Rather than simply including every gene variant reported, the committee applies very stringent criteria such that only those genes with convincing evidence of disease pathogenicity are classified as causes of novel inborn errors of immunity [17]. The Committee makes informed judgments for including new genetic causes of immunological conditions based on what we believe is most useful for practitioners caring for patients. Our current, and continuously evolving, practice is that criteria for inclusion can be met by several ways, for instance peer-reviewed publication of (1) multiple cases from unrelated kindreds, including detailed immunologic data, or (2) very few cases, or even a single case (see below), for whom compelling mechanistic/pathogenic data is also provided, generally from parallel studies in an animal or cell culture model.
Herein, we provide this latest update. The inborn errors of immunity are listed in 10 tables: Combined immunodeficiencies (Table 1), Combined immunodeficiencies with syndromic features (Table 2), Predominantly antibody deficiencies (Table 3), Diseases of immune dysregulation (Table 4), Congenital defects of phagocytes (Table 5), Defects in intrinsic and innate immunity (Table 6), Autoinflammatory diseases (Table 7), Complement deficiencies (Table 8), and Phenocopies of inborn errors of immunity (Table 10) (Fig. 1b). Since the last update (published January 2018) [1, 2], we have added a new table to consolidate genes that cause bone marrow failure (Table 9). Our division into phenotypes does not imply that the presentation is homogeneous. Rather, we recognize that substantial phenotypic and clinical heterogeneity exists within groups of patients with mutations in the same gene and even between individuals from the same pedigree with the identical gene mutation. To simplify the classification, each disorder has been listed only once, although distinct disorders due to mutations in the same gene, but with different modes of inheritance and pathogenic mechanisms are listed individually. Thus, several genes appear more than once in this update (some examples are listed below). Sub-divisions within each table segregate groups of disorders into coherent phenotypic sets. OMIM numbers are also provided within each table. If a OMIM number has not yet been issued for a particular genetic condition, then the number provided generally refers to the OMIM for that gene. Beneath each table, the new disorders added to this update are highlighted for easy reference.
The advances in our understanding of clinical immunology continue to expand at a vast and remarkable rate, with the addition in this update of many—64, distributed across all tables (Fig. 1b)—novel genetic defects underlying inborn errors of immunity. Perhaps not surprisingly, most if not all of these new variants were identified by NGS, thus highlighting that whole exome/whole genome sequencing has become the gold standard for identifying novel pathogenic gene variants [6,7,8]. Indeed, since the first application of NGS to identify novel inborn errors of immunity was published in 2010 [18], ~ 45% of all currently known disease-causing variants have been discovered by whole exome/genome sequencing. Thus, a typical approach to identifying a pathogenic variant in a new patient might now consist of first sequencing a phenotype-driven panel of genes and advancing to whole exome/genome sequencing if the cause of disease remains elusive.
In this update, we increase the list of immunological diseases to 404, with 430 known genetic defects identified as causing these conditions. The unbiased application of NGS to the discovery and characterization of novel inborn errors of immunity continues to inform clinical and basic immunology. Thus, additional phenotypes have been identified for conditions resulting from variants in known and novel genes; the penetrance of genetic variants on clinical phenotypes has been shown to be highly variable; and clinical entities sharing common phenotypes have been discovered. For example, this update includes the findings that bi-allelic mutations in ZNF341 [19, 20], IL6ST (encoding gp130, a common component of the receptors for IL-6, IL-11, IL-27, LIF, OSM, CNTF) [21, 22], or IL6R [23, 24] all cause conditions that resemble autosomal dominant hyper-IgE syndrome due to dominant negative mutations in STAT3 [15]. Detailed analyses of these patients revealed a novel mechanism of regulating STAT3 signaling (via the transcription factor ZNF341) and defined the exact consequences of impaired IL-6/IL-6R/gp130 and putatively IL-11/IL-11R/gp130 signaling to the phenotype of AD-HIES.
Furthermore, key findings over the past 2 years continue to reveal that distinct mechanisms of disease (GOF, LOF, dominant negative, haploinsufficient), as well as different modes of inheritance (autosomal recessive, autosomal dominant) of variants in the same gene can cause disparate clinical conditions. This is a fascinating aspect of the genetics of human disease, and a salient reminder to be cognizant of the nature of the genetic variants identified from NGS. It is these genes that have several entries in this update. A few recent examples include:
-
1.
Heterozygous variants in CARD11 [25, 26] or STAT5B [27] can be pathogenic due to negative dominance. This was potentially unexpected because autosomal recessive LOF variants in both of these genes were previously reported to cause combined immunodeficiency and severe immune dysregulation, respectively, yet heterozygous relatives of these affected individuals were healthy [28, 29].
-
2.
While heterozygous dominant negative mutations in TCF3, encoding the transcription factor E47, cause B cell deficiency and agammaglobulinemia [30], nonsense mutations in TCF3 have now been identified that are pathogenic only in an autosomal recessive state, as heterozygous carriers of these particular allelic variants remained healthy [31, 32].
-
3.
A heterozygous hypermorphic variant in IKBKB was found to cause a combined immunodeficiency [33] not too dissimilar to the original description of bi-allelic, recessive variants in IKBKB [34]. Similarly, bi-allelic LOF mutations in PIK3CD are now known to cause B cell deficiency and agammaglobulinemia [35–37], which is quite distinct from the immune dysregulated state of individuals with monoallelic activating PIK3CD mutations [1, 37]. This observation nicely parallels the earlier findings of either homozygous or heterozygous mutations in PIK3R1 that clinically phenocopy recessive or activating mutations in PIK3CD respectively [1, 37].
-
4.
Distinct diseases can result from heterozygous mutations in IKZF1 (Ikaros): combined immunodeficiency due to dominant negative alleles [38] or CVID due to haploinsufficiency [39].
-
5.
Similar to STAT1 [40], variants in RAC2 [41,42,43,44,45] or CARD11 [25, 26, 28] can be pathogenic either as monoallelic GOF or LOF or bi-allelic recessive LOF.
Thus, these findings have revealed the fundamental importance of elucidating the impact of a novel variant on the function of the encoded protein and thus the mechanism of pathogenicity. Furthermore, these new entries are an important reminder not to overlook the potential significance of identifying heterozygous variants in genes previously believed to cause disease only in a biallelic manner or to result in a previously defined specific clinical entity. Indeed, there are now at least 35 genes that have multiple entries in the current update, reflecting the distinct mechanisms by which variants result in or cause disease (e.g., STAT1, STAT3, NLRP1, RAC2, ZAP70, CARD11, IKBKB, WAS, JAK1, IFIH1, C3, C1R, C1S–GOF or LOF; STAT5, STAT1, CARD11, ACD, CFH, CFHR1–5, FOXN1, RAC2, TCF3, AICDA, PIK3R1, IFNGR1, TREX1, TICAM1, IRF8–AD or AR; PIK3CD–AD GOF, AR LOF; IKZF1–AD, or haploinsufficient; NLRP3—distinct disease phenotypes despite all resulting from GOF alleles).
As noted above, genetic, biochemical, and functional analyses of putative novel pathogenic variants need to meet stringent criteria to be considered for inclusion in this update [17]. These criteria can make reporting genetic findings from single cases challenging, as often the best evidence that a novel variant is disease-causing is to identify additional, similarly affected but unrelated individuals with the same variants, or functionally similar variants in the same gene. While this can be challenging, particularly in light of the rarity of individual inborn errors of immunity, robust mechanistic laboratory investigations continue to provide compelling data from single patients, with or without evidence from animal models. Specifically, homozygous LOF mutations in IRF9 [46] and IL18BP [47] were identified and rigorously characterized in single patients and found to be the molecular cause of life-threatening influenza and fulminant viral hepatitis, respectively.
The study and discovery of novel inborn errors of immunity can also enable improved patient management by implementing gene-specific targeted therapies. Thus, JAK inhibitors are being used to treat disorders of immune dysregulation resulting from GOF mutations in JAK1, STAT1 or STAT3 [11], while mTOR inhibitors such as rapamycin or PI3K p110δ-specific inhibitors have been reported for the treatment of individuals with PIK3CD GOF or PIK3R1 LOF mutations [37]. Regarding novel gene defects, immune dysregulation due to DEF6 deficiency was successfully treated with abatacept (CTLA4-Ig) [48]. This correlated with impaired CTLA4 expression and function in DEF6-deficient T cells [48] and parallels the therapeutic use of abatacept and belatacept for LRBA-deficiency and CTLA4 haploinsufficiency, both of which are characterized by reduced CTLA4 expression in affected regulatory T cells [49, 50]. From a theoretical perspective, the finding that MSMD can be caused by mutations in IL12RB2, IL23R or SPPL2A and that these mutations are associated with impaired production of IFNγ—a requisite of anti-mycobacterial immunity—implies that IFNγ administration could be therapeutically beneficial in these clinical settings [51, 52]. Similarly, recombinant IL18BP could potentially ameliorate viral-induced liver toxicity due to IL18BP deficiency [47].
The goals of the IUIS Expert Committee on Inborn Errors of Immunity are to increase awareness, facilitate recognition, promote optimal treatment, and support research in the field of disorders of immunity. Thus, this 2019 Update and the accompanying “Phenotypical IUIS Classification” publications are intended as resources for clinicians and researchers. Importantly, these tables underpin the design of panels used for targeted gene sequencing to facilitate genetic diagnoses or inborn errors. In the past 5 years, the number of gene defects underlying inborn errors of immunity has nearly doubled from ~ 250 to 430 (Fig. 1a). The human genome contains 1800–2000 genes that are known to be involved in immune responses [13]. Thus, the discovery and study of inborn errors of immunity has elegantly illustrated that > 20% of these immune genes play non-redundant roles in host defense and immune regulation. With the improved identification and phenotyping of patients with rare diseases, combined with high throughput genome sequencing, the number of genes fundamentally required for immunity will no doubt continue to increase, further revealing critical and novel roles for specific genes, molecules, pathways and cell types in immune responses, as well as mechanisms of disease pathogenesis and targets for immunotherapies. The field of inborn errors of immunity, and the global clinical and research communities, will therefore continue to provide key insights into basic and clinical immunology.
Change history
22 February 2020
The original version of this article unfortunately contained mistakes in reference numbers. The in-text citations and the references were mismatched. The original article has been corrected.
References
Picard C, Bobby Gaspar H, Al-Herz W, Bousfiha A, Casanova JL, Chatila T, et al. International Union of Immunological Societies: 2017 primary immunodeficiency diseases committee report on inborn errors of immunity. J Clin Immunol. 2018;38(1):96–128. https://doi.org/10.1007/s10875-017-0464-9.
Bousfiha A, Jeddane L, Picard C, Ailal F, Bobby Gaspar H, Al-Herz W, et al. The 2017 IUIS phenotypic classification for primary Immunodeficiencies. J Clin Immunol. 2018;38(1):129–43. https://doi.org/10.1007/s10875-017-0465-8.
Casanova JL, Abel L. Human genetics of infectious diseases: unique insights into immunological redundancy. Semin Immunol. 2018;36:1–12. https://doi.org/10.1016/j.smim.2017.12.008.
Fischer A, Rausell A. What do primary immunodeficiencies tell us about the essentiality/redundancy of immune responses? Semin Immunol. 2018;36:13–6. https://doi.org/10.1016/j.smim.2017.12.001.
Zhang SY, Jouanguy E, Zhang Q, Abel L, Puel A, Casanova JL. Human inborn errors of immunity to infection affecting cells other than leukocytes: from the immune system to the whole organism. Curr Opin Immunol. 2019;59:88–100. https://doi.org/10.1016/j.coi.2019.03.008.
Bucciol G, Moens L, Bosch B, Bossuyt X, Casanova JL, Puel A, et al. Lessons learned from the study of human inborn errors of innate immunity. J Allergy Clin Immunol. 2019;143(2):507–27. https://doi.org/10.1016/j.jaci.2018.07.013.
Meyts I, Bosch B, Bolze A, Boisson B, Itan Y, Belkadi A, et al. Exome and genome sequencing for inborn errors of immunity. J Allergy Clin Immunol. 2016;138(4):957–69. https://doi.org/10.1016/j.jaci.2016.08.003.
Picard C, Fischer A. Contribution of high-throughput DNA sequencing to the study of primary immunodeficiencies. Eur J Immunol. 2014;44(10):2854–61. https://doi.org/10.1002/eji.201444669.
Zhang Q, Frange P, Blanche S, Casanova JL. Pathogenesis of infections in HIV-infected individuals: insights from primary immunodeficiencies. Curr Opin Immunol. 2017;48:122–33. https://doi.org/10.1016/j.coi.2017.09.002.
Kerner G, Ramirez-Alejo N, Seeleuthner Y, Yang R, Ogishi M, Cobat A, et al. Homozygosity for TYK2 P1104A underlies tuberculosis in about 1% of patients in a cohort of European ancestry. Proc Natl Acad Sci U S A. 2019;116(21):10430–4. https://doi.org/10.1073/pnas.1903561116.
Leiding JW, Forbes LR. Mechanism-based precision therapy for the treatment of primary immunodeficiency and primary Immunodysregulatory diseases. J Allergy Clin Immunol Pract. 2019;7(3):761–73. https://doi.org/10.1016/j.jaip.2018.12.017.
Conley ME, Dobbs AK, Farmer DM, Kilic S, Paris K, Grigoriadou S, et al. Primary B cell immunodeficiencies: comparisons and contrasts. Annu Rev Immunol. 2009;27:199–227. https://doi.org/10.1146/annurev.immunol.021908.132649.
Fischer A, Rausell A. Primary immunodeficiencies suggest redundancy within the human immune system. Sci Immunol. 2016;1(6). https://doi.org/10.1126/sciimmunol.aah5861.
Gayko U, Fung M, Clow F, Sun S, Faust E, Price S, et al. Development of the Bruton's tyrosine kinase inhibitor ibrutinib for B cell malignancies. Ann N Y Acad Sci. 2015;1358:82–94. https://doi.org/10.1111/nyas.12878.
Ma CS, Tangye SG. Flow Cytometric-based analysis of defects in lymphocyte differentiation and function due to inborn errors of immunity. Front Immunol. 2019;10:2108. https://doi.org/10.3389/fimmu.2019.02108.
Bruton OC. Agammaglobulinemia Pediatrics. 1952;9(6):722–8.
Casanova JL, Conley ME, Seligman SJ, Abel L, Notarangelo LD. Guidelines for genetic studies in single patients: lessons from primary immunodeficiencies. J Exp Med. 2014;211(11):2137–49. https://doi.org/10.1084/jem.20140520.
Byun M, Abhyankar A, Lelarge V, Plancoulaine S, Palanduz A, Telhan L, et al. Whole-exome sequencing-based discovery of STIM1 deficiency in a child with fatal classic Kaposi sarcoma. J Exp Med. 2010;207(11):2307–12. https://doi.org/10.1084/jem.20101597.
Beziat V, Li J, Lin JX, Ma CS, Li P, Bousfiha A, et al. A recessive form of hyper-IgE syndrome by disruption of ZNF341-dependent STAT3 transcription and activity. Sci Immunol. 2018;3(24). https://doi.org/10.1126/sciimmunol.aat4956.
Frey-Jakobs S, Hartberger JM, Fliegauf M, Bossen C, Wehmeyer ML, Neubauer JC, et al. ZNF341 controls STAT3 expression and thereby immunocompetence. Sci Immunol. 2018;3(24). https://doi.org/10.1126/sciimmunol.aat4941.
Shahin T, Aschenbrenner D, Cagdas D, Bal SK, Conde CD, Garncarz W, et al. Selective loss of function variants in IL6ST cause hyper-IgE syndrome with distinct impairments of T-cell phenotype and function. Haematologica. 2019;104(3):609–21. https://doi.org/10.3324/haematol.2018.194233.
Schwerd T, Twigg SRF, Aschenbrenner D, Manrique S, Miller KA, Taylor IB, et al. A biallelic mutation in IL6ST encoding the GP130 co-receptor causes immunodeficiency and craniosynostosis. J Exp Med. 2017;214(9):2547–62. https://doi.org/10.1084/jem.20161810.
Spencer S, Kostel Bal S, Egner W, Lango Allen H, Raza SI, Ma CA, et al. Loss of the interleukin-6 receptor causes immunodeficiency, atopy, and abnormal inflammatory responses. J Exp Med. 2019;216(9):1986–98. https://doi.org/10.1084/jem.20190344.
Nahum A, Sharfe N, Broides A, Dadi H, Naghdi Z, Mandola AB, et al. Defining the biological responses of IL-6 by the study of a novel IL-6 receptor chain (IL6R) immunodeficiency. J Allergy Clin Immunol. 2019. https://doi.org/10.1016/j.jaci.2019.11.015.
Ma CA, Stinson JR, Zhang Y, Abbott JK, Weinreich MA, Hauk PJ, et al. Germline hypomorphic CARD11 mutations in severe atopic disease. Nat Genet. 2017;49(8):1192–201. https://doi.org/10.1038/ng.3898.
Dorjbal B, Stinson JR, Ma CA, Weinreich MA, Miraghazadeh B, Hartberger JM, et al. Hypomorphic caspase activation and recruitment domain 11 (CARD11) mutations associated with diverse immunologic phenotypes with or without atopic disease. J Allergy Clin Immunol. 2019;143(4):1482–95. https://doi.org/10.1016/j.jaci.2018.08.013.
Klammt J, Neumann D, Gevers EF, Andrew SF, Schwartz ID, Rockstroh D, et al. Dominant-negative STAT5B mutations cause growth hormone insensitivity with short stature and mild immune dysregulation. Nat Commun. 2018;9(1):2105. https://doi.org/10.1038/s41467-018-04521-0.
Lu HY, Bauman BM, Arjunaraja S, Dorjbal B, Milner JD, Snow AL, et al. The CBM-opathies-A rapidly expanding Spectrum of human inborn errors of immunity caused by mutations in the CARD11-BCL10-MALT1 complex. Front Immunol. 2018;9:2078. https://doi.org/10.3389/fimmu.2018.02078.
Nadeau K, Hwa V, Rosenfeld RG. STAT5b deficiency: an unsuspected cause of growth failure, immunodeficiency, and severe pulmonary disease. J Pediatr. 2011;158(5):701–8. https://doi.org/10.1016/j.jpeds.2010.12.042.
Boisson B, Wang YD, Bosompem A, Ma CS, Lim A, Kochetkov T, et al. A recurrent dominant negative E47 mutation causes agammaglobulinemia and BCR(−) B cells. J Clin Invest. 2013;123(11):4781–5. https://doi.org/10.1172/JCI71927.
Ben-Ali M, Yang J, Chan KW, Ben-Mustapha I, Mekki N, Benabdesselem C, et al. Homozygous transcription factor 3 gene (TCF3) mutation is associated with severe hypogammaglobulinemia and B-cell acute lymphoblastic leukemia. J Allergy Clin Immunol. 2017;140(4):1191–4 e4. https://doi.org/10.1016/j.jaci.2017.04.037.
Qureshi S, Sheikh MDA, Qamar FN. Autosomal recessive Agammaglobulinemia - first case with a novel TCF3 mutation from Pakistan. Clin Immunol. 2019;198:100–1. https://doi.org/10.1016/j.clim.2018.07.016.
Cardinez C, Miraghazadeh B, Tanita K, da Silva E, Hoshino A, Okada S, et al. Gain-of-function IKBKB mutation causes human combined immune deficiency. J Exp Med. 2018;215(11):2715–24. https://doi.org/10.1084/jem.20180639.
Pannicke U, Baumann B, Fuchs S, Henneke P, Rensing-Ehl A, Rizzi M, et al. Deficiency of innate and acquired immunity caused by an IKBKB mutation. N Engl J Med. 2013;369(26):2504–14. https://doi.org/10.1056/NEJMoa1309199.
Sogkas G, Fedchenko M, Dhingra A, Jablonka A, Schmidt RE, Atschekzei F. Primary immunodeficiency disorder caused by phosphoinositide 3-kinase delta deficiency. J Allergy Clin Immunol. 2018;142(5):1650–3 e2. https://doi.org/10.1016/j.jaci.2018.06.039.
Cohen SB, Bainter W, Johnson JL, Lin TY, Wong JCY, Wallace JG, et al. Human primary immunodeficiency caused by expression of a kinase-dead p110delta mutant. J Allergy Clin Immunol. 2019;143(2):797–9 e2. https://doi.org/10.1016/j.jaci.2018.10.005.
Tangye SG, Bier J, Lau A, Nguyen T, Uzel G, Deenick EK. Immune Dysregulation and disease pathogenesis due to activating mutations in PIK3CD-the Goldilocks' effect. J Clin Immunol. 2019;39(2):148–58. https://doi.org/10.1007/s10875-019-00612-9.
Boutboul D, Kuehn HS, Van de Wyngaert Z, Niemela JE, Callebaut I, Stoddard J, et al. Dominant-negative IKZF1 mutations cause a T, B, and myeloid cell combined immunodeficiency. J Clin Invest. 2018;128(7):3071–87. https://doi.org/10.1172/JCI98164.
Kuehn HS, Boisson B, Cunningham-Rundles C, Reichenbach J, Stray-Pedersen A, Gelfand EW, et al. Loss of B cells in patients with heterozygous mutations in IKAROS. N Engl J Med. 2016;374(11):1032–43. https://doi.org/10.1056/NEJMoa1512234.
Toubiana J, Okada S, Hiller J, Oleastro M, Lagos Gomez M, Aldave Becerra JC, et al. Heterozygous STAT1 gain-of-function mutations underlie an unexpectedly broad clinical phenotype. Blood. 2016;127(25):3154–64. https://doi.org/10.1182/blood-2015-11-679902.
Alkhairy OK, Rezaei N, Graham RR, Abolhassani H, Borte S, Hultenby K, et al. RAC2 loss-of-function mutation in 2 siblings with characteristics of common variable immunodeficiency. J Allergy Clin Immunol. 2015;135(5):1380-4 e1-5. https://doi.org/10.1016/j.jaci.2014.10.039.
Hsu AP, Donko A, Arrington ME, Swamydas M, Fink D, Das A, et al. Dominant activating RAC2 mutation with lymphopenia, immunodeficiency, and cytoskeletal defects. Blood. 2019;133(18):1977–88. https://doi.org/10.1182/blood-2018-11-886028.
Lougaris V, Chou J, Beano A, Wallace JG, Baronio M, Gazzurelli L, et al. A monoallelic activating mutation in RAC2 resulting in a combined immunodeficiency. J Allergy Clin Immunol. 2019;143(4):1649–53 e3. https://doi.org/10.1016/j.jaci.2019.01.001.
Sharapova SO, Haapaniemi E, Sakovich IS, Kostyuchenko LV, Donko A, Dulau-Florea A, et al. Heterozygous activating mutation in RAC2 causes infantile-onset combined immunodeficiency with susceptibility to viral infections. Clin Immunol. 2019;205:1–5. https://doi.org/10.1016/j.clim.2019.05.003.
Smits BM, Lelieveld PHC, Ververs FA, Turkenburg M, de Koning C, van Dijk M, et al. A dominant activating RAC2 variant associated with immunodeficiency and pulmonary disease. Clin Immunol. 2019;108248. https://doi.org/10.1016/j.clim.2019.108248.
Hernandez N, Melki I, Jing H, Habib T, Huang SSY, Danielson J, et al. Life-threatening influenza pneumonitis in a child with inherited IRF9 deficiency. J Exp Med. 2018;215(10):2567–85. https://doi.org/10.1084/jem.20180628.
Belkaya S, Michailidis E, Korol CB, Kabbani M, Cobat A, Bastard P, et al. Inherited IL-18BP deficiency in human fulminant viral hepatitis. J Exp Med. 2019;216(8):1777–90. https://doi.org/10.1084/jem.20190669.
Serwas NK, Hoeger B, Ardy RC, Stulz SV, Sui Z, Memaran N, et al. Human DEF6 deficiency underlies an immunodeficiency syndrome with systemic autoimmunity and aberrant CTLA-4 homeostasis. Nat Commun. 2019;10(1):3106. https://doi.org/10.1038/s41467-019-10812-x.
Lo B, Zhang K, Lu W, Zheng L, Zhang Q, Kanellopoulou C, et al. AUTOIMMUNE DISEASE. Patients with LRBA deficiency show CTLA4 loss and immune dysregulation responsive to abatacept therapy. Science. 2015;349(6246):436–40. https://doi.org/10.1126/science.aaa1663.
Schwab C, Gabrysch A, Olbrich P, Patino V, Warnatz K, Wolff D, et al. Phenotype, penetrance, and treatment of 133 cytotoxic T-lymphocyte antigen 4-insufficient subjects. J Allergy Clin Immunol. 2018;142(6):1932–46. https://doi.org/10.1016/j.jaci.2018.02.055.
Martinez-Barricarte R, Markle JG, Ma CS, Deenick EK, Ramirez-Alejo N, Mele F, et al. Human IFN-gamma immunity to mycobacteria is governed by both IL-12 and IL-23. Sci Immunol. 2018;3(30). https://doi.org/10.1126/sciimmunol.aau6759.
Kong XF, Martinez-Barricarte R, Kennedy J, Mele F, Lazarov T, Deenick EK, et al. Disruption of an antimycobacterial circuit between dendritic and helper T cells in human SPPL2a deficiency. Nat Immunol. 2018;19(9):973–85. https://doi.org/10.1038/s41590-018-0178-z.
Roussel L, Landekic M, Golizeh M, Gavino C, Zhong MC, Chen J, et al. Loss of human ICOSL results in combined immunodeficiency. J Exp Med. 2018;215(12):3151–64. https://doi.org/10.1084/jem.20180668.
Conde CD, Petronczki OY, Baris S, Willmann KL, Girardi E, Salzer E, et al. Polymerase delta deficiency causes syndromic immunodeficiency with replicative stress. J Clin Invest. 2019;129(10):4194–206. https://doi.org/10.1172/JCI128903.
Cui Y, Keles S, Charbonnier LM, Jule AM, Henderson L, Celik SC, et al. Combined immunodeficiency due to a loss of function mutation in DNA Polymerase Delta 1. J Allergy Clin Immunol. 2019. https://doi.org/10.1016/j.jaci.2019.10.004.
Badran YR, Dedeoglu F, Leyva Castillo JM, Bainter W, Ohsumi TK, Bousvaros A, et al. Human RELA haploinsufficiency results in autosomal-dominant chronic mucocutaneous ulceration. J Exp Med. 2017;214(7):1937–47. https://doi.org/10.1084/jem.20160724.
Comrie WA, Faruqi AJ, Price S, Zhang Y, Rao VK, Su HC, et al. RELA haploinsufficiency in CD4 lymphoproliferative disease with autoimmune cytopenias. J Allergy Clin Immunol. 2018;141(4):1507–10 e8. https://doi.org/10.1016/j.jaci.2017.11.036.
Beaussant-Cohen S, Jaber F, Massaad MJ, Weeks S, Jones J, Alosaimi MF, et al. Combined immunodeficiency in a patient with c-Rel deficiency. J Allergy Clin Immunol. 2019;144(2):606–8 e4. https://doi.org/10.1016/j.jaci.2019.05.003.
Calzoni E, Platt CD, Keles S, Kuehn HS, Beaussant-Cohen S, Zhang Y, et al. F-BAR domain only protein 1 (FCHO1) deficiency is a novel cause of combined immune deficiency in human subjects. J Allergy Clin Immunol. 2019;143(6):2317–21 e12. https://doi.org/10.1016/j.jaci.2019.02.014.
Maffucci P, Chavez J, Jurkiw TJ, O'Brien PJ, Abbott JK, Reynolds PR, et al. Biallelic mutations in DNA ligase 1 underlie a spectrum of immune deficiencies. J Clin Invest. 2018;128(12):5489–504. https://doi.org/10.1172/JCI99629.
Bosticardo M, Yamazaki Y, Cowan J, Giardino G, Corsino C, Scalia G, et al. Heterozygous FOXN1 variants cause low TRECs and severe T cell Lymphopenia, revealing a crucial role of FOXN1 in supporting early Thymopoiesis. Am J Hum Genet. 2019;105(3):549–61. https://doi.org/10.1016/j.ajhg.2019.07.014.
Lyons JJ, Liu Y, Ma CA, Yu X, O'Connell MP, Lawrence MG, et al. ERBIN deficiency links STAT3 and TGF-beta pathway defects with atopy in humans. J Exp Med. 2017;214(3):669–80. https://doi.org/10.1084/jem.20161435.
Schepers D, Tortora G, Morisaki H, MacCarrick G, Lindsay M, Liang D, et al. A mutation update on the LDS-associated genes TGFB2/3 and SMAD2/3. Hum Mutat. 2018;39(5):621–34. https://doi.org/10.1002/humu.23407.
Fabre A, Charroux B, Martinez-Vinson C, Roquelaure B, Odul E, Sayar E, et al. SKIV2L mutations cause syndromic diarrhea, or trichohepatoenteric syndrome. Am J Hum Genet. 2012;90(4):689–92. https://doi.org/10.1016/j.ajhg.2012.02.009.
Huppke P, Weissbach S, Church JA, Schnur R, Krusen M, Dreha-Kulaczewski S, et al. Activating de novo mutations in NFE2L2 encoding NRF2 cause a multisystem disorder. Nat Commun. 2017;8(1):818. https://doi.org/10.1038/s41467-017-00932-7.
Rodriguez R, Fournier B, Cordeiro DJ, Winter S, Izawa K, Martin E, et al. Concomitant PIK3CD and TNFRSF9 deficiencies cause chronic active Epstein-Barr virus infection of T cells. J Exp Med. 2019. https://doi.org/10.1084/jem.20190678.
Anzilotti C, Swan DJ, Boisson B, Deobagkar-Lele M, Oliveira C, Chabosseau P, et al. An essential role for the Zn(2+) transporter ZIP7 in B cell development. Nat Immunol. 2019;20(3):350–61. https://doi.org/10.1038/s41590-018-0295-8.
Broderick L, Yost S, Li D, McGeough MD, Booshehri LM, Guaderrama M, et al. Mutations in topoisomerase IIbeta result in a B cell immunodeficiency. Nat Commun. 2019;10(1):3644. https://doi.org/10.1038/s41467-019-11570-6.
Bouafia A, Lofek S, Bruneau J, Chentout L, Lamrini H, Trinquand A, et al. Loss of ARHGEF1 causes a human primary antibody deficiency. J Clin Invest. 2019;129(3):1047–60. https://doi.org/10.1172/JCI120572.
Keller B, Shoukier M, Schulz K, Bhatt A, Heine I, Strohmeier V, et al. Germline deletion of CIN85 in humans with X chromosome-linked antibody deficiency. J Exp Med. 2018;215(5):1327–36. https://doi.org/10.1084/jem.20170534.
Schubert D, Klein MC, Hassdenteufel S, Caballero-Oteyza A, Yang L, Proietti M, et al. Plasma cell deficiency in human subjects with heterozygous mutations in Sec61 translocon alpha 1 subunit (SEC61A1). J Allergy Clin Immunol. 2018;141(4):1427–38. https://doi.org/10.1016/j.jaci.2017.06.042.
Mauhin W, Habarou F, Gobin S, Servais A, Brassier A, Grisel C, et al. Update on Lysinuric protein intolerance, a multi-faceted disease retrospective cohort analysis from birth to adulthood. Orphanet J Rare Dis. 2017;12(1):3. https://doi.org/10.1186/s13023-016-0550-8.
Fernandez IZ, Baxter RM, Garcia-Perez JE, Vendrame E, Ranganath T, Kong DS, et al. A novel human IL2RB mutation results in T and NK cell-driven immune dysregulation. J Exp Med. 2019;216(6):1255–67. https://doi.org/10.1084/jem.20182015.
Zhang Z, Gothe F, Pennamen P, James JR, McDonald D, Mata CP, et al. Human interleukin-2 receptor beta mutations associated with defects in immunity and peripheral tolerance. J Exp Med. 2019;216(6):1311–27. https://doi.org/10.1084/jem.20182304.
Has C, Castiglia D, del Rio M, Diez MG, Piccinni E, Kiritsi D, et al. Kindler syndrome: extension of FERMT1 mutational spectrum and natural history. Hum Mutat. 2011;32(11):1204–12. https://doi.org/10.1002/humu.21576.
Kotlarz D, Marquardt B, Baroy T, Lee WS, Konnikova L, Hollizeck S, et al. Human TGF-beta1 deficiency causes severe inflammatory bowel disease and encephalopathy. Nat Genet. 2018;50(3):344–8. https://doi.org/10.1038/s41588-018-0063-6.
Cuchet-Lourenco D, Eletto D, Wu C, Plagnol V, Papapietro O, Curtis J, et al. Biallelic RIPK1 mutations in humans cause severe immunodeficiency, arthritis, and intestinal inflammation. Science. 2018;361(6404):810–3. https://doi.org/10.1126/science.aar2641.
Li Y, Fuhrer M, Bahrami E, Socha P, Klaudel-Dreszler M, Bouzidi A, et al. Human RIPK1 deficiency causes combined immunodeficiency and inflammatory bowel diseases. Proc Natl Acad Sci U S A. 2019;116(3):970–5. https://doi.org/10.1073/pnas.1813582116.
Alosaimi MF, Hoenig M, Jaber F, Platt CD, Jones J, Wallace J, et al. Immunodeficiency and EBV-induced lymphoproliferation caused by 4-1BB deficiency. J Allergy Clin Immunol. 2019;144(2):574–83 e5. https://doi.org/10.1016/j.jaci.2019.03.002.
Somekh I, Thian M, Medgyesi D, Gulez N, Magg T, Gallon Duque A, et al. CD137 deficiency causes immune dysregulation with predisposition to lymphomagenesis. Blood. 2019. https://doi.org/10.1182/blood.2019000644.
Carapito R, Konantz M, Paillard C, Miao Z, Pichot A, Leduc MS, et al. Mutations in signal recognition particle SRP54 cause syndromic neutropenia with Shwachman-diamond-like features. J Clin Invest. 2017;127(11):4090–103. https://doi.org/10.1172/JCI92876.
Bellanne-Chantelot C, Schmaltz-Panneau B, Marty C, Fenneteau O, Callebaut I, Clauin S, et al. Mutations in the SRP54 gene cause severe congenital neutropenia as well as Shwachman-diamond-like syndrome. Blood. 2018;132(12):1318–31. https://doi.org/10.1182/blood-2017-12-820308.
Dhanraj S, Matveev A, Li H, Lauhasurayotin S, Jardine L, Cada M, et al. Biallelic mutations in DNAJC21 cause Shwachman-diamond syndrome. Blood. 2017;129(11):1557–62. https://doi.org/10.1182/blood-2016-08-735431.
Arnadottir GA, Norddahl GL, Gudmundsdottir S, Agustsdottir AB, Sigurdsson S, Jensson BO, et al. A homozygous loss-of-function mutation leading to CYBC1 deficiency causes chronic granulomatous disease. Nat Commun. 2018;9(1):4447. https://doi.org/10.1038/s41467-018-06964-x.
Thomas DC, Charbonnier LM, Schejtman A, Aldhekri H, Coomber EL, Dufficy ER, et al. EROS/CYBC1 mutations: decreased NADPH oxidase function and chronic granulomatous disease. J Allergy Clin Immunol. 2019;143(2):782–5 e1. https://doi.org/10.1016/j.jaci.2018.09.019.
de Jong SJ, Crequer A, Matos I, Hum D, Gunasekharan V, Lorenzo L, et al. The human CIB1-EVER1-EVER2 complex governs keratinocyte-intrinsic immunity to beta-papillomaviruses. J Exp Med. 2018;215(9):2289–310. https://doi.org/10.1084/jem.20170308.
Hernandez N, Bucciol G, Moens L, Le Pen J, Shahrooei M, Goudouris E, et al. Inherited IFNAR1 deficiency in otherwise healthy patients with adverse reaction to measles and yellow fever live vaccines. J Exp Med. 2019;216(9):2057–70. https://doi.org/10.1084/jem.20182295.
Ogunjimi B, Zhang SY, Sorensen KB, Skipper KA, Carter-Timofte M, Kerner G, et al. Inborn errors in RNA polymerase III underlie severe varicella zoster virus infections. J Clin Invest. 2017;127(9):3543–56. https://doi.org/10.1172/JCI92280.
Carter-Timofte ME, Hansen AF, Mardahl M, Fribourg S, Rapaport F, Zhang SY, et al. Varicella-zoster virus CNS vasculitis and RNA polymerase III gene mutation in identical twins. Neurol Neuroimmunol Neuroinflamm. 2018;5(6):e500. https://doi.org/10.1212/NXI.0000000000000500.
Zhang SY, Clark NE, Freije CA, Pauwels E, Taggart AJ, Okada S, et al. Inborn errors of RNA lariat metabolism in humans with brainstem viral infection. Cell. 2018;172(5):952–65 e18. https://doi.org/10.1016/j.cell.2018.02.019.
Guerin A, Kerner G, Marr N, Markle JG, Fenollar F, Wong N, et al. IRF4 haploinsufficiency in a family with Whipple's disease. Elife. 2018;7. https://doi.org/10.7554/eLife.32340.
Brehm A, Liu Y, Sheikh A, Marrero B, Omoyinmi E, Zhou Q, et al. Additive loss-of-function proteasome subunit mutations in CANDLE/PRAAS patients promote type I IFN production. J Clin Invest. 2015;125(11):4196–211. https://doi.org/10.1172/JCI81260.
Rodero MP, Tesser A, Bartok E, Rice GI, Della Mina E, Depp M, et al. Type I interferon-mediated autoinflammation due to DNase II deficiency. Nat Commun. 2017;8(1):2176. https://doi.org/10.1038/s41467-017-01932-3.
Al-Mayouf SM, Sunker A, Abdwani R, Abrawi SA, Almurshedi F, Alhashmi N, et al. Loss-of-function variant in DNASE1L3 causes a familial form of systemic lupus erythematosus. Nat Genet. 2011;43(12):1186–8. https://doi.org/10.1038/ng.975.
Ozcakar ZB, Foster J 2nd, Diaz-Horta O, Kasapcopur O, Fan YS, Yalcinkaya F, et al. DNASE1L3 mutations in hypocomplementemic urticarial vasculitis syndrome. Arthritis Rheum. 2013;65(8):2183–9. https://doi.org/10.1002/art.38010.
Carbonella A, Mancano G, Gremese E, Alkuraya FS, Patel N, Gurrieri F, et al. An autosomal recessive DNASE1L3-related autoimmune disease with unusual clinical presentation mimicking systemic lupus erythematosus. Lupus. 2017;26(7):768–72. https://doi.org/10.1177/0961203316676382.
Cho K, Yamada M, Agematsu K, Kanegane H, Miyake N, Ueki M, et al. Heterozygous mutations in OAS1 cause infantile-onset pulmonary alveolar Proteinosis with Hypogammaglobulinemia. Am J Hum Genet. 2018;102(3):480–6. https://doi.org/10.1016/j.ajhg.2018.01.019.
Zhong FL, Mamai O, Sborgi L, Boussofara L, Hopkins R, Robinson K, et al. Germline NLRP1 mutations cause skin inflammatory and Cancer susceptibility syndromes via Inflammasome activation. Cell. 2016;167(1):187–202 e17. https://doi.org/10.1016/j.cell.2016.09.001.
Drutman SB, Haerynck F, Zhong FL, Hum D, Hernandez NJ, Belkaya S, et al. Homozygous NLRP1 gain-of-function mutation in siblings with a syndromic form of recurrent respiratory papillomatosis. Proc Natl Acad Sci U S A. 2019;116(38):19055–63. https://doi.org/10.1073/pnas.1906184116.
Parlato M, Charbit-Henrion F, Pan J, Romano C, Duclaux-Loras R, Le Du MH, et al. Human ALPI deficiency causes inflammatory bowel disease and highlights a key mechanism of gut homeostasis. EMBO Mol Med. 2018;10(4). https://doi.org/10.15252/emmm.201708483.
Li Q, Lee CH, Peters LA, Mastropaolo LA, Thoeni C, Elkadri A, et al. Variants in TRIM22 that affect NOD2 signaling are associated with very-early-onset inflammatory bowel disease. Gastroenterology. 2016;150(5):1196–207. https://doi.org/10.1053/j.gastro.2016.01.031.
de Jesus AA, Brehm A, VanTries R, Pillet P, Parentelli AS, Montealegre Sanchez GA, et al. Novel proteasome assembly chaperone mutations in PSMG2/PAC2 cause the autoinflammatory interferonopathy CANDLE/PRAAS4. J Allergy Clin Immunol. 2019;143(5):1939–43 e8. https://doi.org/10.1016/j.jaci.2018.12.1012.
Gayden T, Sepulveda FE, Khuong-Quang DA, Pratt J, Valera ET, Garrigue A, et al. Germline HAVCR2 mutations altering TIM-3 characterize subcutaneous panniculitis-like T cell lymphomas with hemophagocytic lymphohistiocytic syndrome. Nat Genet. 2018;50(12):1650–7. https://doi.org/10.1038/s41588-018-0251-4.
Polprasert C, Takeuchi Y, Kakiuchi N, Yoshida K, Assanasen T, Sitthi W, et al. Frequent germline mutations of HAVCR2 in sporadic subcutaneous panniculitis-like T-cell lymphoma. Blood Adv. 2019;3(4):588–95. https://doi.org/10.1182/bloodadvances.2018028340.
Kapferer-Seebacher I, Pepin M, Werner R, Aitman TJ, Nordgren A, Stoiber H, et al. Periodontal Ehlers-Danlos syndrome is caused by mutations in C1R and C1S, which encode subcomponents C1r and C1s of complement. Am J Hum Genet. 2016;99(5):1005–14. https://doi.org/10.1016/j.ajhg.2016.08.019.
Acknowledgments
The members of the Inborn Errors of Immunity committee would like to thanks the International Union of Immunological Societies (IUIS) for funding, as well as CSL Behring, Baxalta and Shire/Takeda for providing educational grants to enable us to compile this classification update.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: The in-text citations and the references were mismatched.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tangye, S.G., Al-Herz, W., Bousfiha, A. et al. Human Inborn Errors of Immunity: 2019 Update on the Classification from the International Union of Immunological Societies Expert Committee. J Clin Immunol 40, 24–64 (2020). https://doi.org/10.1007/s10875-019-00737-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-019-00737-x