Abstract
Background
MicroRNAs (miRNAs) are short single-stranded RNA molecules that regulate gene expression post-transcriptionally. Several hundred miRNAs exist in the mammalian genome and regulate developmental processes, cell cycle, and survival.
Methods
In this review, we highlight general modes of miRNA function and relate them to how such regulation can be beneficial for immune homeostasis and the prevention of autoimmune diseases. We highlight examples of experimentally verified miRNA function and their target genes in the immune system and place them in context of concepts relevant to an understanding of autoimmune pathogenesis. Where available, we refer to clinical correlations. Finally, we speculate how emerging knowledge about miRNA function in the immune system might be used diagnostically and therapeutically.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The regulation of protein expression by (small) RNAs was first observed ∼25 years ago. In the mid-1980s, several studies reported that transgenic antisense RNA could inhibit gene expression in Drosophila, mammalian cells, and plants [1–4]. This phenomenon was termed post-transcriptional gene silencing in the plant biology world. In the 1990s, it was demonstrated that the worm Caenorhabditis elegans uses an endogenous small RNA to inhibit the synthesis of a protein through sequence complementarity to the 3′ untranslated region (3′ UTR) of the targeted mRNA [5]. Fire and Mello discovered that exogenous, ∼21–23 nucleotide double-stranded RNAs (siRNAs), interfered with protein-encoding genes and coined the term RNA interference (RNAi) [6]. A class of endogenous, small single-stranded non-protein-coding RNA genes, comprising hundreds of members, was termed microRNAs (miRNA) [7–9]. Today, this class of genes is thought to function as important regulators of development, cell differentiation, proliferation, and cell death in most multi-cellular organisms [10–12]. Other classes of small RNAs that are generated via different molecular machinery have been described, although their role in the immune regulation is less clear [13–17].
Mammalian miRNAs are transcribed as long precursors, which are processed by the microprocessor complex formed by Drosha/DGCR8 and other proteins. The precursors can encode a single miRNA or a cluster of several polycistronic miRNAs. The microprocessor complex recognizes the precursors and processes them into ∼70 nt precursors. The stem-loop precursor molecules are then exported through exportin 5 into the cytoplasm where they are processed by an enzyme called Dicer. The resulting short double-stranded RNA molecules are separated into single-stranded molecules of which one (guide strand) is loaded into the RNA-induced silencing complex formed by members of the Argonaute (Ago) family, Dicer, TRBP, and others. miRNAs pair with complementary sequences of their target mRNA through a region called “seed” at the 5′ end of the miRNA. Other regions and determinants have been described recently that guide miRNA function as well [18–20].
The bulk of current studies suggest that the large majority of miRNAs inhibit protein-coding genes. However, there are a few reported exceptions where miRNAs positively affect gene expression [21–23]. How inhibition of protein expression is achieved is still a matter of debate. mRNA degradation and translational inhibition have been put forth [24–28] with the most recent data, suggesting that the effect on mRNA degradation is more pronounced than translational inhibition [26].
Despite intensive research, miRNA biogenesis and the maintenance of a functional miRNA pool is still incompletely understood [29]. Particular proteins can regulate the generation of subsets of miRNAs [30] and active degradation of miRNAs has been described in plants and worms [31, 32]. Interestingly, several cytokines with known important functions in the immune system can control miRNA maturation for a subset of miRNAs at the post-transcriptional level. In this regard, transforming growth factor beta (TGF-ß) and bone morphogenetic proteins (BMPs) have been shown to regulate miRNA processing in human vascular smooth muscle cells [33]. Since it is known that TGF-ß can induce FoxP3 in conventional T cells, it will be interesting to determine if TGF-ß controls the high miR-21 expression observed in regulatory T cells (Treg) which are highly TGF-ß dependent and depend on Dicer for FoxP3 induction (ref [34] and Jeker & Bluestone, unpublished).
It is now clearly established that miRNAs play important functions in the immune system [35]. Convincing data has come from studies where individual miRNAs were over-expressed or disrupted and miRNA binding sites in target mRNAs have been genetically deleted. These studies demonstrated that the miR-17∼92 cluster, miR-150, miR-155, and miR-181a are important regulators of B and T cell development and play crucial roles for the establishment of a functional adaptive immune system [36–46].
Excellent comprehensive reviews on individual miRNAs, cell systems, or diseases have recently been published [35, 47–50]. Thus, in this review, we will cover general principles how miRNAs function as rheostats that control overall immune homeostasis. We will discuss modes of miRNA action and highlight examples of how the immune system relies on miRNAs to control lymphocyte development, differentiation, and control of immune responses. Although we will focus on miRNAs expressed in lymphocytes, we speculate that the general principles described will be applicable to the important role of these small RNAs throughout the immune system [35, 51–53]. Finally, we attempt to highlight how the emerging knowledge of miRNA function in the immune system might be exploited diagnostically and therapeutically.
Functional Evidence for the Importance of RNAi Machinery in Lymphocytes and Autoimmunity
The discovery of several key proteins in the biogenesis of small RNAs (Drosha/DGCR8, Dicer, and Argonautes in mammals) have allowed the simultaneous non-selective or partially selective ablation of hundreds of miRNAs while leaving protein-coding genes intact [29, 54–56]. Although constitutive deletion of Dicer is embryonic lethal in mice [10] and zebrafish [55], selective deletion of Dicer [57, 58], Drosha [59], DGCR8 [60, 61], or Ago2 [56] in individual immune subsets has been used to determine that miRNAs are critical for B, NK, NKT, and T cell development, function, and lineage stability of terminally differentiated lymphocytes [59, 62–68]. Dicer ablation in T cells demonstrated its essential role in αβ-T cell receptor (TCR) thymocyte development as Lck-cre-mediated, T cell-specific, Dicer deficiency led to severe thymic hypoplasia. Although there were no apparent changes in the CD4/CD8 lineage choice [65], the severity of this thymic dysplasia prevented a more detailed analysis of the peripheral T cell pool. Dicer ablation using a CD4-cre transgene, which is expressed at a later developmental time point, did not affect thymocyte maturation but resulted in mild T cell lymphopenia as Dicer-deficient T cells underwent apoptosis and proliferated less than wild type T cells [34, 66]. However, after primary stimulation, IL-2 production by Dicer-deficient T cells was normal but demonstrated reduced mRNA stability resulting in rapid decay after prolonged culture. In contrast to Dicer deletion in B cells, the Dicer-deficient T cells could not be rescued with a bcl-2 transgene, suggesting that Dicer was controlling more than the apoptotic program [34, 62].
Most importantly, T cell-specific Dicer-deficiency led to multiorgan autoimmunity of lung, liver, and colon and lymphoproliferative disease with lymphadenopathy and splenomegaly [34] concordant with a significant reduction of thymically derived, natural regulatory T cells (nTreg). Thus, three groups selectively ablated RNAi key enzymes in Treg using FoxP3-cre transgenes [59, 67, 68]. These studies, using different cre transgenic lines, showed largely overlapping results and demonstrated phenotypes resembling the scurfy phenotype where Treg completely lacks FoxP3 function [69]. Disease onset was very early with most mice dying within a few weeks of life. Deletion of Dicer using the CD4-cre strategy resulted in organ infiltration only after 3–4 months with incomplete penetrance [34, 59]. This was most likely due to combined autoimmunity and immunodeficiency resulting from Dicer ablation in all CD4 and CD8 T effector cells, as illustrated by the reduced capability to induce IL-17 in vitro [34].
In the experimental system used by Liston et al., female mice harbor Dicer-sufficient and Dicer-deficient Treg populations due to X-inactivation [68]. These mice remained healthy, indicating that tolerance could be preserved if a fraction of Treg cells remained wild type. The proportion of Dicer-sufficient and Dicer-deficient Treg cells was equal in thymus but favored wild type Tregs in the periphery. There was no proliferation defect. Treg cells remained anergic, but in vitro suppressive capacity was somewhat impaired, and in vitro TCR ligation led to decreased proliferation and increased apoptosis in the Dicer-deficient Treg. Importantly, Treg cells isolated from sick mice lost their hallmark anergy and were completely devoid of in vitro suppressive capacity [68]. These results indicate that miRNAs might be more important under stress conditions like inflammation than under homeostatic conditions.
Our studies provided insights into Treg lineage instability in the absence of Dicer [67]. Using genetic lineage tracing, we demonstrated that miRNA-deficient Treg can survive, but lose FoxP3 to a significant degree with at times >50% of Treg cells having lost FoxP3 expression. Furthermore, we demonstrated that about 5% of actively FoxP3 transcribing Dicer-deficient Treg cells had lost their anergic phenotype and begun to secrete the pathogenic cytokine IFN-γ. Thus, miRNA-deficient Treg cells lose lineage identity and take on Th1-like features. Finally, the indistinguishable phenotype between Treg-specific Dicer and Drosha ablation underscores the conclusion that indeed canonical miRNAs are critical to Treg homeostasis because Drosha is specific for miRNA biogenesis while Dicer also processes other classes of small RNAs [29, 59].
Mechanistically, Dicer-deficient T cells showed deficient induction of FoxP3 by TGF-β in vitro and IL-17 by TGF-β and IL-6 [34]. Of note, it appeared that the FoxP3 fluorescence was lower in the absence of Dicer [34] and the Dicer-deficiency altered T cell differentiation. Dicer-deficient cells were prone to develop a Th1 phenotype in various settings including normally Th2-driving conditions [66]. Thus, Dicer regulates peripheral terminal T cell differentiation and lineage stabilization and miRNA deficiency leads to severely skewed cytokine profiles. Most importantly, unlike most Dicer-deficient cell types, which showed decreased cell proliferation and viability, only T cell-specific Dicer deletion led to autoimmunity.
In summary, miRNAs are functionally important in virtually all lymphocytes examined; however, ablation of Dicer selectively in T cells leads to spontaneous autoimmune disease. This clinical outcome can be attributed to the development of non-functional Treg and a scurfy-like autoimmunity. The underlying mechanisms include miRNA regulation of Treg proliferation, apoptosis, and lineage identity, as well as an alteration of the Treg-defining transcriptome, FoxP3 expression, and ultimately suppressive function. As it has become clear that T cell plasticity is far more common than previously acknowledged [70–74], the importance of miRNAs in regulating and maintaining T cell terminal differentiation programs and a coordinated cytokine response, especially under stress conditions, is likely to play an essential role for proper immune homeostasis.
miRNA's Function as a Rheostat of Gene Expression
Whole proteome analyses have taught us that regulation of protein abundance of individual target genes by miRNAs is modest at best (often <50% of levels without miRNA regulation) [25, 27]. Despite strong evolutionary conservation, a systematic ablation of 83% of known miRNAs in C. elegans revealed that the majority of miRNAs was neither required for viability nor development [75]. This can be explained partially by functional redundancy because many miRNAs exist in families with identical seed sequences, and deletion of whole miRNA families can sometimes be functionally rescued by re-expression of a single miRNA [76, 77]. However, systematic ablation of whole families of sequence-related miRNAs were frequently dispensable when studying development or viability [77]. Nevertheless, individual miRNAs are important regulators of gene expression as absence or sometimes even only a reduction of individual miRNAs or miRNA clusters can have significant effects for the affected individual as demonstrated for miR-1 in the heart, miR-155 in the immune system, and the miR-17∼92 cluster in lung, heart, and B cell development [41, 43, 78, 79]. Thus, miRNAs are generally fine-tuners or rheostats that sharpen gene expression, which in some cases can control essential cellular functions [19, 25, 27]. So how can miRNAs have minor effects on individual proteins yet be important regulators whose absence in small subsets of immune cells (e.g., Treg) results in a fatal disease? Computational algorithms for conserved mammalian miRNAs predict an average of 300 targets per miRNA family [19]. This has experimentally been validated as over-expression of a single tissue-specific miRNA in HeLa cells down-regulated ∼100 transcripts within 12 h. Over-expression of miR-124, a brain-enriched miRNA, induced a shift towards a brain gene expression signature, whereas over-expression of miR-1, a miRNA enriched in muscle cells, induced a muscle-like mRNA transcriptome [24]. Thus, miRNAs can rapidly and simultaneously regulate entire protein networks or signaling pathways, thus shaping cell fate and identity. Furthermore, almost 50% of target genes have two or more miRNA binding sites for different miRNAs allowing for synergistic effects [80] and vice-versa, several miRNAs can be induced by the same stimulus synergizing and/or antagonizing similar pathways [35].
The precise regulation of gene expression by miRNAs has had an important impact on evolution [19, 80–82]. During evolution, miRNA target genes either enriched or avoided miRNA binding sites by changing their 3′ UTR length and/or miRNA binding site density [80]. Ubiquitously expressed genes tend to avoid miRNA regulation while miRNA-regulated genes show tissue-specific expression. In contrast, systematic analysis of miRNA binding sites in 3′ UTRs in Drosophila revealed that the only tissue-specific set of genes that avoids binding sites for a muscle-specific miRNA (miR-1) are muscle-specific genes [80]. In other words, by equipping non-muscle genes with binding sites for the muscle-specific miR-1 and co-expressing miR-1, Drosophila achieves muscle-specific gene expression by repressing non-muscle genes. This mutual exclusion might sharpen gene expression and confer robustness [80].
miRNAs as Guardians of Immune Homeostasis
As up to 50% of protein-coding genes are regulated to some degree by miRNAs, it is not surprising that autoimmunity can be affected by dysregulated miRNA function [83–85]. However, miRNAs are important for regulating immunity not only by the shear number of genes they regulate but because they have balancing effects that are central to immune homeostasis. A recent review suggested that only three miRNAs together are predicted to target >50% of lupus susceptibility genes [50]. The immune system has the ability to react quickly and vigorously but must be prevented from overshooting. After an infection has been successfully controlled, a return to homeostatic conditions must be achieved quickly; an accumulation of previously activated cells need to be avoided. Balance is difficult to achieve since small changes in the homeostatic equilibrium can lead to significant consequences because of the possibility of enormous amplification of rare precursors which underlies the exponential nature of immune responses [86]. Hence, balance is the key in the immune system. As a result, discrimination of noise (i.e., non-specific stimulation) from important information is critical. Therefore, stochastic transcription needs to be repressed, and parallel signaling through opposing cytokines needs to be integrated using checkpoints and feedback mechanisms. Since miRNAs are negative regulators of genetic networks, they are well suited to achieve these tasks.
With a few exceptions, most autoimmune diseases are genetically complex polygenic diseases. It is believed that several defects at various checkpoints need to accumulate before full-blown autoimmunity becomes apparent [87, 88]. Genome-wide association studies (GWAS) have been used to identify multiple genetic risk factors in multigenic immune diseases such as multiple sclerosis and type 1 diabetes (T1D) [89–92]. However, individual genes other than the major histocompatibility cluster (MHC) genes contribute to genetically complex diseases by odds ratios of <2-fold [91], suggesting that genetic elements that function as biologic “rheostats” are critical for the proper control of the immune system consistent with a role for miRNAs as potential key players. Importantly, in many instances, the same GWAS-identified genes that fine tune autoimmune susceptibility, result in catastrophic disease when fully deficient (e.g., the interleukin-2 receptor alpha (CD25) or the cytotoxicity T lymphocyte antigen 4 (CTLA-4)) [93, 94]. This implies that protein levels and functional activity of key genes in the immune system need to be tightly controlled. Indeed, haplo-insufficiency, i.e., reduction of certain gene products by 50%, can predispose to autoimmune disease [95, 96]. In B cells, reducing the amount of c-Myb by 25–50% results in impaired B cell development, an observation also shown in mice mildly over-expressing miR-150 where the transgenic miRNA targets c-Myb. On the other hand, miR-150-deficient mice display increased humoral immune responses. Hence, subtle changes in expression of this miRNA and its target(s) can affect normal development and immune function [39].
As outlined above, “sharpening” of mRNA expression would allow the immune system to regulate gene expression in closely related cell types like lymphocyte subsets during development or immune responses. Temporal control of gene expression machinery in lymphocytes is very tight [97, 98], and related cells of similar origin are aligned spatially (in the thymus for instance) before migrating to inflamed tissues. In addition, different cell subsets (i.e., conventional T cells (Tconv) and Treg cells) share similar transcriptomes and proteomes. With the exception of very few genes, these differences are quantitative rather than qualitative [99]. Thus, the abundance of proteins in closely related cell subsets must be tightly controlled, as small changes result in functionally distinct lymphocyte populations that in many cases redirect the cell to take on distinct or even opposing functions. Therefore, miRNAs are candidates as critical regulators of cell subset differentiation and maintenance and, perhaps Tregs, which themselves function as rheostats of the immune system, are poised to reflect the fine tuning manifested by miRNAs. For instance, a limited comparison of miRNA expression in nTreg, FoxP3-transduced, and activated CD4+ murine T cells demonstrated quantitative differences in expression of a common set of miRNAs [34] suggesting that miRNAs regulate T lymphocyte fate specification. Of note, clonal heterogeneity contributes to hematopoietic lineage choice and differentiation [100], and small differences in protein concentrations or activity can decide over digital responses of cell survival versus apoptosis [101]. miRNAs likely contribute to immune homeostasis by tuning the proteome to reduce the stochastic aspects of these important decisions. Thus, we propose that miRNAs serve to sharpen the transcriptome and proteome of lymphocyte subsets to help define and maintain lineage identity, fine tune antigen-receptor signaling, to set response thresholds, and help convert analog into digital signals.
What is the Evidence for Our Hypotheses in the Immune System?
Immune Cell Lineage Stabilization by miRNAs
We have demonstrated that loss of Dicer exclusively in nTreg led to increased loss of the lineage-defining transcription factor FoxP3, and some FoxP3 expressing Treg inappropriately secreted IFN-γ or IL-17 [67]. Several reports have highlighted the plasticity in T cell subset diversity, that genes can be “poised” for transcription, i.e., epigenetic changes associated with activation can be present without detectable transcripts [70–72, 74, 102], and fully differentiated T cell subsets can re-differentiate under inflammatory or lymphopenic conditions [103–107]. Interestingly, individual Th subsets concomitantly have activating and inactivating histone marks for transcription factors, while histone marks at cytokine genes are mutually exclusive, either activating or inactivating [102]. It has been argued that mixed histone marking could allow for plasticity, consistent with the notion that an additional layer of negative regulation is necessary for lineage-determining transcription factors. miRNAs could provide such regulation. In this regard, nTreg have recently been proposed to exist in Th1-, Th2-, and Th17-like subtypes that regulate their respective effector T cell subsets as “class control”. Th1, Th2, and Th17 subtypes of Tregs suppress Th1, Th2, or Th17 effector cells, respectively, but fail to suppress the other classes [108–110]. We propose that subsets of Treg-expressed miRNAs suppress the generation of the other subsets. In support of this model, Tbx21, GATA3, IRF4, Stat6, and RORγt are targeted by at least one miRNA that is highly expressed in Treg (www.targetscan.org and Jeker & Bluestone, unpublished result). This type of miRNA regulation has been termed “binary off-switch” as opposed to the “tuning” regulation described above [19, 81].
miRNAs as Stabilizers of Genetic Networks Under Stress such as Lymphopenia or Inflammation
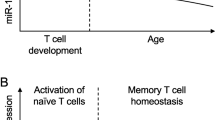
It has been proposed that microRNAs reduce variation of developmental programs and contribute to the development of diverse regulatory networks, particularly under stress conditions such as temperature fluctuations [111]. Such network-stabilizing activity may be important as lymphocytes are constantly exposed to a changing microenvironment during migration by exposure to various cytokines and inflammation with increased blood flow and fluctuations in body temperature. Therefore, miRNAs might both regulate coordinated lymphocyte differentiation [35] and stabilize it under environmental stress [70] (Fig. 1). As highlighted above, Dicer-deficient Treg function was more severely impaired in the presence of inflammation [68]. Similarly, under stress conditions, miR-155−/− Treg had a competitive disadvantage, whereas germline deletion of miR-155 resulted in a surprisingly mild Treg phenotype under homeostatic conditions [42, 45]. In the immune stress setting, miR-155 served to repress suppressor of cytokine signaling 1 (SOCS1), an inhibitor of IL-2 signaling. However, miR-155 is not always essential as IL-2 signaling could be restored in miR-155-deficient Treg in vitro by higher doses of IL-2. Yet, because Treg survival is strongly dependent on IL-2 signaling [112, 113], fine tuning in this pathway is critical. In humans and autoimmune disease susceptible mice, multiple genes of the IL-2/IL-2R pathway are known susceptibility genes for type 1 diabetes (T1D) [89–91]. IL-2 treatment can lead to stabilization of FoxP3 expression in Treg, cure diabetes in mice [114], and restore FoxP3 levels in cultured human Treg [115]. In summary, miR-155 expression in Tregs is not essential to control effector T cells under homeostatic conditions but is important for competitive fitness under stress conditions.
miRNA-mediated network stabilization against stress. A model is presented based on a simple genetic network comprised of signaling molecules S1 and S2 and transcription factor TF that is designed to yield constant transcription irrespective of environmental cues. Under homeostatic conditions, the genetic network leads to gene transcription of gene A because the low miRNA/target mRNA ratio has no measurable effect (a); however, inflammatory signals enhance activity of signaling molecule S2 but also of miRNA miR which dampens the signal S2 to ensure constant transcription of gene A despite inflammation (b). The absence of miR miRNA buffering does not affect the network under homeostatic conditions (c), but under inflammatory conditions, leads to increased transcription of gene A due to defective negative feedback regulation (d). Various modifications can be imagined where miRNAs are not induced but constantly expressed to provide a protective buffer against overshooting protein production of parts of the network. miRNAs may also act on more complex networks with positive or negative feedback loops to enhance, dampen, or stabilize signals. Thus, miRNAs increase precision of genetic networks through fine-tuning expression of network members
miRNA-Mediated Immunologic Threshold Setting
miRNAs can also function to define a certain threshold for signaling. A high miRNA/target ratio can completely repress expression of the target despite transcription [19], thus repressing stochastic transcriptional “noise” during cell fate decisions. A lower miRNA/target ratio can lead to a reduction of gene expression (tuning), and miRNAs can repress expression of target genes without changing their transcription. Arguably, the most sensitive and versatile signal a lymphocyte can receive occurs through its antigen receptor. Cognate peptide–MHC interactions with the corresponding antigen-receptor leads to dramatic consequences: survival and maturation versus death during thymic selection; activation versus quiescence and tolerance in the periphery; and effector cell fate versus memory cell fate during an inflammatory response [116]. How a single antigen receptor can sense an analogous spectrum of ligands with various affinities and convert those into a digital survival or death response has long remained enigmatic. Among many regulatory pathways, miRNAs add an additional layer of regulation by directly influencing the TCR signal [117] and controlling regulatory pathways (e.g., CD28, CTLA-4, PTEN [40], Cbl-b). Recently, several models have been described that attempt to reconcile the paradox that small affinity differences among TCR ligands can have dramatic effects that create a digital response [118–120]. In these models, minor changes in TCR signal strength near the selection threshold are likely to result in selection of a skewed and dangerous TCR repertoire with potentially autoreactive lymphocytes. For instance, miR-181a augments sensitivity to peptide antigens by targeting multiple TCR-related phosphatases, thus affecting thymic selection [117]. Moreover, miR-181a is involved in preventing selection of “forbidden” clones [121]. Finally, miRNA-mediated TCR tuning may contribute to Tconv versus Treg differentiation choices as it has been demonstrated that TCR signal strength regulates FoxP3 expression [122, 123]. Individuals with autoimmune susceptibility might have defective Ag-receptor signal dampening, resulting in a shift of the net activation towards the point of no return for activation allowing a few cells to cross the activation threshold [124] (Fig. 2a). As miRNAs can contribute to setting thresholds, dysregulation of such key miRNAs could underlie autoimmune pathogenesis. PTEN expression, a negative regulator of PI3 kinase and hence TCR signaling, is regulated by the miR-17∼92 cluster. Transgenic overexpression of this miRNA cluster increases T cell proliferation, likely due to enhanced TCR signaling [40]. Moreover, the transgenic mice develop increased autoantibody titers. In this regard, PTEN heterozygous mice display an autoimmune phenotype [95], illustrating that expression of this key-negative regulator of TCR signaling needs to be tightly controlled. It remains to be determined how tolerance is broken in these models and if the autoantibodies are a consequence of uncontrolled proliferation or if the selection of self-reactive lymphocytes is also affected. Nevertheless, overexpression of miR-17-5p, has been described in CD4+ cells from patients with multiple sclerosis [125] suggesting a potential role for miRNAs in these settings.
miRNA-mediated threshold setting in the immune system: preventing unintentional lymphocyte activation. The net activation signal for self-peptides of T cells varies physiologically. a The threshold for activation is not reached by any cell of a given T cell population in healthy (black line) individuals perhaps dependent on TCR dampening by miRNAs. In susceptible individuals (grey line) with impaired miRNA-mediated TCR dampening, the whole T cell population is shifted to the right such that a few cells cross the threshold (dashed line) and become activated to start an autoimmune response. b Alternatively, rather than decreasing net activation of the whole T cell population, the expression of miRNAs reshape the receptor activation curve such that a more narrow range of signals trigger T cells. In this scenario, loss of miRNA expression may broaden the edge of the distribution such that a subset of cells crosses the threshold for activation
In addition to shifting the activation threshold for a given population (Fig. 2a), miRNAs might limit activation signals by shaping the distribution of activation states of a population of cells (Fig. 2b). In resistant individuals, this layer of protection limits the number of cells that “stochastically” cross the threshold of overt self-reactivity. Absence of this buffering would increase the risk of some cells reaching and crossing the tolerance threshold unintentionally leading to autoimmunity.
Conversion of analog signals into digital responses is key in the immune system. Digital responses are important during thymic selection [120]. miRNAs may play a key role in determining the outcome of these events (Fig. 3). miRNAs could prevent or delay a selecting signal under normal conditions; however, when the ratio of miRNA/target is reduced due to target activation by the selecting stimulus, the miRNA no longer suppresses the target efficiently allowing a response. Additional regulatory loops can lead to a very sharp response that overcomes the threshold set by the miRNA feedback. Another set of miRNAs then prevents the response from overshooting its threshold by setting a maximum response level. The effect of a stimulating analog signal into a cell is illustrated in Fig. 3. The presence of miRNAs allows the cell's response to convert into a digital outcome. Absence of such sharpening of the response could result in a skewed Ag-receptor repertoire or dysregulated T cell activation and survival, ultimately contributing to autoimmune disease.
miRNAs can facilitate the conversion of analog signals into digital responses. Through sharpening of gene expression, miRNAs could help to convert analog signals into digital signals. In this scenario, a low stimulus does not lead to a response due to repression by a miRNA (miR-a). If the ratio of miRNA/target(s) decreases because the target is induced, a response occurs rapidly because positive feedback loops have been activated. Overshooting of the response is prevented by other miRNAs (miR-b) that set a threshold at higher protein expression levels limiting the maximum net response. This allows responses to convert from analog into digital signals
Potential Diagnostic and Therapeutic Applications of RNAi, miRNAs, and Artificial Small RNAs (or mimetics)
Several diseases have been probed for differential miRNA expression compared to healthy individuals. miRNA dysregulation and potential function in rheumatoid arthritis and lupus erythematosus has recently been reviewed [126]. miR-146a is upregulated in peripheral blood mononuclear cell (PBMCs) from rheumatoid arthritis patients [127], and patients suffering from multiple sclerosis differentially express miRNAs in their PBMCs and CD4+ T cells [125, 128, 129]. It will also be important to test if accumulations of mutations in miRNA loci (or their mRNA targets) are found in samples from patients with autoimmune disease similar to cancer genomes [130, 131]. Thus, pinpointing individual miRNAs or miRNA signatures dysregulated in autoimmune diseases will be further studied, seeking mechanistic insight into disease pathogenesis, facilitating clinical diagnosis and/or monitoring disease activity, and miRNA modulators might be considered as therapeutics.
It will be important to find the miRNAs that are responsible for stabilizing FoxP3 [67] as these cells are major regulators of autoimmunity. The design of miRNA mimics or inhibitors that stabilize the Treg phenotype may be useful in the treatment of autoimmune diseases as FoxP3 instability has been correlated to autoimmune inflammation [104, 132]. In this regard, small RNA-based therapies might become available in the near future and such therapies might become cell-type specific [133, 134]. Experimental data suggest that small RNA-based therapeutics can be extremely challenging but powerful [135–138]. Recently, a clinical phase Ia study with small synthetic RNAs directed at miR-122 was initiated as a treatment for hepatitis C (www.clinicaltrials.gov NCT00979927). In addition, endogenous miRNA expression is also being targeted to guide transgene expression [139]. Thus, miRNA technology is being exploited on many levels including gene therapy applications where a proof of principle study demonstrated that miRNA-guided gene therapy can induce T cell tolerance [140].
Conclusions
It is now clearly established that miRNAs are critical for proper immune regulation. However, our knowledge of the contribution and mechanisms of individual miRNAs in autoimmune disease and immunity is at a very early stage. Despite lymphocytes being one of the best characterized cell types in mice and humans with extensive knowledge about genetic programs regulating lineage differentiation, a comprehensive small RNA signature is not known for many types of immune cells, with a few exceptions. However, the rapid pace of technologic advance to assess even a single cell's genome-wide transcription combined with sensitive techniques to measure entire proteomes will deliver exciting insight in the coming years. By regulating immune cell development and lineage stabilization, sharpening gene expression, setting thresholds, buffering fluctuations of gene expression to balance genetic networks under stress, miRNAs provide a protective layer to immune homeostasis. Thus, elimination of individual miRNA function will likely not lead to full blown autoimmune disease but may very well increase the susceptibility to autoimmune disease. Thus, miRNAs are likely pieces in the mosaic of autoimmune pathogenesis. Without any doubt, the next decade will be exciting when we start to understand how miRNAs function in lymphocytes and how this knowledge can be exploited diagnostically and therapeutically.
References
Izant JG, Weintraub H. Inhibition of thymidine kinase gene expression by anti-sense RNA: a molecular approach to genetic analysis. Cell. 1984;36:1007–15. doi:0092-8674(84)90050-3 [pii].
Rosenberg UB, Preiss A, Seifert E, Jackle H, Knipple DC. Production of phenocopies by Kruppel antisense RNA injection into Drosophila embryos. Nature. 1985;313:703–6.
Ecker JR, Davis RW. Inhibition of gene expression in plant cells by expression of antisense RNA. Proc Natl Acad Sci U S A. 1986;83:5372–6.
Izant JG, Weintraub H. Constitutive and conditional suppression of exogenous and endogenous genes by anti-sense RNA. Science. 1985;229:345–52.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–54. doi:0092-8674(93)90529-Y [pii].
Fire A et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–11. doi:10.1038/35888.
Lee RC, Ambros V. An extensive class of small RNAs in Caenorhabditis elegans. Science. 2001;294:862–4. doi:10.1126/science.1065329 294/5543/862 [pii].
Lau NC, Lim LP, Weinstein EG, Bartel DP. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science. 2001;294:858–62. doi:10.1126/science.1065062294/5543/858 [pii].
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T. Identification of novel genes coding for small expressed RNAs. Science. 2001;294:853–8. doi:10.1126/science.1064921294/5543/853 [pii].
Bernstein E et al. Dicer is essential for mouse development. Nat Genet. 2003;35:215–7.
Landgraf P et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell. 2007;129:1401–14.
Bartel DP. MicroRNAs, genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Babiarz JE, Ruby JG, Wang Y, Bartel DP, Blelloch R. Mouse ES cells express endogenous shRNAs, siRNAs, and other microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 2008;22:2773–85. doi:22/20/2773 [pii] 10.1101/gad.1705308.
Ruby JG, Jan CH, Bartel DP. Intronic microRNA precursors that bypass Drosha processing. Nature. 2007;448:83–6. doi:nature05983 [pii] 10.1038/nature05983.
Okamura K et al. The Drosophila hairpin RNA pathway generates endogenous short interfering RNAs. Nature. 2008;453:803–6.
Okamura K, Hagen JW, Duan H, Tyler DM, Lai EC. The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell. 2007;130:89–100. doi:S0092-8674(07)00795-7 [pii] 10.1016/j.cell.2007.06.028.
Berezikov E, Chung WJ, Willis J, Cuppen E, Lai EC. Mammalian mirtron genes. Mol Cell. 2007;28:328–36.
Lal A et al. miR-24 inhibits cell proliferation by targeting E2F2, MYC, and other cell-cycle genes via binding to “seedless” 3'UTR microRNA recognition elements. Mol Cell. 2009;35:610–25. doi:S1097-2765(09)00600-5 [pii] 10.1016/j.molcel.2009.08.020.
Bartel DP. Cell. 2009;136:215–33. doi:S0092-8674(09)00008-7 [pii] 10.1016/j.cell.2009.01.002.
Grimson A et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell. 2007;27:91–105. doi:S1097-2765(07)00407-8 [pii] 10.1016/j.molcel.2007.06.017.
Orom UA, Nielsen FC, Lund AH. MicroRNA-10a binds the 5'UTR of ribosomal protein mRNAs and enhances their translation. Mol Cell. 2008;30:460–71. doi:S1097-2765(08)00328-6 [pii] 10.1016/j.molcel.2008.05.001.
Vasudevan S, Tong Y, Steitz JA. Switching from repression to activation: microRNAs can up-regulate translation. Science. 2007;318:1931–4. doi:1149460 [pii] 10.1126/science.1149460.
Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc Natl Acad Sci U S A. 2008;105:1608–13. doi:0707594105 [pii] 10.1073/pnas.0707594105.
Lim LP et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature. 2005;433:769–73. doi:nature03315 [pii] 10.1038/nature03315.
Baek D et al. The impact of microRNAs on protein output. Nature. 2008;455:64–71. doi:nature07242 [pii] 10.1038/nature07242.
Hendrickson DG et al. Concordant regulation of translation and mRNA abundance for hundreds of targets of a human microRNA. PLoS Biol. 2009;7:e1000238. doi:10.1371/journal.pbio.1000238.
Selbach M et al. Widespread changes in protein synthesis induced by microRNAs. Nature. 2008;455:58–63.
Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet. 2008;9:102–14.
Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10:126–39. doi:nrm2632 [pii] 10.1038/nrm2632.
Trabucchi M et al. The RNA-binding protein KSRP promotes the biogenesis of a subset of microRNAs. Nature. 2009;459:1010–14. doi:nature08025 [pii] 10.1038/nature08025.
Chatterjee S, Grosshans H. Active turnover modulates mature microRNA activity in Caenorhabditis elegans. Nature. 2009;461:546–9. doi:nature08349 [pii] 10.1038/nature08349.
Ramachandran V, Chen X. Degradation of microRNAs by a family of exoribonucleases in Arabidopsis. Science. 2008;321:1490–2. doi:321/5895/1490 [pii] 10.1126/science.1163728.
Davis BN, Hilyard AC, Lagna G, Hata A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature. 2008;454:56–61. doi:nature07086 [pii] 10.1038/nature07086.
Cobb BS et al. A role for Dicer in immune regulation. J Exp Med. 2006;203:2519–27.
O'Connell RM, Rao DS, Chaudhuri AA, Baltimore D. Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol. 2010;10:111–22. doi:nri2708 10.1038/nri2708.
Zhou B, Wang S, Mayr C, Bartel DP, Lodish HF. miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc Natl Acad Sci U S A. 2007;104:7080–5. doi:0702409104 [pii] 10.1073/pnas.0702409104.
Chen CZ, Li L, Lodish HF, Bartel DP. MicroRNAs modulate hematopoietic lineage differentiation. Science. 2004;303:83–6.
Vigorito E et al. microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity. 2007;27:847–59. doi:S1074-7613(07)00503-1 [pii] 10.1016/j.immuni.2007.10.009.
Xiao C et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell. 2007;131:146–59.
Xiao C et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat Immunol. 2008;9:405–14.
Rodriguez A et al. Requirement of bic/microRNA-155 for normal immune function. Science. 2007;316:608–11.
Lu L et al. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity. 2009;30:80–91. doi:S1074-7613(08)00556-6 [pii] 10.1016/j.immuni.2008.11.010.
Thai TH et al. Regulation of the germinal center response by microRNA-155. Science. 2007;316:604–8.
Dorsett Y et al. MicroRNA-155 suppresses activation-induced cytidine deaminase-mediated Myc-Igh translocation. Immunity. 2008;28:630–8.
Kohlhaas S et al. Cutting edge: the Foxp3 target miR-155 contributes to the development of regulatory T cells. J Immunol. 2009;182:2578–82. doi:182/5/2578 [pii] 10.4049/jimmunol.0803162.
Teng G et al. MicroRNA-155 is a negative regulator of activation-induced cytidine deaminase. Immunity. 2008;28:621–9. doi:S1074-7613(08)00188-X [pii] 10.1016/j.immuni.2008.03.015.
Baltimore D, Boldin MP, O'Connell RM, Rao DS, Taganov KD. MicroRNAs: new regulators of immune cell development and function. Nat Immunol. 2008;9:839–45. doi:ni.f.209 [pii] 10.1038/ni.f.209.
Xiao C, Rajewsky K. MicroRNA control in the immune system: basic principles. Cell. 2009;136:26–36. doi:S0092-8674(08)01633-4 [pii] 10.1016/j.cell.2008.12.027.
Lindsay MA. microRNAs and the immune response. Trends Immunol. 2008;29:343–51. doi:S1471-4906(08)00131-2 [pii] 10.1016/j.it.2008.04.004.
Vinuesa CG, Rigby RJ, Yu D. Logic and extent of miRNA-mediated control of autoimmune gene expression. Int Rev Immunol. 2009;28:112–38. doi:10.1080/08830180902934909 [pii].
Lang KS et al. Toll-like receptor engagement converts T-cell autoreactivity into overt autoimmune disease. Nat Med. 2005;11:138–45. doi:nm1176 [pii] 10.1038/nm1176.
Lang KS, Burow A, Kurrer M, Lang PA, Recher M. The role of the innate immune response in autoimmune disease. J Autoimmun. 2007;29:206–12. doi:S0896-8411(07)00093-5 [pii] S0896-8411(07)00093-5.
Iwasaki A, Medzhitov R. Regulation of adaptive immunity by the innate immune system. Science. 2010;327:291–5. doi:327/5963/291 [pii] 10.1126/science.1183021.
Martinez J, Busslinger M. Life beyond cleavage: the case of Ago2 and hematopoiesis. Genes Dev. 2007;21:1983–8. doi:10.1101/gad.1591407 [pii] 10.1101/gad.1591407.
Wienholds E, Koudijs MJ, van Eeden FJ, Cuppen E, Plasterk RH. The microRNA-producing enzyme Dicer1 is essential for zebrafish development. Nat Genet. 2003;35:217–8. doi:10.1038/ng1251 [pii].
O’Carroll D et al. A Slicer-independent role for Argonaute 2 in hematopoiesis and the microRNA pathway. Genes Dev. 2007;21:1999–2004. doi:gad.1565607 [pii] 10.1101/gad.1565607.
Harfe BD, McManus MT, Mansfield JH, Hornstein E, Tabin CJ. The RNaseIII enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb. Proc Natl Acad Sci U S A. 2005;102:10898–903.
Kanellopoulou C et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev. 2005;19:489–501.
Chong MM, Rasmussen JP, Rudensky AY, Littman DR. The RNAseIII enzyme Drosha is critical in T cells for preventing lethal inflammatory disease. J Exp Med. 2008;205:2005–17. doi:em.20081219 [pii] 10.1084/jem.20081219.
Wang Y, Medvid R, Melton C, Jaenisch R, Blelloch R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat Genet. 2007;39:380–5. doi:ng1969 [pii] 10.1038/ng1969.
Yi R et al. DGCR8-dependent microRNA biogenesis is essential for skin development. Proc Natl Acad Sci U S A. 2009;106:498–502. doi:0810766105 [pii] 10.1073/pnas.0810766105.
Koralov SB et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell. 2008;132:860–74.
Fedeli M et al. Dicer-dependent microRNA pathway controls invariant NKT cell development. J Immunol. 2009;183:2506–12. doi:jimmunol.0901361 [pii] 10.4049/jimmunol.0901361.
Zhou L et al. Tie2cre-induced inactivation of the miRNA-processing enzyme Dicer disrupts invariant NKT cell development. Proc Natl Acad Sci U S A. 2009;106:10266–71. doi:0811119106 [pii] 10.1073/pnas.0811119106.
Cobb BS et al. T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. J Exp Med. 2005;201:1367–73.
Muljo SA et al. Aberrant T cell differentiation in the absence of Dicer. J Exp Med. 2005;202:261–9.
Zhou X et al. Selective miRNA disruption in T reg cells leads to uncontrolled autoimmunity. J Exp Med. 2008;205(9):1983–91.
Liston A, Lu LF, O'Carroll D, Tarakhovsky A, Rudensky AY. Dicer-dependent microRNA pathway safeguards regulatory T cell function. J Exp Med. 2008;205(9):1993–2004.
Brunkow ME et al. Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat Genet. 2001;27:68–73.
Zhou X, Bailey-Bucktrout S, Jeker LT, Bluestone JA. Plasticity of CD4(+) FoxP3(+) T cells. Curr Opin Immunol. 2009;21:281–5. doi:S0952-7915(09)00087-9 [pii] 10.1016/j.coi.2009.05.007 S0952-7915(09)00087-9 [pii] 10.1016/j.coi.2009.05.007.
Bluestone JA, Mackay CR, O'Shea JJ, Stockinger B. The functional plasticity of T cell subsets. Nat Rev Immunol. 2009;9:811–6. doi:nri2654 [pii] 10.1038/nri2654.
O'Shea JJ, Paul WE. Mechanisms underlying lineage commitment and plasticity of helper CD4+ T cells. Science. 2010;327:1098–102. doi:327/5969/1098 [pii] 10.1126/science.1178334.
Locksley RM. Nine lives: plasticity among T helper cell subsets. J Exp Med. 2009;206:1643–6. doi:jem.20091442 [pii] 10.1084/jem.20091442.
Zhou L, Chong MM, Littman DR. Plasticity of CD4+ T cell lineage differentiation. Immunity. 2009;30:646–55. doi:S1074-7613(09)00198-8 [pii] 10.1016/j.immuni.2009.05.001.
Miska EA et al. Most Caenorhabditis elegans microRNAs are individually not essential for development or viability. PLoS Genet. 2007;3:e215.
Wang Y et al. Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nat Genet. 2008;40:1478–83. doi:ng.250 [pii] 10.1038/ng.250.
Alvarez-Saavedra E, Horvitz H. Many families of C. elegans micrornas are not essential for development or viability. Curr Biol. 2010;20:367–73. doi:S0960-9822(09)02214-3 [pii] 10.1016/j.cub.2009.12.051.
Zhao Y et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell. 2007;129:303–17. doi:S0092-8674(07)00398-4 [pii] 10.1016/j.cell.2007.03.030.
Ventura A et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell. 2008;132:875–86. doi:S0092-8674(08)00267-5 [pii] 10.1016/j.cell.2008.02.019.
Stark A, Brennecke J, Bushati N, Russell RB, Cohen SM. Animal microRNAs confer robustness to gene expression and have a significant impact on 3′UTR evolution. Cell. 2005;123:1133–46. doi:S0092-8674(05)01272-9 [pii] 10.1016/j.cell.2005.11.023.
Bartel DP, Chen CZ. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet. 2004;5:396–400.
Grimson A et al. Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals. Nature. 2008;455:1193–7. doi:nature07415 [pii] 10.1038/nature07415.
Sharp PA. The centrality of RNA. Cell. 2009;136:577–80. doi:S0092-8674(09)00143-3 [pii] 10.1016/j.cell.2009.02.007.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20. doi:S0092867404012607 [pii] 10.1016/j.cell.2004.12.035.
Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19:92–105. doi:gr.082701.108 [pii] 10.1101/gr.082701.108.
Kaech SM, Wherry EJ, Ahmed R. Effector and memory T-cell differentiation: implications for vaccine development. Nat Rev Immunol. 2002;2:251–62. doi:10.1038/nri778.
Rioux JD, Abbas AK. Paths to understanding the genetic basis of autoimmune disease. Nature. 2005;435:584–9. doi:nature03723 [pii] 10.1038/nature03723.
Goodnow CC, Sprent J, Fazekas de St Groth B, Vinuesa CG. Cellular and genetic mechanisms of self tolerance and autoimmunity. Nature. 2005;435:590–7.
Lowe CE et al. Large-scale genetic fine mapping and genotype–phenotype associations implicate polymorphism in the IL2RA region in type 1 diabetes. Nat Genet. 2007;39:1074–82.
Barrett JC et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet. 2009. doi:ng.381 [pii] 10.1038/ng.381.
Concannon P, Rich SS, Nepom GT. Genetics of type 1A diabetes. N Engl J Med. 2009;360:1646–54. doi:360/16/1646 [pii] 10.1056/NEJMra0808284.
Hafler DA et al. Risk alleles for multiple sclerosis identified by a genomewide study. N Engl J Med. 2007;357:851–62. doi:NEJMoa073493 [pii] 10.1056/NEJMoa073493.
Willerford DM et al. Interleukin-2 receptor alpha chain regulates the size and content of the peripheral lymphoid compartment. Immunity. 1995;3:521–30. doi:1074-7613(95)90180-9 [pii].
Tivol EA et al. Loss of CTLA-4 leads to massive lymphoproliferation and fatal multiorgan tissue destruction, revealing a critical negative regulatory role of CTLA-4. Immunity. 1995;3:541–7.
Di Cristofano A et al. Impaired Fas response and autoimmunity in Pten+/- mice. Science. 1999;285:2122–5. doi:7853 [pii].
Bouillet P et al. Proapoptotic Bcl-2 relative Bim required for certain apoptotic responses, leukocyte homeostasis, and to preclude autoimmunity. Science. 1999;286:1735–8. doi:8026 [pii].
Rothenberg EV, Taghon T. Molecular genetics of T cell development. Annu Rev Immunol. 2005;23:601–49.
Rothenberg EV, Moore JE, Yui MA. Launching the T-cell-lineage developmental programme. Nat Rev Immunol. 2008;8:9–21. doi:nri2232 [pii] 10.1038/nri2232.
Feuerer M, Hill JA, Mathis D, Benoist C. Foxp3+ regulatory T cells: differentiation, specification, subphenotypes. Nat Immunol. 2009;10:689–95. doi:ni.1760 [pii] 10.1038/ni.1760.
Chang HH, Hemberg M, Barahona M, Ingber DE, Huang S. Transcriptome-wide noise controls lineage choice in mammalian progenitor cells. Nature. 2008;453:544–7. doi:nature06965 [pii] 10.1038/nature06965.
Spencer SL, Gaudet S, Albeck JG, Burke JM, Sorger PK. Non-genetic origins of cell-to-cell variability in TRAIL-induced apoptosis. Nature. 2009;459:428–32. doi:nature08012 [pii] 10.1038/nature08012.
Wei G et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells. Immunity. 2009;30:155–67. doi:S1074-7613(08)00555-4 [pii] 10.1016/j.immuni.2008.12.009.
Yang XO et al. Molecular antagonism and plasticity of regulatory and inflammatory T cell programs. Immunity. 2008;29:44–56. doi:S1074-7613(08)00270-7 [pii] 10.1016/j.immuni.2008.05.007.
Zhou X et al. Instability of the transcription factor Foxp3 leads to the generation of pathogenic memory T cells in vivo. Nat Immunol. 2009;10:1000–7. doi:ni.1774 [pii] 10.1038/ni.1774.
Lal G et al. Epigenetic regulation of Foxp3 expression in regulatory T cells by DNA methylation. J Immunol. 2009;182:259–73. doi:182/1/259 [pii].
Tsuji M et al. Preferential generation of follicular B helper T cells from Foxp3+ T cells in gut Peyer's patches. Science. 2009;323:1488–92. doi:323/5920/1488 [pii] 10.1126/science.1169152.
Oldenhove G et al. Decrease of Foxp3+ Treg cell number and acquisition of effector cell phenotype during lethal infection. Immunity. 2009;31:772–86. doi:S1074-7613(09)00454-3 [pii] 10.1016/j.immuni.2009.10.001.
Zheng Y et al. Regulatory T-cell suppressor program co-opts transcription factor IRF4 to control T(H)2 responses. Nature. 2009;458:351–6. doi:nature07674 [pii] 10.1038/nature07674.
Koch MA et al. The transcription factor T-bet controls regulatory T cell homeostasis and function during type 1 inflammation. Nat Immunol. 2009;10:595–602. doi:ni.1731 [pii] 10.1038/ni.1731.
Chaudhry A et al. CD4+ regulatory T cells control TH17 responses in a Stat3-dependent manner. Science. 2009;326:986–91. doi:1172702 [pii] 10.1126/science.1172702.
Li X, Cassidy JJ, Reinke CA, Fischboeck S, Carthew RW. A microRNA imparts robustness against environmental fluctuation during development. Cell. 2009;137:273–82. doi:S0092-8674(09)00315-8 [pii] 10.1016/j.cell.2009.01.058.
Malek TR, Bayer AL. Tolerance, not immunity, crucially depends on IL-2. Nat Rev Immunol. 2004;4:665–74. doi:10.1038/nri1435nri1435 [pii].
Setoguchi R, Hori S, Takahashi T, Sakaguchi S. Homeostatic maintenance of natural Foxp3(+) CD25(+) CD4(+) regulatory T cells by interleukin (IL)-2 and induction of autoimmune disease by IL-2 neutralization. J Exp Med. 2005;201:723–35.
Tang Q et al. Central role of defective interleukin-2 production in the triggering of islet autoimmune destruction. Immunity. 2008;28:687–97.
Long SA et al. Defects in IL-2R signaling contribute to diminished maintenance of FOXP3 expression in CD4(+)CD25(+) regulatory T-cells of type 1 diabetic subjects. Diabetes. 2010;59:407–15. doi:db09-0694 [pii] 10.2337/db09-0694.
Teixeiro E et al. Different T cell receptor signals determine CD8+ memory versus effector development. Science. 2009;323:502–5. doi:323/5913/502 [pii] 10.1126/science.1163612.
Li QJ et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell. 2007;129:147–61.
Naeher D et al. A constant affinity threshold for T cell tolerance. J Exp Med. 2007;204:2553–9. doi:jem.20070254 [pii] 10.1084/jem.20070254.
Daniels MA et al. Thymic selection threshold defined by compartmentalization of Ras/MAPK signalling. Nature. 2006;444:724–9. doi:nature05269 [pii] 10.1038/nature05269.
Palmer E, Naeher D. Affinity threshold for thymic selection through a T-cell receptor–co-receptor zipper. Nat Rev Immunol. 2009;9:207–13. doi:nri2469 [pii] 10.1038/nri2469.
Ebert PJ, Jiang S, Xie J, Li QJ, Davis MM. An endogenous positively selecting peptide enhances mature T cell responses and becomes an autoantigen in the absence of microRNA miR-181a. Nat Immunol. 2009;10:1162–9. doi:ni.1797 [pii] 10.1038/ni.1797.
Sauer S et al. T cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc Natl Acad Sci U S A. 2008;105:7797–802. doi:0800928105 [pii] 10.1073/pnas.0800928105.
Haxhinasto S, Mathis D, Benoist C. The AKT-mTOR axis regulates de novo differentiation of CD4+Foxp3+ cells. J Exp Med. 2008;205:565–74. doi:jem.20071477 [pii] 10.1084/jem.20071477.
Germain RN. The art of the probable: system control in the adaptive immune system. Science. 2001;293:240–5. doi:10.1126/science.1062946 293/5528/240 [pii].
Lindberg RL, Hoffmann F, Mehling M, Kuhle J, Kappos L. Altered expression of miR-17-5p in CD4(+) lymphocytes of relapsing–remitting multiple sclerosis patients. Eur J Immunol. 2010;40:888–98. doi:10.1002/eji.200940032.
Pauley KM, Cha S, Chan EK. MicroRNA in autoimmunity and autoimmune diseases. J Autoimmun. 2009;32:189–94. doi:S0896-8411(09)00027-4 [pii] 10.1016/j.jaut.2009.02.012.
Pauley KM et al. Upregulated miR-146a expression in peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Res Ther. 2008;10:R101. doi:ar2493 [pii] 10.1186/ar2493.
Du C et al. MicroRNA miR-326 regulates TH-17 differentiation and is associated with the pathogenesis of multiple sclerosis. Nat Immunol. 2009;10:1252–9. doi:ni.1798 [pii] 10.1038/ni.1798.
Otaegui D et al. Differential micro RNA expression in PBMC from multiple sclerosis patients. PLoS ONE. 2009;4:e6309. doi:10.1371/journal.pone.0006309.
Pleasance ED et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature. 2010;463:191–6. doi:nature08658 [pii] 10.1038/nature08658.
Pleasance ED et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature. 2010;463:184–90. doi:nature08629 [pii] 10.1038/nature08629.
Wan YY, Flavell RA. Regulatory T-cell functions are subverted and converted owing to attenuated Foxp3 expression. Nature. 2007;445:766–70. doi:nature05479 [pii] 10.1038/nature05479.
Peer D, Shimaoka M. Systemic siRNA delivery to leukocyte-implicated diseases. Cell Cycle. 2009;8:853–9. doi:7936 [pii].
Kumar P et al. T cell-specific siRNA delivery suppresses HIV-1 infection in humanized mice. Cell. 2008;134:577–86. doi:S0092-8674(08)00821-0 [pii] 10.1016/j.cell.2008.06.034.
Krutzfeldt J et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature. 2005;438:685–9.
Elmen J et al. LNA-mediated microRNA silencing in non-human primates. Nature. 2008;452:896–9. doi:nature06783 [pii] 10.1038/nature06783.
Kota J et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009;137:1005–17. doi:S0092-8674(09)00446-2 [pii] 10.1016/j.cell.2009.04.021.
Castanotto D, Rossi JJ. The promises and pitfalls of RNA-interference-based therapeutics. Nature. 2009;457:426–33. doi:nature07758 [pii] 10.1038/nature07758.
Brown BD et al. Endogenous microRNA can be broadly exploited to regulate transgene expression according to tissue, lineage and differentiation state. Nat Biotechnol. 2007;25:1457–67. doi:nbt1372 [pii] 10.1038/nbt1372.
Annoni A et al. In vivo delivery of a microRNA-regulated transgene induces antigen-specific regulatory T cells and promotes immunologic tolerance. Blood. 2009;114:5152–61. doi:blood-2009-04-214569 [pii] 10.1182/blood-2009-04-214569.
Acknowledgments
We thank K. Mark Ansel and Ed Palmer for constructive criticism and helpful suggestions on this manuscript. This work was supported by a JDRF Scholar award and NIH grants U19 AI056388 to JAB and fellowships from the Novartis Foundation (formerly Ciba-Geigy Jubilee Foundation) and the Swiss Foundation for Grants in Biology and Medicine (SFGBM) PASMP3-124274/1 and Swiss National Science Foundation to LTJ.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s10875-010-9428-z
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Jeker, L.T., Bluestone, J.A. Small RNA Regulators of T Cell-Mediated Autoimmunity. J Clin Immunol 30, 347–357 (2010). https://doi.org/10.1007/s10875-010-9392-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-010-9392-7