Abstract
Reducing sugars can spontaneously react with free amines in protein side chains leading to posttranslational modifications (PTMs) called glycation. In contrast to glycosylation, glycation is a non-enzymatic modification with consequences on the overall charge, solubility, aggregation susceptibility and functionality of a protein. Glycation is a critical quality attribute of therapeutic monoclonal antibodies. In addition to glucose, also disaccharides like maltose can form glycation products. We present here a detailed NMR analysis of the Amadori product formed between proteins and maltose. For better comparison, data collection was done under denaturing conditions using 7 M urea-d4 in D2O. The here presented correlation patterns serve as a signature and can be used to identify maltose-based glycation in any protein that can be denatured. In addition to the model protein BSA, which can be readily glycated, we present data of the biotherapeutic abatacept containing maltose in its formulation buffer. With this contribution, we demonstrate that NMR spectroscopy is an independent method for detecting maltose-based glycation, that is suited for cross-validation with other methods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
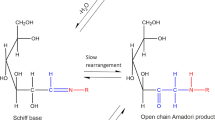
Carbohydrates with a free reducing end can react spontaneously with amines of proteins. This process is called glycation and stands in sharp contrast to glycosylation, which comprises all enzymatic attachments of mono- or oligosaccharides to amino acid side chains in proteins. Spontaneous glycation between an aldehyde group of an aldose moiety leads via a Schiff base and the Amadori rearrangement to Amadori products (Fig. 1), which typically exist as a mixture of different forms (Fig. 2). For example, in the case of glucose-based glycation the Amadori product exists as a mixture of 70% β-pyranose, 13% α-furanose, 13% β-furanose and 4% α-pyranose form (Mossine et al. 1994; Kaufmann et al. 2016; Moises et al. 2022). Glycation is a reversible process (Xu et al. 2022) unless the Amadori products react further to advanced glycation end products (AGEs) (Goldin et al. 2006).
Glycation of proteins with glucose is very abundant. It occurs for example in the bloodstream, in which always a certain glucose concentration is present (Xu et al. 2022), or during the production of therapeutic proteins due to glucose in the medium and cytosol (Quan et al. 2008; Beck and Liu 2019). Glucose-based glycation is very well studied, also because it is a critical quality attribute in therapeutic proteins (Alt et al. 2016; Sjögren et al. 2016). It is mainly detected and monitored by mass-spectrometry coupled with high-performance liquid chromatography (Schmutzler and Hoffmann 2022; Rabbani et al. 2016; Soboleva et al. 2017; Zhang et al. 2011) but also recently by NMR spectroscopy (Kaufmann et al. 2016; Moises et al. 2022).
Glycation with maltose is much less investigated (Leblanc et al. 2016; Montgomery et al. 2010; Krause et al. 2003), although it is also highly relevant for protein therapeutics, as high maltose concentrations are used in drug formulation buffers of e.g. abatacept and tositumomab (Strickley and Lambert 2021; Lynaugh et al. 2013).
After observing unknown NMR correlation patterns in a sample of abatacept, an Fc fusion protein with maltose in the formulation buffer, we suspected glycation by maltose as origin. Indeed, maltose-based glycation in abatacept was previously detected by HPLC-MS (Lynaugh et al. 2013). To find potential characteristic NMR correlation patterns, we undertook a systematic NMR analysis of Amadori products formed from maltose and the model protein BSA.
2D NMR spectroscopy developed into a versatile technique for the identification of different PTMs. For example glycans in small or even large proteins can be directly observed with 1H-13C correlations due to fast tumbling or their intrinsic flexibility (Unione et al. 2019; de Beer et al. 1994). However, to ensure that the signals of the PTMs are not impaired by line broadening especially in large proteins an approach for denaturing was developed that is compatible with 1H-13C correlations (Schubert et al. 2015). Using deuterated urea in D2O, in which the lyophilized protein of interest is dissolved, and adding reducing agent for eliminating disulfide bonds results in simplified spectra showing signals with random coil chemical shifts. Ideally, each PTM is recognized by a unique chemical shift correlation pattern that is sufficiently different from random coil correlations of the proteinogenic amino acids within a protein as illustrated for such diverse modifications as specific glyco epitopes (Unione et al. 2019; Schubert et al. 2015; Hinterholzer et al. 2022; Hargett et al. 2021; Peng et al. 2018), oxidation products (Hinterholzer et al. 2020), deamidation products (Grassi et al. 2017), pyroglutamate (Hinterholzer et al. 2019), aspartate isomerization (Hinterholzer et al. 2021) and glycation (Moises et al. 2022). Typically the observed patterns are so characteristic that an unambiguous identification of a certain modification is achieved, which is very reliable and orthogonal to HPLC-MS2 techniques.
Here we present characteristic NMR correlation patterns of maltose-based glycation in 1H-13C and 1H-1H correlation spectra. These patterns are suitable for an unambiguous detection of glycation by maltose in proteins. Independently we use HPLC-MS/MS to confirm maltose glycation. We illustrate that the presented NMR approach is complementary to MS/MS-based methods and is suited as an independent standard for cross-validation.
Material and methods
Procedure for the glycation of bovine serum albumin with maltose
Bovine serum albumin (BSA, Sigma A7030, 70 mg) was dissolved in 3.3 mL of 50 mM KH2PO4/K2HPO4 pH 7.4, 100 mM NaCl buffer and mixed with 6.7 mL 276.4 mM maltose monohydrate (Fluka 63419). The solution was incubated for 11 days at 40 °C. After incubation the buffer was exchanged by ultrafiltration to ddH2O using Amicon (Sigma Aldrich Amicon Ultra-15, UFC9030) with a cutoff of 30 kDa. 100 µL of the obtained 1.5 mL solution was taken for MS/MS analysis. Subsequently, the remaining sample was lyophilized and then dissolved in 550 µL of a 7 M urea-d4 (Armar Chemicals 049500,3041) solution in D2O (Armar Chemicals 014400,0010) for NMR analysis. The pH* (uncorrected readout measured in D2O) was adjusted to 7.4 by adding 3% DCl in D2O (Armar Chemicals 042100.0035). To reduce the disulfide bonds DTT-d10 (Cambridge Isotope Laboratories DLM-2622-0) was added to a concentration of 67 mM. The sample was heated to 60 °C for 15 min. The final protein concentration was ~ 1.9 mM.
Sample preparation of abatacept for NMR analysis
The sample preparation was previously described (Hinterholzer et al. 2022). In brief, abatacept (ORENCIA®, Bristol Myers Squibb; Lot. OE61132, exp. 08/2012) 60 mg in 2.4 mL formulation buffer was dialyzed twice against 4 L ddH2O overnight using a SpectraPor membrane with a cut-off of 3.5 kDa. After lyophilization the sample was dissolved in 650 μL of a 7 M urea-d4 (98 atom%D, ARMAR Chemicals) solution in D2O. DTT-d10 (Cambridge Isotope Laboratories) was added to a concentration of 15 mM, and after an incubation for 15 min at 60 °C, the pH* was adjusted to 7.4 using NaOD (Armar Chemicals).
NMR spectroscopy
Spectra were measured on a 600 MHz Bruker Avance III HD spectrometer equipped with a 1H/13C/15N/31P quadruple-resonance room temperature probe at 298 K. All samples were measured in a standard 5 mm NMR tube (Armar, Type 5TA) with a volume of 500 µL or 550 µL. For assigning the resonances of the Amadori products, the following 2D experiments were recorded: 1H–13C HSQC, 1H–13C HMBC (hmbcgpndqf), 1H–1H TOCSY with mixing times of 100 ms and 12 ms, 1H–1H COSY (cosygpppqf), 1H–13C HSQC–TOCSY (hsqcdietgpsisp.2) with mixing times of 13 ms and 100 ms. More details of the experimental parameters are given in the Figure captions. The data was processed with Topspin 3.6.2 (Bruker) and analyzed with Sparky 1.470 (Lee et al. 2015).
Sample preparation for HPLC–MS/MS analysis
Ultrapure water was produced with a MilliQ® Integral 3 instrument (Millipore, Billerica, MA, USA). Triethylammonium bicarbonate buffer (TEAB, pH 8.50 ± 0.10, 1 mol L−1), sodium dodecyl sulfate (SDS, ≥ 99.5%), tris(2-carboxyethyl)phosphin-hydrochlorid (TCEP, ≥ 98.0%), iodoacetamide (IAA, ≥ 99.0%), formic acid (FA, 98.0–100%) and trifluoroacetic acid (TFA, ≥ 99.0%) were acquired from Sigma-Aldrich (Vienna, Austria). Methanol (MeOH, LiChrosolv®) and ortho-phosphoric acid (85%) were purchased from Merck (Darmstadt, Germany). Acetonitrile (ACN, LC–MS grade) was purchased from “VWR International” (Vienna, Austria). Trypsin (sequencing grade modified, porcine) was acquired from Promega (Madison, WI, USA). 74 µg of glycated BSA (after buffer exchange against ddH2O, pH 7.4, conc.: 7.4 mg.mL−1) were diluted to a concentration of 1.6 µg.µL−1 in 50 mmol L−1 TEAB (pH 8.50) buffer containing 5% (w/w) SDS and denatured by heating for 5 min at 95 °C. Next, disulfides were reduced by addition of TCEP to a concentration of 5.0 mmol L−1 and incubation at 55 °C for 15 min, followed by alkylation of the cysteine residues by addition of IAA to a concentration of 40 mmol L−1 and incubation at 22 °C in the dark for 10 min. Following, the protein was precipitated at a pH ≤ 1 with 12% (v/v) ortho-phosphoric acid and by adding 7:1 (v/v) of 100 mmol L−1 TEAB (pH 7.55) in 90% MeOH (v/v). Next, the proteins were purified by suspension trapping employing S-Trap mini columns (Protifi, Huntington, NY, USA) according to the manufacturer’s instructions, and digested to peptides employing trypsin at a protease/protein ratio of 1:10 (w/w) at 37 °C for 12 h. The obtained peptides were dried at 50 °C in a vacuum centrifuge and resuspended in 1% ACN + 0.10% FA to a concentration of 3.3 µg µL−1.
High-performance liquid chromatography coupled to MS/MS
Chromatographic separation of 1.0 µg peptides was carried out in five technical replicates employing reversed-phase HPLC on an UltiMate™ 3000 RSLCnano System (Thermo Fisher Scientific, Germering, Germany), on a DNV PepMap™ Neo column (150 × 0.075 mm i.d.) from Thermo Fisher Scientific, Germering, Germany. The mobile phases used for the separation were 0.10% aqueous FA (solvent A) and 0.10% FA in ACN (solvent B), pumped at a flow rate of 200 nL.min−1 in the following order: 1.0% B for 5.0 min, a linear gradient from 1.0 to 5.0% B in 5 min, a second linear gradient from 5.0 to 35.0% B in 60.0 min, and a third linear gradient from 35.0 to 45.0% B in 20.0 min. This was followed by flushing at 99.0% B for 10 min and column re-equilibration at 1.0% B for 35 min. The column temperature for the separation was kept constant at 50 °C. The nanoHPLC system was hyphenated to a Q Exactive™ Hybrid Quadrupole-Orbitrap™ mass spectrometer via a Nanospray Flex™ ion source (both from Thermo Fisher Scientific, Bremen, Germany). The source was equipped with a SilicaTip emitter with 360 µm o.d., 20 µm i.d. and a tip i.d. of 10 µm from CoAnn Technologies Inc. (Richland, WA, USA). The spray voltage was set to 1.5 kV, S-lens RF level to 60.0 and capillary temperature to 250 °C. Each scan cycle consisted of a full scan at a scan range of m/z 350–2000 and a resolution setting of 70,000 at m/z 200, followed by 5 data-dependent higher-energy collisional dissociation (HCD) scans in a 2.0 m/z isolation window at 28% normalized collision energy at a resolution setting of 17,500 at m/z 200. For the full scan, the automatic gain control (AGC) target was set to 3e6 charges with a maximum injection time of 100 ms, for the HCD scans the AGC target was 1e5 charges with a maximum injection time of 150 ms. Already fragmented precursor ions were excluded for 10 s. Data acquisition was conducted using Thermo Scientific™ Chromeleon™ 7.2 CDS (Thermo Fisher Scientific, Germering, Germany). For the identification of modification sites, as well as for sequence coverage mapping, Byonic 3.11.3 (Protein Metrics, Cupertino, CA, USA) was used with a precursor and a fragment mass tolerance of 10 ppm, applying a 1% false discovery rate. Relative quantification of the modified peptides was done using MaxQuant 2.0.1.0 (Cox and Mann 2008) with the setting Label-free quantification, applying a 1% false discovery rate.
Results
Assignment of NMR correlation patterns of Amadori products of maltose in the model protein BSA
To obtain a suitable sample for studying the NMR correlation patterns of maltose-glycation products, we incubated bovine serum albumin (BSA) with a high concentration of maltose at pH 7.4. We chose BSA, because it produced earlier high amounts of glycation products with glucose (Moises et al. 2022). BSA contains several lysines with low pKa values, which are susceptible to glycation. Specifically, we achieved glycation by incubating a solution of 11 mM BSA with 185 mM maltose for 11 days at 40 °C at pH 7.4. After buffer exchange to ddH2O, the treated protein was analyzed under denaturing conditions in a completely deuterated solution (7 M urea-d4 in D2O, 67 mM DTT-d10). Although not relevant for this study, we observed phase separation after buffer exchange of the glycated protein to ddH2O. However, after lyophilization, the sample completely dissolved under denaturing conditions.
The signal-to-noise was sufficient to obtain high-quality 2D NMR data for all species including less sensitive 1H-13C correlation spectra. Figure 3 shows 2D 1H-13C HSQC spectra comparing untreated BSA, glucose-based glycated BSA and maltose-based glycated BSA. Both glycated forms showed many new signals in the carbohydrate region between 71 and 105 ppm. The new sets of signals are very different for glucose-based glycated BSA compared to maltose-based glycated BSA. The anomeric region of the 2D 1H-13C HSQC spectrum (90–105 ppm) showed only in the case of maltose-based glycation three C1-H1 correlations of the terminal Glc moieties of the glycated forms in addition to signals of free sugar.
2D 1H-13C correlations comparing untreated BSA with glucose-based glycated BSA and maltose-based glycated BSA. a 1H-13C HSQC spectrum of untreated BSA. b 1H-13C HSQC spectrum of glucose-based glycated BSA. Grey arrows indicate prominent signals of free glucose. The region indicated by the red arrow is typical for the furanose forms of the Amadori product, the region marked by the cyan arrow indicates the β-pyranose form of the Amadori product. c 1H-13C HSQC spectrum of maltose-based glycated BSA. The same color code as in section b is used for the arrows. Interestingly, three additional anomeric signals of the Amadori product are visible indicated by a magenta arrow
A detailed 2D 1H-13C HSQC spectrum of maltose-based glycated BSA with all relevant signals labeled is shown in Fig. 4. Three anomeric signals of the free maltose are visible (Glc1′, Glc1α and Glc1β) likely due to the back reaction of the reversible glycation reaction. Besides signals of free maltose in the sample, we observed three forms of Amadori products: the dominating β-pyranose form and two furanose forms with populations of 60%, 22% and 18% (Fig. 4a, b), respectively. The populations were estimated from peak volumes of the C1-H1 correlations in the 2D 1H-13C HSQC spectrum. Further, a comparison of the peak volumes with the Cβ-Hβ signal of all isoleucines (BSA contains 14 Ile residues) we could estimate an averaged glycation of 3.7 sites per BSA molecule.
Maltose-based glycation products observed with the model protein BSA. a 1H-.13C HSQC spectrum of glycated BSA at a concentration of 2 mM. The sample was dialyzed against ddH2O, lyophilized and dissolved in 7 M urea-d4 in D2O pH* 7.4. The β-pyranose form, the most dominant form of the Amadori product, is labeled in cyan. The furanose forms are labeled in red and purple. The chemical structures of these species are shown on the right using the same color code. Signals of free maltose, which can form in a back-reaction, are shown with grey labels. b Chemical structures of the observed Amadori products with their observed abundances and symbol presentation (Varki et al. 2015; Neelamegham et al. 2019)
The major form, the β-pyranose, could be readily assigned using 1H-1H TOCSY, 1H-1H COSY, 1H-13C HSQC and 1H-13C HMBC spectra (Figs. 4, 5 and Suppl. Fig. S1). The chemical shift assignment of the β-pyranose agrees with the previously reported data by (Krause et al. 2003), but the chemical shift assignments for the distal Glc moiety disagree (Table 1). However, with 1H-1H COSY, 1H-1H TOCSY and 1H-13C HSQC correlations we could unambiguously assign the distal Glc moiety (see Figs. 4 and 5). HMBC correlations between H1′ and C4 and between H4 and C1′ confirmed the linkage between the terminal Glc moiety and the first fructolysine moiety (Fig. S1).
Chemical shift assignment of maltose-based glycation products observed with the model protein BSA. a 2D 1H-1H TOCSY spectrum with correlations of the anomeric H1 protons of the distal Glc residue of the Amadori products (cyan, red, purple) and a rest of free maltose (grey). b Comparable region of a 2D 1H-1H COSY spectrum with correlations of the anomeric H1 protons of the distal Glc residue of the Amadori products (cyan, red, purple) and a rest of free maltose (grey). c Schematic presentation of the shown TOCSY correlations on the chemical structures of all three Amadori products indicated as arrows. d Observed COSY correlations schematically illustrated on the chemical structures
For the assignment of the minor forms, we recorded in addition 1H-1H NOESY (Fig. 6) and 1H-13C HSQC-TOCSY spectra with different mixing times (data not shown). The 1H-13C HSQC-TOCSY spectrum with a short mixing time (13 ms) shows typically at one 13C frequency correlations to three 1H frequencies—of the directly attached 1H and of the next 1H neighbors. Almost complete chemical shift assignments were obtained, and the three sets of products could be unambiguously assigned to the β-pyranose, β-furanose and α-furanose forms. In addition, the random coil chemical shifts of the modified lysines could be assigned (Table 1). Here especially the Cε–Hε correlation is characteristic. It is very similar to glucose-based glycation.
Key NOE correlations of maltose-based glycation products observed with the model protein BSA. a 2D 1H-1H NOESY spectrum of the anomeric H1 protons of the distal Glc residue showing correlations to the proximal fucose residues for all three observed Amadori products. b Observed NOE correlations schematically shown on the chemical structures
Independent confirmation of maltose-based glycation by MS/MS analysis
BSA glycated by maltose was independently analyzed by HPLC-MS/MS analysis. Figure 7 shows exemplary data of the peptide (Ala249 –Lys263) unmodified and glycated at Lys256. A mass difference of 324.106 Da was observed for the parent ions corresponding to a modification of maltose consisting of two hexoses. The sequence of the peptide is almost completely covered by y ions in both cases. In the maltosylated variant (Fig. 7b), the fragments from y8 to y14 are observed as furylium (+ 78 Da) and pyrylium (+ 108 Da), products of typical dissociation pathways for peptides glycated with aldohexoses (Corzo-Martinez et al. 2009). The sequence of BSA was well covered (Supplementary Fig. S2) providing evidence for the glycation of 51 out of all 59 lysine residues (Supplementary Fig. S3 and Table S1). The individual abundances of glycation ranged from 0.4 to 98.2% (Suppl. Table S1).
Exemplary MS/MS spectra of the BSA peptide (Ala249 –Lys263) unmodified [uncharged mass 1691.935 Da] (a) and glycated at Lys256 [uncharged mass 2016.040 Da] (b), corresponding to a delta mass of 324.106 Da. The sequence of the peptide is almost completely covered by y ions in both cases. In the maltosylated variant (b), the fragments from y8-y14 are observed as furylium (+ 78 Da) and pyrilium (+ 108 Da), representing typical dissociation pathways for peptides glycated with aldohexoses
Detection of maltose-based glycation in a biotherapeutic Fc-fusion protein
In contrast to glycation with Glc, whose Amadori product lacked an anomeric proton, the anomeric signals of the terminal Glc moiety of the maltose-Amadori product are well separated for the different forms in a 1H-1H TOCSY (Fig. 5). More importantly, they occur in a characteristic region, which is normally quite empty and not disturbed by neither protein nor glycosylation signals. These patterns of chemical shift correlations can be used to unambiguously identify the presence of maltose-based glycation in any protein sample.
We decided to analyze the biotherapeutic abatacept, which is stored with a high amount of maltose in its formulation buffer and of which maltose-glycation was reported earlier (Lynaugh et al. 2013). Figure 8a shows the relevant region of a 2D 1H-1H TOCSY of abatacept under denaturing conditions in comparison to the here assigned maltose-based glycated BSA (Fig. 8b).
Unambiguous identification of maltose-based glycation in the biotherapeutic abatacept. a 2D 1H-1H TOCSY spectrum of the dialyzed and lyophilized abatacept dissolved in 7 M urea-d4 in D2O (pH* 7.4) showing the anomeric region. Correlations of all three forms of maltose-based glycation are visible despite the busy background of n-glycosylation signals. b Comparable region of a 2D 1H-1H TOCSY spectrum of maltose-glycated BSA
The presence of maltose-based glycation in a protein is indicated by a characteristic pattern of 4 strong signals of the β-pyranose form. Both fucose forms give additionally very characteristic patterns consisting of 6 signals each, however, with lower intensity due to their lower abundance. Monitoring this region of a 2D 1H-1H TOCSY is an ideal method to unambiguously recognize maltose adducts in proteins, e.g. in biotherapeutic proteins or proteins within foods. 1H-13C HSQC correlations are also very characteristic and unique. However, such 1H-13C spectra are less sensitive due to the low natural abundance of 13C of only 1.1% and require therefore much longer measurement times like 24 h to obtain a sufficient signal-to-noise ratio. From comparing the weak C4–H4 signal of the proximal ring of the β-pyranose form in the 1H-13C HSQC spectrum and Cβ-Hβ signal of all Ile residues (abatacept dimer contains 22 Ile residues), we estimate that on average 16.7% of all Fc-fusion dimers are glycated.
Discussion
The chemical shift patterns of the maltose-based glycation products in 2D NMR spectra are clearly different than those of glucose-based glycation. This allows the unambiguous distinction between these two different modifications due to glycation resulting from different sugars with free reducing ends.
The characteristic signals of the terminal Glc of maltose, which are well isolated in a sensitive 1H-1H TOCSY spectrum, enable the straightforward and sensitive detection of maltose glycation. This stands in contrast to glycation by the monosaccharide glucose whose anomeric signal is lost after the Amadori rearrangement, and whose presence is only detectable by characteristic NMR correlation patterns using 1H-13C HSQC correlations, which are much less sensitive due to the low natural abundance of 13C of 1.1%. Here 1H-1H TOCSY spectra can be recorded within the time of approx. 1 h depending on the recycle delay and the resolution chosen in the indirect dimension. It is not so common to have isolated and characteristic signals of PTMs in 1H-1H TOCSY spectra, because dispersion in 1H is much smaller than in 13C, but rare cases like characteristic signals of α-Gal epitopes (Hinterholzer et al. 2022) and now maltose-glycation allow the unambiguous detection in a time efficient and easy to interpret manner.
The observed populations of the different Amadori products were different compared to glucose-based glycation. We observed here 60% β-pyranose, 22% β-furanose and 18% α-furanose for maltose-based glycation, whereas the α-pyranose was either absent or too weak to be observed. For glucose-based glycation the populations were 70% β-pyranose, 13% β-furanose, 13% α-furanose and 4% α-pyranose (Mossine et al. 1994; Moises et al. 2022). This indicates that the α1,4-linked glucose substitution influences the equilibrium of the Amadori products: although the β-pyranose form is still dominating, it is less populated; both furanose forms are more abundant, but the β-furanose is favored.
The lower detection limit of the approach is in principle independent of the kind of modification. We determined in an earlier study an absolute amount of ~ 28 nmol for the oxidation product of methionine using 1H-13C HSQC spectra recorded in 24 h on a 900 MHz spectrometer with cryogenic probe (Hinterholzer et al. 2020). However, this depends on the sensitivity of the spectrometer and the measurement time. A similar detection limit was reported by Peng et al. (Peng et al. 2018). In the case of the denatured abatacept sample with a concentration of 1.2 mM (dimer), this lower limit corresponds to an average modification of 5% of all dimers. For 1H-1H correlations it will be even lower.
The strength of the presented NMR approach is the unambiguous identification of maltose-based glycation and the possibility of quantifying the total amount. A weakness is that it does not provide sequence specific localization and that the detection limit of NMR is orders of magnitudes higher than MS-based techniques. However, it is quite complementary to HPLC-MS2 methods, which are much more sensitive and can provide sequence specific modifications, but quantification is less accurate and identification of a modification is sometimes ambiguous. The same mass difference can have several origins, e.g. a glucose-based glycation site or an extra hexose within a glycan, or a maltose-based glycation within a peptide versus two glucose-based glycation sites. Another disadvantage of HPLC-MS2 methods is that the investigated modification should typically be known beforehand to set up the experiments appropriately. Another advantage of NMR spectroscopy is that with either a single spectrum (HSQC) or two spectra (TOCSY in addition) one gets an overview of many different modifications that are present in a sample, e.g. a therapeutic protein. Even unknown modifications could be detected if they result in characteristic patterns. The complementarity of the methods allows the unambiguous identification of a certain type of modification with NMR, to detect it with much lower sensitivity with MS-based techniques and to cross-validate or even calibrate MS-based quantifications.
It did not escape our attention that the maltose-glycation of a protein might serve as a very simple model system to study detailed mechanisms of reductive amination. The proximal glucose at the reducing end might serve as a model for any glucose at the reducing end and the distal glucose is a very simple substitution so that the system can still be easily studied. That is much different compared to studying larger oligosaccharides or long polysaccharides linked to proteins by reductive amination, particles or surfaces (Gildersleeve et al. 2008; Munster et al. 2017). In the case of a polysaccharide-protein conjugate the abundance and signal intensity of the reducing end is typically very low and NMR line widths are much larger at the connection point of two polymers making it practically impossible to observe small populations of reaction intermediates or the products at the linkage site. Functionalized particles or surfaces are even more difficult to study.
In conclusion, we found characteristic NMR correlation patterns of maltose-based glycation in 1H-13C HSQC and in 1H-1H TOCSY spectra that are suitable to unambiguously identify the presence of glycation by maltose. The detection by very sensitive 1H-1H TOCSY spectra is very competitive, because it relies exclusively on 1H nuclei with 99.99% abundance in contrast to 13C with only 1.11%. This approach is complementary to MS-based methods, it is suited as an independent standard for cross-validation.
References
Alt N et al (2016) Determination of critical quality attributes for monoclonal antibodies using quality by design principles. Biologicals 44:291–305
Beck A, Liu HC (2019) Macro- and micro-heterogeneity of natural and recombinant IgG antibodies. Antibodies 8:18
Corzo-Martinez M, Lebron-Aguilar R, Villamiel M, Quintanilla-Lopez JE, Moreno FJ (2009) Application of liquid chromatography-tandem mass spectrometry for the characterization of galactosylated and tagatosylated beta-lactoglobulin peptides derived from in vitro gastrointestinal digestion. J Chromatogr A 1216:7205–7212
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26:1367–1372
de Beer T et al (1994) Rapid and simple approach for the NMR resonance assignment of the carbohydrate chains of an intact glycoprotein - application of gradient-enhanced natural-abundance 1H-13C HSQC and HSQC-TOCSY to the alpha-subunit of human chorionic-gonadotropin. FEBS Lett 348:1–6
Gildersleeve JC, Oyelaran O, Simpson JT, Allred B (2008) Improved procedure for direct coupling of carbohydrates to proteins via reductive amination. Bioconjugate Chem 19:1485–1490
Goldin A, Beckman JA, Schmidt AM, Creager MA (2006) Advanced glycation end products. Circulation 114:597–605
Grassi L et al (2017) Complete NMR assignment of succinimide and its detection and quantification in peptides and intact proteins. Anal Chem 89:11962–11970
Hargett AA et al (2021) Glycosylation states on intact proteins determined by NMR spectroscopy. Molecules 26:4308
Hinterholzer A, Stanojlovic V, Cabrele C, Schubert M (2019) Unambiguous identification of pyroglutamate in full-length biopharmaceutical monoclonal antibodies by NMR spectroscopy. Anal Chem 91:14299–14305
Hinterholzer A et al (2020) Identification and quantification of oxidation products in full-length biotherapeutic antibodies by NMR spectroscopy. Anal Chem 92:9666–9673
Hinterholzer A et al (2021) Detecting aspartate isomerization and backbone cleavage after aspartate in intact proteins by NMR spectroscopy. J Biomol NMR 75:71–82
Hinterholzer A et al (2022) Unambiguous identification of alpha-Gal epitopes in intact monoclonal antibodies by NMR spectroscopy. mAbs 14:2132977
Kaufmann M, Meissner PM, Pelke D, Mügge C, Kroh LW (2016) Structure–reactivity relationship of Amadori rearrangement products compared to related ketoses. Carbohydr Res 428:87–99
Krause R, Knoll K, Henle T (2003) Studies on the formation of furosine and pyridosine during acid hydrolysis of different Amadori products of lysine. Eur Food Res Technol 216:277–283
Leblanc Y et al (2016) Glycation of polyclonal IgGs: Effect of sugar excipients during stability studies. Eur J Pharm Biopharm 102:185–190
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327
Lynaugh H, Li HJ, Gong B (2013) Rapid Fc glycosylation analysis of Fc fusions with IdeS and liquid chromatography mass spectrometry. mAbs 5:641–645
Moises JE, Regl C, Hinterholzer A, Huber CG, Schubert M (2022) Unambiguous identification of glucose-induced glycation in mAbs and other proteins by NMR spectroscopy. Pharm Res. 40:1341–1353. https://doi.org/10.1007/s11095-022-03454-0
Montgomery H, Tanaka K, Belgacem O (2010) Glycation pattern of peptides condensed with maltose, lactose and glucose determined by ultraviolet matrix-assisted laser desorption/ionization tandem mass spectrometry. Rapid Commun Mass Sp 24:841–848
Mossine VV, Glinsky GV, Feather MS (1994) The preparation and characterization of some Amadori compounds (1-amino-1-deoxy-D-fructose derivatives) derived from a series of aliphatic omega-amino acids. Carbohydr Res 262:257–270
Neelamegham S et al (2019) Updates to the symbol nomenclature for glycans guidelines. Glycobiology 29:620–624
Peng J, Patil SM, Keire DA, Chen K (2018) Chemical structure and composition of major glycans covalently linked to therapeutic monoclonal antibodies by middle-down nuclear magnetic resonance. Anal Chem 90:11016–11024
Quan C et al (2008) A study in glycation of a therapeutic recombinant humanized monoclonal antibody: where it is, how it got there, and how it affects charge-based behavior. Anal Biochem 373:179–191
Rabbani N, Ashour A, Thornalley PJ (2016) Mass spectrometric determination of early and advanced glycation in biology. Glycoconjugate J 33:553–568
Schmutzler S, Hoffmann R (2022) Chromatographic separation of glycated peptide isomers derived from glucose and fructose. Anal Bioanal Chem 414:6801–6812
Schubert M, Walczak MJ, Aebi M, Wider G (2015) Posttranslational modifications of intact proteins detected by NMR spectroscopy: application to glycosylation. Angew Chem Int Ed 54:7096–7100
Sjögren J, Olsson F, Beck A (2016) Rapid and improved characterization of therapeutic antibodies and antibody related products using IdeS digestion and subunit analysis. Analyst 141:3114–3125
Soboleva A, Vikhnina M, Grishina T, Frolov A (2017) Probing protein glycation by chromatography and mass spectrometry: analysis of glycation adducts. Int J Mol Sci 18:2557
Strickley RG, Lambert WJ (2021) A review of formulations of commercially available antibodies. J Pharm Sci-US 110:2590–2608
Unione L et al (2019) Glycoprofile analysis of an intact glycoprotein as inferred by NMR spectroscopy. ACS Cent Sci 5:1554–1561
van Munster JM et al (2017) Application of carbohydrate arrays coupled with mass spectrometry to detect activity of plant-polysaccharide degradative enzymes from the fungus Aspergillus niger. Sci Rep 7:43117
Varki A et al (2015) Symbol nomenclature for graphical representations of glycans. Glycobiology 25:1323–1324
Xu X et al (2022) Low pKa of Lys promotes glycation at one complementarity-determining region of a bispecific antibody. Biophys J 121:1081–1093
Zhang Q et al (2011) Comprehensive identification of glycated peptides and their glycation motifs in plasma and erythrocytes of control and diabetic subjects. J Proteome Res 10:3076–3088
Acknowledgements
We acknowledge Dr. Urs Lohrig and Novartis Kundl for generously providing a sample of abatacept. The authors thank Fionna Loughlin (Monash University), Ryan McKay (University of Alberta) and Tim Keys (ETH Zurich) for fruitful discussions and Lawrence P. McIntosh (University of British Columbia) Sebastian Schwap (Technical University of Munich) for carefully reading the manuscript and comments.
Funding
Open access funding provided by Paris Lodron University of Salzburg.
Author information
Authors and Affiliations
Contributions
PD, CR and MS planned and carried out the experiments and analyzed data. NMR spectra were recorded by MS. NMR resonances were assigned by PD under the supervision of MS. The manuscript was written by MS, CR and CH and PD. The Figures were prepared by CR and MS.
Corresponding author
Ethics declarations
Conflict of interest
The Author declares that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Defant, P., Regl, C., Huber, C.G. et al. The NMR signature of maltose-based glycation in full-length proteins. J Biomol NMR 78, 61–72 (2024). https://doi.org/10.1007/s10858-023-00432-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-023-00432-5