Abstract
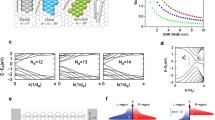
The effects of histidine and its imidazole ring adsorption on the electronic transport properties of graphene were investigated by first-principles calculations within a combination of density functional theory and non-equilibrium Greens functions. Firstly, we report adsorption energies, adsorption distances, and equilibrium geometrical configurations with no bias voltage applied. Secondly, we model a device for the transport properties study: a central scattering region consisting of a finite graphene sheet with the adsorbed molecule sandwiched between semi-infinite source (left) and drain (right) graphene electrode regions. The electronic density, electrical current, and electronic transmission were calculated as a function of an applied bias voltage. Studying the adsorption of the two systems, i.e., the histidine and its imidazole ring, allowed us to evaluate the importance of including the carboxyl (–COOH) and amine (–\(\hbox {NH}_{2}\)) groups. We found that the histidine and the imidazole ring affects differently the electronic transport through the graphene sheet, posing the possibility of graphene-based sensors with an interesting sensibility and specificity.







Similar content being viewed by others
References
Geim, A.K., Novoselov, K.S.: The rise of graphene. Nat. Mater. 6, 183–191 (2007)
Ferrari, A.C.: Science and technology roadmap for graphene, related two-dimensional crystals, and hybrid systems. Nanoscale 112, 1–343 (2014)
Baraket, M., Stine, R., Lee, S., Robinson, J., Tamanaha, C.R., Sheehab, P.E., Walton, S.G.: Aminated graphene for DNA attachment produced via plasma functionalization. Appl. Phys. Lett. 100, 233123 (2012)
Huang, B., Li, Z., Liu, Z., Zhou, G., Hao, S., Wu, J., Gu, B.L., Duan, W.: Adsorption of gas molecules on graphene nanoribbons and its implication for nanoscale molecule sensor. J. Phys. Chem. C 112, 13442–13446 (2008)
Georgakilas, V., Otyepka, M., Bourlinos, A.B., Chandra, V., Kim, N., Kemp, K.C., Hobza, P., Zboril, R., Kim, S.: Functionalization of graphene: covalent and non-covalent approaches, derivates and applications. Chem. Rev. 1, 1–58 (2012)
Milowska, K., Majewski, J.: Graphene-based sensors: theoretical study. J. Phys. Chem. C 118, 17395–17401 (2014)
Shao, Y., Wang, J., Wu, H., Liu, J., Aksay, I.A., Lin, Y.: Graphene based electrochemical sensors and biosensors: a review. Electroanalysis 22, 1027–1036 (2010)
Liu, Y., Yu, D., Zeng, C., Miao, Z., Dail, L.: Biocompatible graphene oxide-based glucose biosensors. Langmuir 26, 6158–6160 (2010)
Yang, K., Feng, L., Shi, X., Liu, Z.: Nano-graphene in biomedicine: theranostic applications. Chem. Soc. Rev. 42, 530–547 (2013)
Nelson, T., Zhang, B., Prezhdo, O.: Detection of nucleic acids with graphene nanopores: ab initio characterization of a novel sequencing device. Nano Lett. 10, 3237–3242 (2010)
Huang, Y., Dong, X., Liu, Y., Li, L.J., Chen, P.: Graphene-based biosensors for detection of bacteria and their metabolic activities. J. Mater. Chem. 21, 12358–12362 (2011)
Notley, S.M., Crawford, R., Ivanova, E.P.: Bacterial interaction with graphene particles and surfaces. Nanotechnol. Nanomater. 5, 99–118 (2013)
Song, B., Cuniberti, G., Sanvito, S., Fang, H.: Nucleobase adsorbed at graphene devices: enhance bio-sensorics. Appl. Phys. Lett. 100, 063101 (2012)
Lee, E.C.: Effects of DNA nucleotide adsorption on the conductance of graphene nanoribbons from first principles. Appl. Phys. Lett. 100, 153117 (2012)
Zhang, Y.H., Zhou, K.G., Xie, K.F., Zeng, J., Zhang, H., Peng, Y.: Tuning the electronic structure and transport properties of graphene by noncovalent functionalization: effects of organic donor, acceptor and metal atoms. Nanotechnology 21, 065201 (2010)
Rodríguez, S., Makinistian, L., Albanesi, E.: Theoretical study of the adsorption of histidine amino acid on graphene. J. Phys. Conf. Ser. 705, 012012 (2016)
Ortmann, F., Schmidt, W., Bechstedt, F.: Attracted by long-range electron correlation: adenine on graphite. Phys. Rev. Lett. 95, 186101 (2005)
Lee, J., Choi, Y., Kim, H., Scheicher, R., Cho, J.: Physisorption of DNA nucleobases on h-BN and graphene: vdW-corrected DFT calculations. J. Phys. Chem. C 117, 13435–13441 (2013)
Grimme, S.: Semiempirical GGA-type density functional constructed with a long-range dispersion correction. J. Comput. Chem. 27, 1787 (2006)
Ozaki, T.: Numerical atomic basis orbitals from H to Kr. Phys. Rev. B. 67, 155108 (2003)
Ozaki, T., Nishio, K., Kino, H.: Efficient implementation of the nonequilibrium Green function method for electronic transport. Phys. Rev. B 81, 035116 (2010)
Rajesh, C., Majumder, C., Mizuseki, H., Kawazoe, Y.: A theoretical study on the interaction of aromatic amino acids with graphene and single walled carbon nanotube. J. Chem. Phys. 130, 124911 (2009)
You, X., Pak, J.J.: Graphene-based field effect transistor enzymatic glucose biosensor using silk protein for enzyme immobilization and device substrate. Sens. Actuators B 202, 1357–1365 (2014)
Cai, B., Wang, S., Huang, L., Ning, Y., Zhang, Z., Zhang, G.J.: Ultrasensitive label-free detection of PNA-DNA hybridization by reduced graphene oxide field-effect transistor biosensor. ACS Nano 8, 2632–2638 (2014)
Zhang, T., Nix, M.B., Yoo, B.Y., Deshusses, M.A., Myung, N.V.: Electrochemically functionalized single-walled carbon nanotube gas sensor. Electroanalysis 12, 1153–1158 (2006)
Dong, X., Fu, D., Xu, Y., Wei, J., Shi, Y., Chen, P., Li, J.: Label-free electronic detection of DNA using simple double-walled carbon nanotube resistors. J. Phys. Chem. 112, 9891–9895 (2008)
Mao, S., Lu, G., Yu, K., Bo, Z., Chen, J.: Specific protein detection using thermally reduced graphene oxide sheet decorated with gold nanoparticle-antibody conjugates. Adv. Mater. 22, 3521–3526 (2010)
Acknowledgements
The authors acknowledge financial support from Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), through grants PIP 11220150100124CO (S.J.R. and E.A.A.) and 112-201101-00615 (L.M.). Also, L.M. and E.A.A. acknowledge financial support from the Universidad Nacional de San Luis (PROICO 3-10314), and the Universidad Nacional de Entre Ríos, respectively.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rodríguez, S.J., Makinistian, L. & Albanesi, E. Computational study of transport properties of graphene upon adsorption of an amino acid: importance of including –\(\hbox {NH}_{2}\) and –COOH groups. J Comput Electron 16, 127–132 (2017). https://doi.org/10.1007/s10825-016-0943-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10825-016-0943-x