Abstract
In vivo associations of respiratory complexes forming higher supramolecular structures are generally accepted nowadays. Supercomplexes (SC) built by complexes I, III and IV and the so-called respirasome (I/III2/IV) have been described in mitochondria from several model organisms (yeasts, mammals and green plants), but information is scarce in other lineages. Here we studied the supramolecular associations between the complexes I, III, IV and V from the secondary photosynthetic flagellate Euglena gracilis with an approach that involves the extraction with several mild detergents followed by native electrophoresis. Despite the presence of atypical subunit composition and additional structural domains described in Euglena complexes I, IV and V, canonical associations into III2/IV, III2/IV2 SCs and I/III2/IV respirasome were observed together with two oligomeric forms of the ATP synthase (V2 and V4). Among them, III2/IV SC could be observed by electron microscopy. The respirasome was further purified by two-step liquid chromatography and showed in-vitro oxygen consumption independent of the addition of external cytochrome c.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
ATP production by oxidative phosphorylation (OXPHOS) is a key process in eukaryotic energetic metabolism. In this process, the respiratory chain complexes NADH:ubiquinone oxidoreductase (complex I), succinate:ubiquinone oxidoreductase (complex II), ubiquinol:cytochrome c oxidoreductase (complex III) and cytochrome c oxidase (complex IV) transfer electrons from NADH or succinate to oxygen and, except for complex II, establish an electrochemical proton gradient across the inner mitochondrial membrane (proton-motive force). Two mobile electron carriers, ubiquinone and cytochrome c, connect the electron flow between complex I or II with complex III, and complex III with IV, respectively. An additional complex, the ATP synthase (complex V), utilizes the energy of the proton-motive force to synthetize ATP.
The organization of the OXPHOS complexes is generally discussed in terms of two extreme models, the “fluid state” where all membrane proteins and redox components are in constant and independent diffusional motion (Hackenbrock et al. 1986) in agreement with the “fluid mosaic model” (Singer and Nicolson 1972) and the “solid state” model which proposes that all the complexes are associated in one functional unit (Keilin and Hartree 1947; Slater 2003). The first suggestion that the OXPHOS complexes can associate with each other in larger structures named supercomplexes (SC) was brought to light based on pioneering blue native electrophoresis experiments (Schägger and Pfeiffer 2000). With the advance of techniques to obtain larger macromolecular protein structures, e.g. cryo-electron microscopy and cryotomography, nowadays, the existence of mitochondrial SC is generally accepted (Wittig and Schägger 2009; Chaban et al. 2014; Genova and Lenaz 2014; Letts et al. 2016; Lobo-Jarne and Ugalde 2017). The reasons or advantages of these association are still in debate, they include a more efficient transport of electrons to minimize the generation of reactive oxygen species during electron transfer reactions (Winge 2012), the regulation of the mitochondrial metabolism in response to different stimuli, carbon sources, or stress conditions (Acin-Perez and Enriquez 2014; Genova and Lenaz 2014) or provide a kinetic advantage by substrate channeling specially maintaining a dedicated quinol pool (Genova and Lenaz 2014; Lenaz et al. 2016; Milenkovic et al. 2017).
The family of the rotary ATPases originates from a common evolutionary ancestor (Cross and Taiz 1990; Cross and Müller 2004). This superfamily comprises the vacuolar H+-translocating V1Vo-ATPases (V-ATPase), the archaeal A1Ao-ATPases (A-ATPase) and the bacterial, plastid and mitochondrial F1Fo-ATPases (F-ATPase) (Muench et al. 2011). The overall structure and subunit composition of the bacterial and mitochondrial F-ATPases from Opisthokonts (i.e. Fungi/Metazoa group), is overall well conserved (Kühlbrandt 2019). The mitochondrial F-ATPase presents a dimeric nature with a V-shape architecture that folds the inner membrane to form the mitochondrial cristae (Strauss et al. 2008; Davies et al. 2011). In contrast, no bacterial or plastidic dimeric enzyme has been reported to date. Accordingly, the most variable structure among the different species is the peripheral stalk while the most conserved regions between bacterial and opisthokont enzymes correspond to the catalytic core (α3β3), the peripheral stalk binding subunit (OSCP), the central rotor and proton-translocation region (γ, δ, ε, a and c-ring) (Colina-Tenorio et al. 2018). The study of mitochondrial F-ATPase of other eukaryotic lineages (e.g. ciliates (Alveolata), chlorophyceae (Archeaeplastida), euglenozoan (Excavata)) revealed highly divergent subunit compositions of the peripheral stalk between the lineages (Zíková et al. 2009; Balabaskaran Nina et al. 2010; Allegretti et al. 2015; Mühleip et al. 2016, 2017; Yadav et al. 2017; Miranda-Astudillo et al. 2018a; Salunke et al. 2018; Colina-Tenorio et al. 2019). These various peripheral stalks are usually more robust, and give rise to highly stable dimers with various geometries.
Euglena gracilis is a secondary photosynthetic flagellate that arose from an endosymbiosis between a green alga and an ancient phagotroph (Gibbs 1981; Turmel et al. 2009). Euglenids, together with other heterotrophic flagellates like Symbiontida (free-living flagellates found in low-oxygen marine sediments), Diplonemea (free-living marine flagellates) and Kinetoplastida (free-living and parasitic flagellates, e.g. Trypanosoma) form the monophyletic Euglenozoa group (Burki 2014; Zakrys et al. 2017). E. gracilis has a mitochondrial electron transfer system constituted by the OXPHOS complexes (Complexes I - IV) and also exhibits alternative electron pathways. These pathways involve an alternative oxidase (AOX) sensitive to diphenylamine, salicylhydroxamic acid (SHAM), n-propyl gallate and disulfiram (Sharpless and Butow 1970a; Benichou et al. 1988; Moreno-Sánchez et al. 2000), a CIII-like complex resistant to antimycin A (Sharpless and Butow 1970b) and an enzyme catalyzing a cytochrome c oxidase activity partially insensitive to cyanide in the presence of L-lactate (Moreno-Sánchez et al. 2000).
The subunit composition of the OXPHOS complexes among the Euglenozoa species includes the conserved canonical subunits, mainly related with the catalytic activity of each complex, but also a series of lineage-specific atypical subunits (Speijer et al. 1997; Morales et al. 2009; Perez et al. 2014; Verner et al. 2015; Yadav et al. 2017; Miranda-Astudillo et al. 2018b). This divergent subunit composition leads notably to the presence of atypical domains observed in the structures of complexes I, IV and V2 (Duarte and Tomás 2014; Mühleip et al. 2017; Yadav et al. 2017; Miranda-Astudillo et al. 2018b; Montgomery et al. 2018). In the present work, we studied the consequences of these atypical structures on the supramolecular association of the OXPHOS complexes in E. gracilis by native electrophoresis and single-particle electron microscopy, additionally, the in-vitro oxygen consumption activity of the purified respirasome complex (i.e. I/III2/IV) was determined.
Materials and methods
Algal strain, growth conditions and mitochondria isolation
E. gracilis (SAG 1224–5/25) was obtained from the University of Göttingen (Sammlung von Algenkulturen, Germany). Cells were grown in liquid mineral Tris-minimum-phosphate medium (TMP) pH 7.0 supplemented with a mix of vitamins (biotin 10−7%, B12 vitamin 10−7% and B1 vitamin 2 × 10−5% (w/v)). Ethanol 1% was used as carbon source. The cultures were grown in the dark under orbital agitation at 25 °C and collected in the middle of the logarithmic phase. Mitochondria were prepared as described in (Yadav et al. 2017) and stored at −80 °C until use. Protein concentration was determined by the Bradford method (Biorad).
Native and denaturing protein electrophoresis
All steps were performed at 4 °C. n-dodecyl-β-D-maltoside (DDM, Sigma), digitonin (Sigma) and the synthetic drop-in substitute for digitonin glyco-diosgenin (GDN101, Anatrace) were used for the solubilization in a 4.0, 8.0 and 8.0 g detergent/mitochondrial protein ratio, respectively. Final concentrations of detergent were 3.2% or 6.4% in solubilization buffer (SB) containing 50 mM Tris-HCl, 1.5 mM MgSO4, 100 mM NaCl, 10% glycerol, 1 mM phenylmethylsulfonyl fluoride (PMSF) and 50 μg/mL tosyl-lysyl-chloromethylketone (TLCK) (pH 8.4). The mixture was incubated with gentle stirring for 30 min, and centrifuged at 90,000×g for 30 min as in (Miranda-Astudillo et al. 2018b). After discarding the insoluble material, the solubilized complexes were subjected to BN-PAGE in 3%–10% acrylamide gradient gels (Schägger 1994a), 0.05% digitonin was added in the acrylamide gradient if digitonin-extracted sample was loaded (Wittig and Schägger 2005).
To determine the molecular masses of the protein bands, the well characterized mitochondrial complexes from the chlorophycean alga Polytomella sp. were used as molecular mass markers (Miranda-Astudillo et al. 2018a). The logarithm of the distance migrated from each complex was interpolated into a Log (distance migrated) versus size (kDa) regression of the molecular markers (R2 = 0.9939) (Fig. S1 Suppl. Information). In-gel ATPase and complex I activities were carried out as in (Yadav et al. 2017; Miranda-Astudillo et al. 2018a). Denaturing 2D SDS Tricine-PAGE was carried out in 12% polyacrylamide gels as reported (Schägger 1994b). Two dimensional BN/BN-PAGE gels were carried out as previously described (Schägger and Pfeiffer 2001; Wittig and Schägger 2008).
Respirasome complex (I/III2/IV) purification by liquid chromatography
All steps were performed at 4 °C. Thirty milligrams of mitochondria were solubilized with GDN101 as described above. The mixture was incubated with gentle stirring for 60 min, and centrifuged at 90,000×g for 30 min. The supernatant was diluted in 3 volumes of SB without NaCl and supplemented with GDN101 0.01%. The sample was loaded onto an anion exchange column (Source 15Q 5/50, Volume column (VC): 1 mL) connected to an ÄKTA monitor UPC-900 Workstation (GE Healthcare Life Sciences) equilibrated with the same buffer and washed until a constant baseline was obtained. The bound proteins were eluted with a linear 0–500 mM NaCl (20 VC) in the same buffer supplemented with 0.01% GDN, 0.5 mL fractions were collected and analyzed by BN-PAGE.
The fractions corresponding to the respirasome (I/III2/IV) together with the respiratory supercomplexes (III2/IV1–2) were pooled and concentrated with an Amicon Ultra-15 Centrifugal Filter 100 kDa (EMD Millipore) to a final volume of 500 μL and injected to a Superose 6 10/300 (GE Healthcare Life Sciences) previously equilibrated with SB buffer containing NaCl 200 mM and GDN 0.01%. The elution was carried out at 0.25 mL/min, 0.5 mL fractions were collected and visualized by BN-PAGE. The samples enriched with mitochondrial respirasome complex were pooled and stored at −70 °C until use.
Differential spectroscopy of the purified respirasome
Absorption spectra from purified respirasome were measured at 25 °C in a Cary 60 UV-Vis spectrophotometer (Agilent Technologies). Differential spectrum was obtained as the sodium dithionite reduced spectrum minus the potassium ferricyanide oxidized spectrum as described in (Mukai et al. 1989).
Oxygen consumption of the purified respirasome
Oxygen consumption was assessed in a YSI model 5300 oxygraph equipped with a Clark-Type electrode as described in (Miranda-Astudillo et al. 2018a). The reaction vessel was a 200 μL water-jacketed chamber maintained at 30 °C. The activity buffer contained MOPS 50 mM, NaCl 100 mM, GDN 0.01% (pH 7.2). NADH 5 mM was used as electron donor and 2,3-dimethoxy-5-methyl-p-benzoquinone (5 mM) was used to complete the electron transfer chain. The reaction was initiated with the addition of 100 μg of the purified respirasome. Specific inhibitors for complex I (rotenone 500 μM) and complex III (antimycin A 100 μM and myxothiazol 100 μM) were evaluated.
Supercomplexes structure modelling
The crystal structures from chicken dimeric complex III (PDB: 4U3F (Hao et al. 2015)) and the monomeric bovine complex IV together with the cytochrome c (PDB: 5IY5 (Shimada et al. 2017)) were aligned with the corresponding chains in the mammalian respirasome model (PDB 5GUP (Wu et al. 2016) and fit into the density in the electronic map obtained from mammalian respirasome (EMD: 9539 (Wu et al. 2016)). Both structures were used together as a unique coupled model to interpret the projections from the Euglena III2/IV SC.
The cryo-EM structures from dimeric ATP synthase from E. gracilis (PDB: 6TDU (Mühleip et al. 2019)) were fit inside the 27.5 Å 3D map from ribbon of ATP synthases (three dimers) obtained by electron cryotomography and subtomogram averaging from intact inner mitochondrial membranes (EMD-3559 (Mühleip et al. 2017)). All the structure fitting and the images were generated using the UCSF Chimera (https://www.cgl.ucsf.edu/chimera/) (Pettersen et al. 2004).
Results
ATPase oligomers and respiratory supercomplexes in E. gracilis
Mitochondria from dark-grown E. gracilis were treated with mild detergents: 4.0 g n-dodecyl-β-D-maltoside (DDM)/g protein (3.2% w/v) or 8.0 g digitonin/g protein (6.4% w/v). The native protein complexes were then subjected to a 3–10% acrylamide gradient BN-PAGE. When DDM was used to solubilize the complexes, four prominent bands ranging from 460 to 2200 kDa were found in the Coomassie-stained gel (Fig. 1 lane 1). They correspond to the dimeric complex V (V2), monomeric complex I (I), dimeric complex III (III2), and monomeric complex IV (IV) (Perez et al. 2014; Yadav et al. 2017; Miranda-Astudillo et al. 2018b). By contrast, when digitonin was used, four additional main bands ranging between 970 kDa and 5.2 MDa were observed (Fig. 1 lane 2), which may represent supramolecular associations between the OXPHOS complexes (i.e. supercomplexes, SC). Additionally, we tested a synthetic digitonin substitute (GDN101, Anatrace). Compared to digitonin, this substitute had a different impact on SCs solubilization: the 0.97, 1.2, 4.2 and 5.2 MDa bands are fainter while the 2.2 MDa band is more prominent (Fig. S2 lane 2 Suppl. Information).
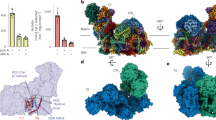
ATPase oligomers and respiratory supercomplexes in E. gracilis. Isolated mitochondria were solubilized with the indicated detergent: n-dodecyl-β-D-maltoside (DDM) at 4.0 g/ g protein or digitonin (Dig) at 8.0 g/g protein, after removing the insoluble material, each sample was resolved by BN-PAGE in a 3–10% polyacrylamide gradient gel. a Coomassie-stained gel showing the main bands with DDM- and digitonin-solubilization. b In-gel NADH-dehydrogenase activity; the BN-gel was incubated in the presence of NADH and Nitro blue tetrazolium chloride (NBT). c Detection of in-gel ATPase activity. The gel was incubated with ATP, MgSO4 and Pb(NO3)2. The determined molecular mass (kDa) of each isolated complex or supercomplexes is indicated. Nomenclature used: I, III2 and IV for the corresponding mitochondrial complexes, V2 and V4 for the dimeric and tetrameric ATP synthase respectively. Supercomplexes were: III2/IV, III2/IV2, the so-called “respirasome” association I/III2/IV and the putative Vx/I association, their stoichiometries are indicated as subindexes
Composition of E. gracilis supercomplexes
To get insight into the composition of the newly identified protein bands, lanes with digitonin- and DDM- solubilized complexes were used to perform in-gel staining for complex I and complex V activities (Fig. 1b and c). In the case of digitonin solubilization, three complex I-stained bands were observed (1.4, 2.2 and 5.2 MDa, respectively), the lower and more intense band matches with the monomeric complex I in DDM lane (1.4 MDa) while the band above (2.2 MDa) probably corresponds to the association of complex I with dimeric complex III or complex IV. This band very often co-localized with dimeric complex V (Fig. 1 lanes 4 and 6). On the other hand, three digitonin-solubilized bands exhibited ATPase activity. The lower one has the same molecular mass as the V2 in DDM lane (2.2 MDa), the second prominent band might correspond to a tetrameric complex V (4.2 MDa) which is also observed as a faint band in DDM solubilisation (Fig. 1c lanes 5 and 6). Notably, the faint upper band at 5.2 MDa presents both complex V and complex I activity staining. Finally, two digitonin-solubilized bands without CI or CV activity are observed below complex I and above dimeric complex III at 1200 kDa and 970 kDa (Fig. 1 lanes 1 and 2), and may correspond to the previously described associations of complex III with IV (III2/IV and III2/IV2) in Euglena mitochondria (Perez et al. 2014). To discard that these supramolecular associations involving complex I or complex V are due to an incomplete solubilisation, Euglena mitochondria were solubilized with increasing digitonin concentrations (up to 12 g digitonin/g protein, 9.6% w/v). All the associations involving CI or CV described above, including putative SC I + III + IV and 5.2 MDa bands were stable at the highest concentration of detergent (Fig. S3, red arrow, Suppl. Information).
To further characterize the composition of these stable supramolecular associations, a second BN-PAGE with 0.02% of DDM in the cathode buffer was performed on the acrylamide lanes comprising separated digitonin- or DDM- solubilised complexes. All DDM-solubilized complexes separated as individual complexes in the 1D BN-PAGE are now found on a diagonal according to their molecular masses in the 2D gel (Fig. S4, Suppl. Information). In contrast, the digitonin-solubilized SCs are dissociated by the DDM present in the cathode buffer during the 2D BN-PAGE (Fig. 2a). As expected, individual complexes dissociated from SCs migrated below the diagonal at positions in the gel that correspond to the migration distances of the four native DDM-solubilized respiratory complexes I, III2, IV or V2 (shown on the left part of Fig. 2a). In the case of complex I, its identity was further confirmed by in-gel staining activity. The band between the monomeric complex I and dimeric complex V bands (Fig. 2a, purple arrowhead) comprises CI, CIII, and CIV and might thus correspond to the so-called “respirasome” with an I/III2/IV stoichiometry (2.2 MDa). The 4.2 MDa spot only comprises dimeric complex V and may thus corresponds to a tetrameric complex (V4) (Fig. 2ayellow arrowhead). In some cases, two faintly ATPase activity bands are also observed in the 1D gel below the V2 band (Figs. 2a and S4, upper lanes), these bands (marked as a and b) correspond to partial dissociation of the dimeric complex V that is also observed when the purified V2 complex is incubated in presence of DDM (Fig. S5, Suppl. Information). Finally, a co-migration involving CI and CV2 is visible, interestingly, CI and CV activities are present in this 5.2 MDa band but no evidence of CIII and CIV presence after the Coomassie blue staining was observed (data not shown), this opens the possibility that these two complexes form a larger supercomplex (Fig. 2a, green arrowhead).
Two-dimensional resolution of OXPHOS complexes and supercomplexes in E. gracilis mitochondria. a Upper panels: The OXPHOS complexes and supercomplexes from Euglena mitochondria were solubilized using digitonin (Dig) and separated by BN-PAGE followed by in-gel NADH-dehydrogenase and ATPase activities. Lower panel: NADH-dehydrogenase activity stain of two-dimensional gel from digitonin-extracted complexes and supercomplexes, the isolated spots show the complexes present in each supercomplex. III2/IV and III2/IV2 (red arrowheads), the respirasome (I/III2/IV, purple arrowhead), V4 oligomer (yellow arrowhead) and the Vx/I association (green arrowhead), isolated complexes were used as molecular mass markers (left lane). b The OXPHOS complexes and supercomplexes from Euglena mitochondria were solubilized using digitonin (upper lane) and separated by BN-PAGE. Lower panel: Two-dimensional SDS-tricine gel from digitonin-extracted complexes and supercomplexes. Representative subunits of each complex NDUFS3/NDUFA6/NDUFA9/NDUFA13/NDTB12/GapC3 for CI (purple arrows), QCR1/QCR2/QCR7 for CIII (red arrows), COX1/COX3/COXTB4/COXTB5/COX6b for CIV (green arrows) and ATPTB1/ATPTB2/Alpha-C/Beta for CV (blue arrows) are indicated. Molecular masses from the molecular mass marker are indicated on left side
To further confirm the composition of these SCs, the digitonin-extracted complexes were subjected to a 2D BN/SDS-PAGE. Based on the electrophoretic profile of subunits constitutive for each isolated complex (Fig. S6, Suppl. Information) (Perez et al. 2014; Yadav et al. 2017; Miranda-Astudillo et al. 2018b), several subunits representative of the individual complexes were identified in each of the SCs (e.g. NDTB12, NDUFA6, NDUFA9, NDUFA13 and GapC3 of CI, QCR1, QCR2 and QCR7 of CIII, COX1, COX3, COXTB4, COX6b and COXTB5 of CIV and ATPTB1, ATPTB2 Alpha-C and Beta of CV) (Fig. 2b). In the largest 5.2 MDa band, CI components are barely visible. This suggests that this band is dominated by a CV oligomer (V5 or V6). Relatively to complex III components, complex IV subunits are more abundant in the 1200 kDa band than in the 970 kDa band. This observation is in line with a greater complex III: complex IV stoichiometry in the 1200 kDa band of the two dimensional BN-PAGE/BN(+DDM)-PAGE (Fig. 2a, red arrowheads), suggesting the existence of two CIII/CIV SCs with two different stoichiometries: III2/IV or III2/IV2 for the 970 and 1200 kDa bands, respectively. Taken together, our results show that four isolated complexes and five larger SCs can be isolated from Euglena mitochondria. The proposed stoichiometry for each SC is also supported by the good correspondence between the determined and the expected molecular masses for all the SCs (linear regression coefficient R2 = 0.9971) (Fig. S7).
Purification of the Euglenoid respirasome
To further characterize the euglenoid respirasome, the GDN101-extracted I/III2/IV SC was purified by a two-step chromatographic procedure (see Material and methods section 2.3 for further details). The fractions containing associations between CI, CIII and CIV were enriched after the anion exchange chromatography (Fig. 3a, lower bracket), and a purified fraction containing the three complexes I/III2/IV association was obtained in the size exclusion chromatographic step (Fig. 3b). The redox differential absorption spectrum of the purified respirasome shows 528 nm and 558 nm peaks (Fig. 3c) which are typical of Euglena cytochrome c (c-558), as reported previously (Pettigrew et al. 1975; Mukai et al. 1989). To estimate the activity of this purified respirasome, in-vitro oxygen consumption was assayed upon addition of NADH as an electron donor. No oxygen consumption occurred at this point (Fig. 3dsegmented line) indicating either the interruption of the electron transport (e.g. loss of electron carriers by the detergent effect) or the damage/loss of function of the complexes during the purification procedure. Nevertheless, addition of external 2,3-dimethoxy-5-methyl-p-benzoquinone together with NADH led to a substantial oxygen consumption (Fig. 3dcontinuous line). Further addition of external horse cytochrome c did not enhance this in-vitro activity. This activity was also inhibited by rotenone (CI inhibitor) and antimycin A (CIII inhibitor) (Fig. 3e). In contrast, it was barely affected by the presence of myxothiazol (CIII inhibitor) (Fig. 3e).
Purification of the Euglenoid respirasome (I/III2/IV) by ion exchange/size exclusion chromatography and in-vitro oxygen consumption. Thirty milligrams of mitochondria were solubilized with glyco-diosgenin (GDN101) and loaded into an anion exchange column, then eluted with a 0–500 mM NaCl linear gradient. a BN- PAGE from the eluted 0.5 mL fractions, fractions containing III2/IV, III2/IV2 and I/III2/IV SCs (lower bracket) were concentrated and subjected to size exclusion column. b BN-PAGE from the pure respirasome (I/III2/IV) and the sample load onto the column (M), the identities of the SCs are indicated. c Differential redox absorption spectrum (520–580 nm) of purified respirasome was obtained as the sodium dithionite reduced spectrum minus the potassium ferricyanide oxidized spectrum. Alpha (558 nm) and beta (528 nm) peaks are signaled. d Oxygen consumption of the purified respirasome. The purified respirasome was incubated in the presence of NADH as electron donor (segmented line). External oxidized 2,3-Dimethoxy-5-methyl-p-benzoquinone was added (continuous line). The asterisk indicates the addition of the protein sample and the arrowhead indicates the addition of the complex I inhibitor rotenone. The lines were moved along the y axis for clarity. e Effect of external cytochrome c and inhibitory effect of antimycin A, myxothiazol and rotenone over the purified Euglena respirasome. The values represent the mean of three independent experiments and the bars represent the standard deviation
Discussion
Canonical association between complexes III & IV in E. gracilis
It has been recently shown that the OXPHOS complexes I, III2, IV and V2 from E. gracilis present atypical subunits which lead to characteristic structural features such as extra domains in complexes I, IV and V2, when comparing them with their homologs in classical model organisms (e.g. mammals and yeast) (Yadav et al. 2017; Miranda-Astudillo et al. 2018b). Digitonin is a widely-used detergent to study the supramolecular association in mitochondrial complexes from several species (Paumard et al. 2002; Bultema et al. 2009; Dudkina et al. 2011). This detergent is known to favour the native associations between membrane complexes from different organelle sources (Schägger 2002; Vonck and Schäfer 2009; Benson et al. 2015). When Euglena mitochondria were treated with digitonin or its synthetic substitute GDN101, four main additional supramolecular associations are observed (Fig. 1). Among them, three correspond to supercomplexes involving complexes III and IV. The association of Euglena complexes III and IV was already observed in native gels with low DDM concentration or digitonin extractions (Perez et al. 2014). Based on their estimated molecular mass of 2200, 1200 and 970 kDa, and on the relative abundance of each complex, these SCs probably correspond to I/III2/IV, III2/IV2 and III2/IV, respectively. Structures of both I/III2/IV, and III2/IV2 SCs have been already characterized in mammals, yeast and plants (Davies et al. 2018; Rathore et al. 2019). We reanalysed transmission electron microscopy (TEM) images obtained from an enriched fraction containing both complexes III and IV purified in presence of DDM (Miranda-Astudillo et al. 2018b). Further single particle analysis from this fraction revealed two additional classes of images that may correspond to an association between complexes III and IV (Fig. 4, upper panels). The atypical “helmet-like” domain of the Euglena cytochrome c oxidase (Miranda-Astudillo et al. 2018b) is however not visible in the two projections. This atypical extra domain exposed to the intermembrane side (p side) (Fig. 4, red arrowheads) was proposed to build a specific cavity for the endogenous cytochrome c (Miranda-Astudillo et al. 2018b). The III2/IV model built from the electronic density map recently obtained from mammalian respirasome (EMD: 9539 (Wu et al. 2016)) (Fig. 4, right panels) explains both EM projections (Fig. 4, lower panels). This suggests that the overall structural association between complex III and IV described for porcine (Wu et al. 2016), bovine (Davies et al. 2018) and yeast (Heinemeyer et al. 2007) mitochondria is conserved in E. gracilis, extending the previous proposed idea that the fundamental features of the supramolecular organisation (i.e., structure, composition, stoichiometry) of the respiratory complexes were conserved in lineages beyond classical mammals, fungi, and flowering plants models (Krause et al. 2004), supporting the idea of a common ancestry of these organelles (Margulis 1971).
2D Projection maps of III2/IV supercomplex from E. gracilis obtained by single particle averaging. Left panels: A fraction containing both complexes was obtained after a two-step chromatographic procedure in presence of β-dodecyl-n-maltoside and analyzed by TEM (a and b). Overlap of the coupled model (see material and methods point 2.6 for further details) built with chicken dimeric complex III (pdb: 4U3F (Hao et al. 2015)) and the monomeric bovine complex together with the cytochrome c (pdb: 5IY5 (Shimada et al. 2017)) over the TEM images was performed (c and d). The membrane region is indicated by the green arrowheads, the cytochrome c binding site is indicated with red arrowheads. Right panels (a-c): model showing the position of the III2/IV supercomplex inside the mammalian respirasome map (EMD: 9539 (Wu et al. 2016)). Orange: dimeric complex III, green monomeric complex IV, light grey: monomeric complex one. The membrane region is signaled (green arrowheads), topology of this supercomplex in the mitochondrial inner membrane is signaled (M: matrix, IM: intermembrane space). The scale bar is 10 nm
The purified respirasome lost the dedicated quinone pool but conserved the endogenous euglenoid cytochrome c
After both steps of anion exchange chromatography and size exclusion chromatography, a stable I/III2/IV respirasome complex was separated from the smaller SCs (III2/IV1 and III2/IV2). This is, to our knowledge, the first purification of a complete and functional respirasome from an organism beyond the classical model organisms (i.e., mammals, yeasts, green plants). The presence of the endogenous cytochrome c bound to the purified respirasome was evidenced (Fig. 3c). As a result, this respirasome was capable of transferring electrons from NADH to oxygen provided that only external quinones were added. This contrasts with the in-vitro reconstituted respirasome from the chlorophycean alga Polytomella sp. which requires further addition of both external electron carriers (quinone and cytochrome c) (Miranda-Astudillo et al. 2018a). Recently, it was also shown that oxygen consumption in presence of NADH of the purified respirasome from U. maydis is enhanced when external electron carries were added (Reyes-Galindo et al. 2019). Our observation thus suggests that association between endogenous cytochrome c and the respirasome in E. gracilis is more stable than in other species. Electron density corresponding to cytochrome c is observed in the III2/IV1 SC projections (Fig. 4red arrow heads). The presence of an identified helmet-like extra domain in isolated Euglena complex IV (Miranda-Astudillo et al. 2018b) might prevent the release of cytochrome c from the respirasome. Incidentally, the existence of such an unusual binding cavity for cytochrome c would be in line with previous works that showed that Euglena CIV is unable to used cytochrome c from other species (Collins et al. 1975; Brönstrup and Hachtel 1989).
Another major difference is the insensitivity of E. gracilis complex III to myxothiazol (Moreno-Sánchez et al. 2000; Perez et al. 2014). Our results (Fig. 3e) are in line with the inhibitors sensitivity previously described for fresh mitochondria (Mukai et al. 1989). It has been suggested earlier that the putative presence of a previously proposed additional CIII-like complex (Moreno-Sánchez et al. 2000) might explain this differential sensitivity. In this respect, we proposed that the presence of atypical QCR1/2 orthologs in E. gracilis (QCRTB1/2) along with the lack of the conserved QCR8 subunit, and the presence of an atypical QCR7 N-terminal extension (~ 100 residues) may affect the binding site of myxothiazol (Perez et al. 2014; Miranda-Astudillo et al. 2018b). Recently, Krnáčová et al. (2015), showed a myxothiazol- and antimycin- sensitive respiration (~50%) in E. gracilis but only one type of CIII has been identified in their study (Krnáčová et al. 2015) as in ours (Perez et al. 2014; Miranda-Astudillo et al. 2018b). These differences of sensitivity to myxothiazol and antimycin A can perhaps be explained by a difference in accessibility of the inhibitors to their binding site depending on the oligomeric state (monomer, dimer, supercomplex), which can be different depending on the culture conditions, the type of samples and the preparative methods (cells, mitochondria, isolated complexes), or their storage conditions (frozen, fresh).
The substrate channeling, specially the dedicated quinol pool, is one of the major advantages proposed for the respirasome formation. Nevertheless, recent spectroscopic and kinetic experiments performed in mammalian mitochondria refute this idea and points towards the existence of an universally accessible ubiquinone/ubiquinol pool that is not partitioned or channelled (Blaza et al. 2014; Fedor and Hirst 2018). These results together with the absence of known supercomplex-mediating factors in extensive structural data of isolated complexes (Davies et al. 2011, 2018; Dudkina et al. 2011) together with the ability to reconstitute functional SCs from isolated complexes (Bazán et al. 2013; Miranda-Astudillo et al. 2018a) stand up for a structural role of these SCs specially working in favour of CI stability (Acin-Perez and Enriquez 2014). Additionally, specific interactions between the complexes may protect against non-specific aggregation in the high protein concentration of the mitochondrial inner membrane (Blaza et al. 2014) and promote higher diffusion rates of the membrane embedded quinones (Kirchhoff 2014; Fedor and Hirst 2018). Putative respiratory strings, formed by an association of respirasomes have been observed by the study of the freeze-fractured and deep-etched inner mitochondrial membranes from Paramecium multimicronucleatum (Allen et al. 1989), and also have been proposed to be present in mammalian and plant mitochondria based on electron microscopy and native electrophoresis analysis (Wittig et al. 2006b; Bultema et al. 2009; Nubel et al. 2009; Wittig and Schägger 2009; Dudkina et al. 2010) where I2/III2/IV2 and I2/(III2)2/IV2 SCs should work as building blocks according to circular or linear models respectively (Bultema et al. 2009; Letts et al. 2016; Guo et al. 2018). Our results when digitonin or GDN101 are used for the solubilisation of membranes indicated that a large amount of CI, CIII and CIV are involved in SCs. This suggests the existence of respirasome strings as well in Euglena.
Tetrameric stable ATP synthase from E. gracilis
The dimeric mitochondrial ATP synthases of opisthokonts (i.e. mammals and fungal enzymes) easily dissociates into monomers in presence of DDM when subjected to BN-PAGE (van Lis et al. 2003, 2007; Wittig et al. 2006a). In contrast, Euglena DDM-extracted ATP synthase remains as a stable dimer and even a remnant of the tetrameric form (V4) is detectable (Fig. 1c, lane 5). Similarly, a highly stable DDM-solubilized tetrameric ATP synthase has already been observed in mitochondria of chlorophycean algae (Miranda-Astudillo et al. 2018a). In contrast, no tetrameric complex from opisthokonts (i.e metazoan and or fungi) has been observed in presence of DDM. Oligomeric forms have been however observed when digitonin is used to extract the complex (e.g in bovine (Strauss et al. 2008; Wittig and Schägger 2009), porcine (Gu et al. 2019) and yeast (Thomas et al. 2008; Habersetzer et al. 2013)). The cryo-EM structure of the mammalian tetrameric enzyme was recently obtained (Gu et al. 2019). In an H-shape, two dimers are bound mainly by their membrane sector as previously proposed based on crosslinking experiments and 2D microscopy images from the isolated tetramers (Thomas et al. 2008; Habersetzer et al. 2013). Additionally, two dimers of the inhibitory subunit IF1 link the F1 sectors from the adjacent dimers (Gu et al. 2019). This ATPase oligomerization leads to ATPase ribbons formation (Strauss et al. 2008; Blum et al. 2019).
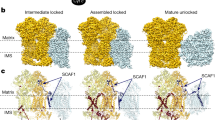
To get some insight into the structure of the tetrameric form of Euglena ATP synthase, the recently obtained cryo-EM structure of the Euglena ATP synthase dimer (Mühleip et al. 2019) were fitted inside the 27.5 Å 3D map (EMD: 3559) from Euglena ATP synthase ribbon determined by electron cryotomography and subtomogram averaging from intact mitochondrial membranes (Mühleip et al. 2017). This comparison let us propose a putative structure of the Euglena tetrameric complex V where the dimer-dimer interaction is present mainly at the membrane level (Fig. 5, green arrowheads), and where the external peripheral stalks together with one of the p18 subunits face outside of the ribbon (Fig. 5, red arrowheads). The possible role of the euglenoid subunit p18 (Zíková et al. 2009; Perez et al. 2014; Yadav et al. 2017) remains obscure, nevertheless, its structural role could be related to its exposed side in the ATP synthase ribbon. The possible dimer-dimer contact zone should be also located mainly among the large membrane-spanning region described for Euglena enzyme (Mühleip et al. 2017; Yadav et al. 2017), especially because in the Euglena enzyme the IF1 peptide is binding the adjacent monomers inside one dimer (Mühleip et al. 2019) and not between adjacent dimers as described for the mammalian tetramer (Gu et al. 2019).
Putative tetrameric structure of the E. gracilis ATP synthase. Putative tetrameric structure of the E. gracilis ATP synthase based in the electron cryotomography images from intact mitochondria membranes (Mühleip et al. 2017). The externally located peripheral stalks are shown in purple, the inter membrane space density below the c-ring is shown in orange, the F1/central rotor sector is shown in cyan, the membrane region is signalled (green arrowheads) and the position of the euglonoid specific subunit p18 is indicated (red arrowheads), the structure of the Euglena ATP synthase dimer (PDB: 6TDU (Mühleip et al. 2019)) is fitted inside the electron density for one dimer. The scale bar is 10 nm
The angle formed between the CV membrane sector (FO) of the two monomers in Euglena dimer is much lower (40°) than in the dimers of opisthokonts (70°), so the effect of these dimers on cristae shape might be different. In this respect, lamellar cristae are found in the supergroup of opisthokonts, while tubular-shaped cristae typify the unicellular supergroup SAR, containing stramenopiles, alveolates, and rhizarians (Mühleip et al. 2016, 2021). Euglenids, members of the protistan supergroup Excavata, generally have discoidal cristae, which exhibit a paddle-like morphology (Kaurov et al. 2018). Electron cryotomography analyses on mitochondria from plants, mammals and yeasts showed that the dimeric ATP synthase rows locate along the cristae curvature, bending the membrane while the rest of the complexes are irregularly distributed confined to flat membrane regions (Davies et al. 2011, 2018). Some CI particles are however located beside the CV row in tomographic slices of isolated cristae membranes from bovine mitochondria (Davies et al. 2011). In Euglena, the comigration of a fraction of CI and oligomeric CV in BN-PAGE experiments, could reflect a similar partitioning in vivo. Similar comigration of CV together with respiratory complexes has been observed recently in pea mitochondria treated with digitonin (Ukolova et al. 2020).
Conserved supercomplexes formation in E. gracilis
The OXPHOS complexes from E. gracilis contain atypical subunits which lead to extra structural domains (Yadav et al. 2017; Miranda-Astudillo et al. 2018b). Nevertheless, they form classical III2/IV1–2 associations and the respirasome (I/III2/IV). The mammalian respirasome present four major structural pivots, two of them related with the CI/CIII2 association, one with CIII2/IV interface and the last with CI membrane extrinsic arm (Letts and Sazanov 2017), leaving a major structural role in the central position of CIII2 whose in situ arrangement is conserved between the opisthokonts and plant mitochondria (Davies et al. 2018). In this respect, despite that Euglena CIII presents four atypical subunits, the overall structure of the dimeric complex is conserved (Miranda-Astudillo et al. 2018b), and a canonical CIII2/IV arrangement is observed in DDM-extracted III2/IV supercomplex (Fig. 4). Additionally, Euglena III2/IV SC can bind a second monomeric CIV forming a III2/IV2 SC that may form a larger respirasome (I/III2/IV2) observed at 2450 kDa band (Fig. 2a, upper bands). This latter species is however less stable in our experimental conditions than the purified 2200 kDa respirasome (I/III2/IV) (Fig. 3b). Similarly, in situ observation inside the inner membrane of mammalian mitochondria showed that I/III2/IV SC is more abundant than I/III2/IV2 SC (Davies et al. 2018). Finally, the data obtained in the present work and in our previous works (Perez et al. 2014; Yadav et al. 2017; Miranda-Astudillo et al. 2018b) let us propose an assembly pathway for Euglena respirasome SC from OXPHOS complexes (Fig. 6) quite similar to the one described in other linages (Lobo-Jarne and Ugalde 2018), despite atypical subunit composition and additional structural domains of the oxidative phosphorylation complexes in Euglena gracilis.
Assembly pathway of mitochondrial respirasome in E. gracilis. Schematic representation of the associations of complexes into supercomplexes. The species observed by BN-PAGE (I, III2, IV, V2, III2/IV, III2/IV2, I/III2/IV, I/III2/IV2) and TEM analysis (I, III2, IV, and III2/IV) from Euglena mitochondria ((Yadav et al. 2017; Miranda-Astudillo et al. 2018b), this work) are shown in green and dark blue boxes, respectively
Change history
07 May 2021
A Correction to this paper has been published: https://doi.org/10.1007/s10863-021-09890-8
References
Acin-Perez R, Enriquez JA (2014) The function of the respiratory supercomplexes: the plasticity model. Biochim Biophys Acta Bioenerg 1837:444–450. https://doi.org/10.1016/j.bbabio.2013.12.009
Allegretti M, Klusch N, Mills DJ, Vonck J, Kühlbrandt W, Davies KM (2015) Horizontal membrane-intrinsic α-helices in the stator a-subunit of an F-type ATP synthase. Nature 521:237–240. https://doi.org/10.1038/nature14185
Allen RD, Schroeder CC, Fok AK (1989) An investigation of mitochondrial inner membranes by rapid-freeze deep-etch techniques. J Cell Biol 108:2233–2240. https://doi.org/10.1083/jcb.108.6.2233
Atteia A, Van Lis R, Mendoza-Hernández G et al (2003) Bifunctional aldehyde/alcohol dehydrogenase (ADHE) in chlorophyte algal mitochondria. Plant Mol Biol 53:175–188. https://doi.org/10.1023/B:PLAN.0000009274.19340.36
Balabaskaran Nina P, Dudkina NV, Kane LA, van Eyk JE, Boekema EJ, Mather MW, Vaidya AB (2010) Highly divergent mitochondrial ATP synthase complexes in Tetrahymena thermophila. PLoS Biol 8:3–6. https://doi.org/10.1371/journal.pbio.1000418
Bazán S, Mileykovskaya E, Mallampalli VKPS, Heacock P, Sparagna GC, Dowhan W (2013) Cardiolipin-dependent reconstitution of respiratory supercomplexes from purified Saccharomyces cerevisiae complexes III and IV. J Biol Chem 288:401–411. https://doi.org/10.1074/jbc.M112.425876
Benichou P, Calvayrac R, Claisse M (1988) Induction by antimycin a of cyanide-resistant respiration in heterotrophic Euglena gracilis: effects on growth, respiration and protein biosynthesis. Planta 175:23–32
Benson SL, Maheswaran P, Ware MA, Hunter CN, Horton P, Jansson S, Ruban AV, Johnson MP (2015) An intact light harvesting complex I antenna system is required for complete state transitions in Arabidopsis. Nat Plants 1:15176. https://doi.org/10.1038/nplants.2015.176
Blaza JN, Serreli R, Jones AJY, Mohammed K, Hirst J (2014) Kinetic evidence against partitioning of the ubiquinone pool and the catalytic relevance of respiratory-chain supercomplexes. Proc Natl Acad Sci 111:15735–15740. https://doi.org/10.1073/pnas.1413855111
Blum TB, Hahn A, Meier T, Davies KM, Kühlbrandt W (2019) Dimers of mitochondrial ATP synthase induce membrane curvature and self-assemble into rows. Proc Natl Acad Sci 116:4250–4255. https://doi.org/10.1073/pnas.1816556116
Brönstrup U, Hachtel W (1989) Cytochrome c oxidase of Euglena gracilis: purification, characterization, and identification of mitochondrially synthesized subunits. J Bioenerg Biomembr 21:359–373. https://doi.org/10.1007/BF00762727
Bultema JB, Braun HP, Boekema EJ, Kouril R (2009) Megacomplex organization of the oxidative phosphorylation system by structural analysis of respiratory supercomplexes from potato. Biochim Biophys Acta Bioenerg 1787:60–67. https://doi.org/10.1016/j.bbabio.2008.10.010
Burki F (2014) The eukaryotic tree of life from a global phylogenomic perspective. Cold Spring Harb Perspect Biol 6:1–18. https://doi.org/10.1101/cshperspect.a016147
Cardol P, Figueroa F, Remacle C et al (2009) Chapter 13 – oxidative phosphorylation: building blocks and related components. The Chlamydomonas Sourcebook, In, pp 469–502
Chaban Y, Boekema EJ, Dudkina NV (2014) Structures of mitochondrial oxidative phosphorylation supercomplexes and mechanisms for their stabilisation. Biochim Biophys Acta Bioenerg 1837:418–426. https://doi.org/10.1016/j.bbabio.2013.10.004
Colina-Tenorio L, Dautant A, Miranda-Astudillo H, Giraud MF, González-Halphen D (2018) The peripheral stalk of rotary ATPases. Front Physiol 9:1–19. https://doi.org/10.3389/fphys.2018.01243
Colina-Tenorio L, Miranda-Astudillo H, Dautant A, Vázquez-Acevedo M, Giraud MF, González-Halphen D (2019) Subunit Asa3 ensures the attachment of the peripheral stalk to the membrane sector of the dimeric ATP synthase of Polytomella sp. Biochem Biophys Res Commun 509:341–347. https://doi.org/10.1016/j.bbrc.2018.12.142
Collins N, Brown RH, Merrett MJ (1975) Oxidative phosphorylation during glycollate metabolism in mitochondria from phototrophic Euglena gracilis. Biochem J 150:373–377
Cross RL, Müller V (2004) The evolution of A-, F-, and V-type ATP synthases and ATPases: reversals in function and changes in the H+/ATP coupling ratio. FEBS Lett 576:1–4. https://doi.org/10.1016/j.febslet.2004.08.065
Cross RL, Taiz L (1990) Gene duplication as a means for altering H+/ATP ratios during the evolution of Fo F1 ATPases and synthases. FEBS Lett 259:227–229. https://doi.org/10.1016/0014-5793(90)80014-A
Davies KM, Strauss M, Daum B, Kief JH, Osiewacz HD, Rycovska A, Zickermann V, Kuhlbrandt W (2011) Macromolecular organization of ATP synthase and complex I in whole mitochondria. Proc Natl Acad Sci 108:14121–14126. https://doi.org/10.1073/pnas.1103621108
Davies KM, Blum TB, Kühlbrandt W (2018) Conserved in situ arrangement of complex I and III 2 in mitochondrial respiratory chain supercomplexes of mammals, yeast, and plants. Proc Natl Acad Sci 115:3024–3029. https://doi.org/10.1073/pnas.1720702115
Duarte M, Tomás AM (2014) The mitochondrial complex I of trypanosomatids - an overview of current knowledge. J Bioenerg Biomembr 46:299–311. https://doi.org/10.1007/s10863-014-9556-x
Dudkina NV, Kouril R, Peters K et al (2010) Structure and function of mitochondrial supercomplexes. Biochim Biophys Acta Bioenerg 1797:664–670. https://doi.org/10.1016/j.bbabio.2009.12.013
Dudkina NV, Kudryashev M, Stahlberg H, Boekema EJ (2011) Interaction of complexes I, III, and IV within the bovine respirasome by single particle cryoelectron tomography. Proc Natl Acad Sci U S A 108:15196–15200. https://doi.org/10.1073/pnas.1107819108
Fedor JG, Hirst J (2018) Mitochondrial Supercomplexes do not enhance catalysis by Quinone channeling. Cell Metab 28:1–7. https://doi.org/10.1016/j.cmet.2018.05.024
Genova ML, Lenaz G (2014) Functional role of mitochondrial respiratory supercomplexes. Biochim Biophys Acta 1837:427–443. https://doi.org/10.1016/j.bbabio.2013.11.002
Gibbs SP (1981) The chloroplasts of some algal groups may have evolved from endosymbiotic eukaryotic algae. Ann N Y Acad Sci 81:193–208. https://doi.org/10.1111/j.1749-6632.1981.tb54365.x
Gu J, Zhang L, Zong S et al (2019) Cryo-EM structure of the mammalian ATP synthase tetramer bound with inhibitory protein IF1. Science (80- ) 364:1068–1075. https://doi.org/10.1126/science.aaw4852
Guo R, Gu J, Zong S, Wu M, Yang M (2018) Structure and mechanism of mitochondrial electron transport chain. Biom J 41:9–20. https://doi.org/10.1016/j.bj.2017.12.001
Habersetzer J, Ziani W, Larrieu I, Stines-Chaumeil C, Giraud MF, Brèthes D, Dautant A, Paumard P (2013) ATP synthase oligomerization: from the enzyme models to the mitochondrial morphology. Int J Biochem Cell Biol 45:99–105. https://doi.org/10.1016/j.biocel.2012.05.017
Hackenbrock CR, Chazotte B, Gupte SS (1986) The random collision model and a critical assessment of diffusion and collision in mitochondrial electron transport. J Bioenerg Biomembr 18:331–368. https://doi.org/10.1007/BF00743010
Hao G-F, Yang S-G, Huang W, Wang L, Shen YQ, Tu WL, Li H, Huang LS, Wu JW, Berry EA, Yang GF (2015) Rational Design of Highly Potent and Slow-Binding Cytochrome bc1 inhibitor as fungicide by computational substitution optimization. Sci Rep 5:13471. https://doi.org/10.1038/srep13471
Heinemeyer J, Braun HP, Boekema EJ, Kouřil R (2007) A structural model of the cytochrome c reductase/oxidase supercomplex from yeast mitochondria. J Biol Chem 282:12240–12248. https://doi.org/10.1074/jbc.M610545200
Kaurov I, Vancová M, Schimanski B et al (2018) The diverged trypanosome MICOS complex as a hub for mitochondrial cristae shaping and protein import. Curr biol 28:3393-3407.e5. https://doi.org/10.1016/j.cub.2018.09.008
Keilin D, Hartree EF (1947) Activity of the cytochrome system in heart muscle preparations. Biochem J 41:500–502. https://doi.org/10.1042/bj0410500
Kirchhoff H (2014) Diffusion of molecules and macromolecules in thylakoid membranes. Biochim Biophys Acta Bioenerg 1837:495–502. https://doi.org/10.1016/j.bbabio.2013.11.003
Krnáčová K, Rýdlová I, Vinarčíková M, et al (2015) Characterization of oxidative phosphorylation enzymes in Euglena gracilis and its white mutant strain WgmZOflL. FEBS Lett 589:687–694. https://doi.org/10.1016/j.febslet.2015.01.035
Krause F, Reifschneider NH, Vocke D, Seelert H, Rexroth S, Dencher NA (2004) “Respirasome”-like supercomplexes in green leaf mitochondria of spinach. J Biol Chem 279:48369–48375. https://doi.org/10.1074/jbc.M406085200
Kühlbrandt W (2019) Structure and mechanisms of F-type ATP synthases. Annu Rev Biochem 88:515–549. https://doi.org/10.1146/annurev-biochem-013118-110903
Lenaz G, Tioli G, Falasca AI, Genova ML (2016) Complex i function in mitochondrial supercomplexes. Biochim Biophys Acta Bioenerg 1857:991–1000. https://doi.org/10.1016/j.bbabio.2016.01.013
Letts JA, Sazanov LA (2017) Clarifying the supercomplex: the higher-order organization of the mitochondrial electron transport chain. Nat Struct Mol Biol 24:800–808. https://doi.org/10.1038/nsmb.3460
Letts JA, Fiedorczuk K, Sazanov LA (2016) The architecture of respiratory supercomplexes. Nature 537:644–648. https://doi.org/10.1038/nature19774
Lobo-Jarne T, Ugalde C (2017) Respiratory chain supercomplexes: structures, function and biogenesis. Semin Cell Dev Biol 76:179–190. https://doi.org/10.1016/j.semcdb.2017.07.021
Lobo-Jarne T, Ugalde C (2018) Respiratory chain supercomplexes: structures, function and biogenesis. Semin Cell Dev Biol 76:179–190. https://doi.org/10.1016/j.semcdb.2017.07.021
Margulis L (1971) The origin of plant and animal cells. Am Sci 59:230–235
Milenkovic D, Blaza JN, Larsson NG, Hirst J (2017) The enigma of the respiratory chain Supercomplex. Cell Metab 25:765–776. https://doi.org/10.1016/j.cmet.2017.03.009
Miranda-Astudillo H, Colina-Tenorio L, Jiménez-Suárez A, Vázquez-Acevedo M, Salin B, Giraud MF, Remacle C, Cardol P, González-Halphen D (2018a) Oxidative phosphorylation supercomplexes and respirasome reconstitution of the colorless alga Polytomella sp. Biochim Biophys Acta Bioenerg 1859:434–444. https://doi.org/10.1016/j.bbabio.2018.03.004
Miranda-Astudillo H, Yadav KNS, Colina-Tenorio L et al (2018b) The atypical subunit composition of respiratory complexes I and IV is associated with original extra structural domains in Euglena gracilis. Sci Rep 8:9698. https://doi.org/10.1038/s41598-018-28039-z
Montgomery MG, Leslie AGW, Zíková A, Walker JE (2018) ATP synthase from Trypanosoma brucei has an elaborated canonical F 1 -domain and conventional catalytic sites. Proc Natl Acad Sci 115:2102–2107. https://doi.org/10.1073/pnas.1720940115
Morales J, Mogi T, Mineki S, Takashima E, Mineki R, Hirawake H, Sakamoto K, Ōmura S, Kita K (2009) Novel mitochondrial complex II isolated from trypanosoma cruzi is composed of 12 peptides including a heterodimeric ip subunit. J Biol Chem 284:7255–7263. https://doi.org/10.1074/jbc.M806623200
Moreno-Sánchez R, Covián R, Jasso-Chávez R, Rodrı́guez-Enrı́quez S, Pacheco-Moisés F́, Torres-Márquez ME (2000) Oxidative phosphorylation supported by an alternative respiratory pathway in mitochondria from Euglena. Biochim Biophys Acta Bioenerg 1457:200–210. https://doi.org/10.1016/S0005-2728(00)00102-X
Muench SP, Trinick J, Harrison MA (2011) Structural divergence of the rotary ATPases
Mühleip AW, Joos F, Wigge C, Frangakis AS, Kühlbrandt W, Davies KM (2016) Helical arrays of U-shaped ATP synthase dimers form tubular cristae in ciliate mitochondria. Proc Natl Acad Sci U S A 113:8442–8447. https://doi.org/10.1073/pnas.1525430113
Mühleip AW, Dewar EC, Schnaufer A et al (2017) In-situ structure of trypanosomal ATP synthase dimer reveals unique arrangement of catalytic subunits. PNAS. 114:992–997. https://doi.org/10.1073/pnas.1612386114
Mühleip A, McComas SE, Amunts A (2019) Structure of a mitochondrial ATP synthase with bound native cardiolipin. Elife 8:1–23. https://doi.org/10.7554/eLife.51179
Mühleip A, Kock Flygaard R, Ovciarikova J, Lacombe A, Fernandes P, Sheiner L, Amunts A (2021) ATP synthase hexamer assemblies shape cristae of toxoplasma mitochondria. Nat Commun 12:120. https://doi.org/10.1038/s41467-020-20381-z
Mukai K, Yoshida M, Toyosaki H et al (1989) An atypical heme-binding structure of cytochrome c1 of Euglena gracilis mitochondrial complex III. Eur J Biochem 178:649–656. https://doi.org/10.1111/j.1432-1033.1989.tb14494.x
Nubel E, Wittig I, Kerscher S et al (2009) Two-dimensional native electrophoretic analysis of respiratory supercomplexes from Yarrowia lipolytica. Proteomics 9:2408–2418. https://doi.org/10.1002/pmic.200800632
Paumard P, Vaillier J, Coulary B, Schaeffer J, Soubannier V, Mueller DM, Brèthes D, di Rago JP, Velours J (2002) The ATP synthase is involved in generating mitochondrial cristae morphology. EMBO J 21:221–230. https://doi.org/10.1093/emboj/21.3.221
Perez E, Lapaille M, Degand H, Cilibrasi L, Villavicencio-Queijeiro A, Morsomme P, González-Halphen D, Field MC, Remacle C, Baurain D, Cardol P (2014) The mitochondrial respiratory chain of the secondary green alga Euglena gracilis shares many additional subunits with parasitic Trypanosomatidae. Mitochondrion 19:338–349. https://doi.org/10.1016/j.mito.2014.02.001
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF chimera - a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Pettigrew GW, Leaver JL, Meyer TE, Ryle a P (1975) Purification, properties and amino acid sequence of atypical cytochrome c from two protozoa, Euglena gracilis and Crithidia oncopelti. Biochem J 147:291–302
Rathore S, Berndtsson J, Marin-Buera L, Conrad J, Carroni M, Brzezinski P, Ott M (2019) Cryo-EM structure of the yeast respiratory supercomplex. Nat Struct Mol Biol 26:50–57. https://doi.org/10.1038/s41594-018-0169-7
Reyes-Galindo M, Suarez R, Esparza-Perusquía M, de Lira-Sánchez J, Pardo JP, Martínez F, Flores-Herrera O (2019) Mitochondrial respirasome works as a single unit and the cross-talk between complexes I, III2 and IV stimulates NADH dehydrogenase activity. Biochim Biophys Acta Bioenerg 1860:618–627. https://doi.org/10.1016/j.bbabio.2019.06.017
Salunke R, Mourier T, Banerjee M, Pain A, Shanmugam D (2018) Highly diverged novel subunit composition of apicomplexan F-type ATP synthase identified from toxoplasma gondii. PLoS Biol 16:e2006128. https://doi.org/10.1371/journal.pbio.2006128
Schägger H (1994a) Native gel electrophoresis. In: Schägger H (ed) von Jagow G. Academic Press, A Practical Guide to Membrane Protein Purification, pp 81–104
Schägger H (1994b) Denaturing electrophoretic techniques. In: Schägger H (ed) von Jagow G. Academic Press, A Practical Guide to Membrane Protein Purification, pp 59–79
Schägger H (2002) Respiratory chain supercomplexes of mitochondria and bacteria. Biochim Biophys Acta Bioenerg 1555:154–159. https://doi.org/10.1016/S0005-2728(02)00271-2
Schägger H, Pfeiffer K (2000) Supercomplexes in the respiratory chains of yeast and mammalian mitochondria. EMBO J 19:1777–1783. https://doi.org/10.1093/emboj/19.8.1777
Schägger H, Pfeiffer K (2001) The ratio of oxidative phosphorylation complexes I-V in bovine heart mitochondria and the composition of respiratory chain Supercomplexes. J Biol Chem 276:37861–37867. https://doi.org/10.1074/jbc.M106474200
Sharpless TK, Butow RA (1970a) An inducible alternate terminal oxidase in Euglena gracilis mitochondria. J Biol Chem 245:58–70
Sharpless TK, Butow RA (1970b) Phosphorylation sites, cytochrome complement, and alternate pathways of coupled electron transport in Euglena gracilis mitochondria. J Biol Chem 245:50–57
Shimada S, Shinzawa-Itoh K, Baba J, et al (2017) Complex structure of cytochrome c –cytochrome c oxidase reveals a novel protein–protein interaction mode. EMBO J 36:291–300. https://doi.org/10.15252/embj.201695021
Singer SJ, Nicolson GL (1972) The fluid mosaic model of the structure cell membranes. Science (80- ) 175:720–731
Slater EC (2003) Keilin, cytochrome, and the respiratory chain. J Biol Chem 278:16455–16461. https://doi.org/10.1074/jbc.X200011200
Speijer D, Breek CKD, Muijsers AO, Hartog AF, Berden JA, Albracht SPJ, Samyn B, van Beeumen J, Benne R (1997) Characterization of the respiratory chain from cultured Crithidia fasciculata. Mol Biochem Parasitol 85:171–186. https://doi.org/10.1016/S0166-6851(96)02823-X
Strauss M, Hofhaus G, Schröder RR, Kühlbrandt W (2008) Dimer ribbons of ATP synthase shape the inner mitochondrial membrane. EMBO J 27:1154–1160. https://doi.org/10.1038/emboj.2008.35
Thomas D, Bron P, Weimann T, Dautant A, Giraud MF, Paumard P, Salin B, Cavalier A, Velours J, Brèthes D (2008) Supramolecular organization of the yeast F1Fo-ATP synthase. Biol Cell 100:591–601. https://doi.org/10.1042/BC20080022
Turmel M, Gagnon MC, O’Kelly CJ et al (2009) The chloroplast genomes of the green algae Pyramimonas, Monomastix, and Pycnococcus shed new light on the evolutionary history of prasinophytes and the origin of the secondary chloroplasts of euglenids. Mol Biol Evol 26:631–648. https://doi.org/10.1093/molbev/msn285
Ukolova I, Kondakova M, Kondratov I et al (2020) New insights into the organisation of the oxidative phosphorylation system in the example of pea shoot mitochondria. Biochim Biophys Acta Bioenerg 1861:148264. https://doi.org/10.1016/j.bbabio.2020.148264
van Lis R, Atteia A, Mendoza-hernández G, González-halphen D (2003) Identification of novel mitochondrial protein components of Chlamydomonas reinhardtii . A proteomic approach 1. Plant Physiol 132:318–330. https://doi.org/10.1104/pp.102.018325.proteins
van Lis R, González-Halphen D, Atteia A (2005) Divergence of the mitochondrial electron transport chains from the green alga Chlamydomonas reinhardtii and its colorless close relative Polytomella sp. Biochim Biophys Acta Bioenerg 1708:23–34. https://doi.org/10.1016/j.bbabio.2004.12.010
van Lis R, Mendoza-Hernández G, Groth G, Atteia A (2007) New insights into the unique structure of the F0F1-ATP synthase from the chlamydomonad algae Polytomella sp. and Chlamydomonas reinhardtii. Plant Physiol 144:1190–1199. https://doi.org/10.1104/pp.106.094060
Verner Z, Basu S, Benz C et al (2015) Malleable mitochondrion of Trypanosoma brucei. Int Rev Cell Mol Biol 315:73–151. https://doi.org/10.1016/bs.ircmb.2014.11.001
Vonck J, Schäfer E (2009) Supramolecular organization of protein complexes in the mitochondrial inner membrane. Biochim Biophys Acta 1793:117–124. https://doi.org/10.1016/j.bbamcr.2008.05.019
Winge DR (2012) Sealing the mitochondrial Respirasome. Mol Cell Biol 32:2647–2652. https://doi.org/10.1128/MCB.00573-12
Wittig I, Schägger H (2005) Advantages and limitations of clear-native PAGE. Proteomics 5:4338–4346. https://doi.org/10.1002/pmic.200500081
Wittig I, Schägger H (2008) Features and applications of blue-native and clear-native electrophoresis. Proteomics 8:3974–3990. https://doi.org/10.1002/pmic.200800017
Wittig I, Schägger H (2009) Supramolecular organization of ATP synthase and respiratory chain in mitochondrial membranes. Biochim Biophys Acta 1787:672–680. https://doi.org/10.1016/j.bbabio.2008.12.016
Wittig I, Braun H-P, Schägger H (2006a) Blue native PAGE. Nat Protoc 1:418–428. https://doi.org/10.1038/nprot.2006.62
Wittig I, Carrozzo R, Santorelli FM, Schägger H (2006b) Supercomplexes and subcomplexes of mitochondrial oxidative phosphorylation. Biochim Biophys Acta Bioenerg 1757:1066–1072. https://doi.org/10.1016/j.bbabio.2006.05.006
Wu M, Gu J, Guo R et al (2016) Structure of mammalian respiratory Supercomplex I1III2IV1. Cell 167:1598–1609.e10. https://doi.org/10.1016/j.cell.2016.11.012
Yadav KNS, Miranda-Astudillo HV, Colina-Tenorio L, Bouillenne F, Degand H, Morsomme P, González-Halphen D, Boekema EJ, Cardol P (2017) Atypical composition and structure of the mitochondrial dimeric ATP synthase from Euglena gracilis. Biochim Biophys Acta Bioenerg 1858:267–275. https://doi.org/10.1016/j.bbabio.2017.01.007
Zakrys B, Rafal M, Karnkowska A (2017) Evolutionary Origin of Euglena. Adv Exp Med Biol 979:3-17. https://doi.org/10.1007/978-3-319-54910-1_1
Zíková A, Schnaufer A, Dalley RA, Panigrahi AK, Stuart KD (2009) The F0F1-ATP synthase complex contains novel subunits and is essential for procyclic Trypanosoma brucei. PLoS Pathog 5:e1000436. https://doi.org/10.1371/journal.ppat.1000436
Funding
P.C. acknowledges financial support from the Belgian Fonds de la Recherche Scientifique F.R.S.-F.N.R.S. (PDR T.0032) and European Research Council (ERC, H2020-EU BEAL project 682580).
Author information
Authors and Affiliations
Contributions
H.V.M.A. and P.C. conceived the research, H.V.M.A. and K.N.S.Y. performed the experiments, H.V.M.A., K.N.S.Y., E.J.B., and P.C. analysed the data, H.V.M.A. and P.C. wrote the manuscript, all authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Figure S1
Estimated molecular masses for the Euglena gracilis supercomplexes. The logarithms of the molecular masses of previously characterized mitochondrial complexes from the colorless algae Polytomella sp. (◆) (V6, V4, V2, I, III2; see (Atteia et al. 2003; van Lis et al. 2003, 2005, 2007; Cardol et al. 2009; Miranda-Astudillo et al. 2018a) for details) were plotted against their migration distance in BN-PAGE. Then, the migration distances of the Euglena digitonin-extracted supercomplexes (○) were interpolated and their corresponding molecular masses inferred. The proposed stoichiometry and the molecular mass of each complex and supercomplex are indicated. (PNG 757 kb)
Figure S2
ATPase oligomers and respiratory supercomplexes in Euglena gracilis extracted with non-ionic detergents. Isolated mitochondria were solubilized with digitonin (Dig) and its synthetic drop-in substitute glyco-diosgenin GDN101 (GDN) at 8.0 g/g protein. Final concentrations of detergent were 6.4%. After removing the insoluble material, each sample was resolved by BN-PAGE in a 3–10% polyacrylamide gradient gel. The determined molecular mass of each isolated complex or supercomplexes are indicated. Nomenclature used: I, III and IV for the corresponding mitochondrial complexes, V2 and V4 for the dimeric and tetrameric ATP synthase respectively. Supercomplexes were: III2/IV, III2/IV2, the so-called “Respirasome” association I/III2/IV and the putative Vx/I association, their stoichiometries are indicated as subindexes. (PNG 1146 kb)
Figure S3
Stability of the Euglena gracilis supercomplexes at increasing concentrations of digitonin. Isolated mitochondria were solubilized with increasing concentrations of digitonin. Left panel: in-gel NADH-dehydrogenase activity; the BN-gel was incubated in the presence of NADH and Nitro blue tetrazolium chloride (NBT). Right panel: BN-gel and detection of in-gel ATPase activity. The gel was incubated with ATP, MgSO4 and Pb(NO3)2. Numbers above represent the g of detergent/g of protein relation. (PNG 1880 kb)
Figure S4
Two-dimensional resolution of OXPHOS complexes in Euglena gracilis mitochondria. Upper lanes: The OXPHOS complexes from Euglena mitochondria were solubilized using n-dodecyl-β-D-maltoside (DDM) and separated by BN-PAGE followed by in-gel NADH-dehydrogenase and ATPase activities. Lower panel: NADH-dehydrogenase activity stain two-dimensional gels from DDM-extracted complexes, the isolated spots in the diagonal show the complexes separated by their molecular masses in each dimension. Isolated complexes were used as molecular mass markers (left lane). (PNG 1702 kb)
Figure S5
ATP synthase dissociation in Euglena gracilis. Purified dimeric ATP synthase was incubated in presence of n-dodecyl-β-D-maltoside (DDM) at room temperature for 10 min. Four species are observed which may correspond to dimeric enzyme V2, dimer without one F1 sector (band a), monomeric enzyme (band b), and the F1 sector. This assignment was done based on size. (PNG 1164 kb)
Figure S6
Two-dimensional resolution of OXPHOS complexes in E. gracilis mitochondria. The OXPHOS complexes from Euglena mitochondria were solubilized using n-dodecyl-β-D-maltoside (DDM) (upper lane) and separated by BN-PAGE. Lower panel: Two-dimensional SDS-tricine gel from DDM-extracted complexes. Identified subunits by tandem mass spectroscopy of each complex are indicated (Yadav et al. 2017; Miranda-Astudillo et al. 2018b). Molecular masses from the molecular mass marker are indicated on left side. (PNG 2180 kb)
Figure S7
Linear regression of calculated versus determined molecular masses in Euglena gracilis supercomplexes. (PNG 568 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Miranda-Astudillo, H.V., Yadav, K.N.S., Boekema, E.J. et al. Supramolecular associations between atypical oxidative phosphorylation complexes of Euglena gracilis. J Bioenerg Biomembr 53, 351–363 (2021). https://doi.org/10.1007/s10863-021-09882-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10863-021-09882-8