Abstract
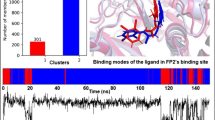
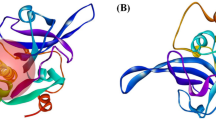
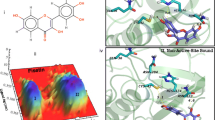
Falcipain-2 (FP-2) is a Plasmodium falciparum hemoglobinase widely targeted in the search for antimalarials. FP-2 can be allosterically modulated by various noncompetitive inhibitors that have been serendipitously identified. Moreover, the crystal structures of two inhibitors bound to an allosteric site, termed site 6, of the homolog enzyme human cathepsin K (hCatK) suggest that the equivalent region in FP-2 might play a similar role. Here, we conduct the rational identification of FP-2 inhibitors through virtual screenings (VS) of compounds into several pocket-like conformations of site 6, sampled during molecular dynamics (MD) simulations of the free enzyme. Two noncompetitive inhibitors, ZINC03225317 and ZINC72290660, were confirmed using in vitro enzymatic assays and their poses into site 6 led to calculated binding free energies matching the experimental ones. Our results provide strong evidence about the allosteric inhibition of FP-2 through binding of small molecules to site 6, thus opening the way toward the discovery of new inhibitors against this enzyme.



Similar content being viewed by others
Availability of data and material
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
MOE, Amber14, Amber18, and Gaussian 09 are commercial software. Autodock-Vina, Gromacs and Avogadro are freely available on the internet.
References
Wells TN, Hooft van Huijsduijnen R, Van Voorhis WC (2015) Nat Rev Drug Discov 14:424–442. https://doi.org/10.1038/nrd4573
World Malaria Report (2021) https://www.who.int/teams/global-malaria-programme/reports/world-malaria-report-2020. Accessed 27 Mar 2021
Bekono BD, Ntie-Kang F, Owono Owono LC et al (2018). Curr Drug Target. https://doi.org/10.2174/1389450117666161221122432
Teixeira C, Gomes JR, Gomes P (2011) Curr Med Chem 18:1555–1572. https://doi.org/10.2174/092986711795328328
Ettari R, Bova F, Zappala M et al (2010) Med Res Rev 30:136–167. https://doi.org/10.1002/med.20163
Roy KK (2017) Int J Antimicrob Agents 50:287–302. https://doi.org/10.1016/j.ijantimicag.2017.04.006
Rosenthal PJ (2011) Adv Exp Med Biol 712:30–48. https://doi.org/10.1007/978-1-4419-8414-2_3
Wagner JR, Lee CT, Durrant JD et al (2016) Chem Rev 116:6370–6390. https://doi.org/10.1021/acs.chemrev.5b00631
Bhat AS, Dustin Schaeffer R, Kinch L et al (2020) Curr Opin Struct Biol 62:183–188. https://doi.org/10.1016/j.sbi.2020.02.004
Bertoldo JB, Chiaradia-Delatorre LD, Mascarello A et al (2015) J Enzyme Inhib Med Chem 30:299–307. https://doi.org/10.3109/14756366.2014.920839
Alberca LN, Chuguransky SR, Alvarez CL et al (2019) Front Chem 7:534. https://doi.org/10.3389/fchem.2019.00534
Marques AF, Esser D, Rosenthal PJ et al (2013) Bioorg Med Chem 21:3667–3673. https://doi.org/10.1016/j.bmc.2013.04.047
Marques AF, Gomes PS, Oliveira PL et al (2015) Arch Biochem Biophys 573:92–99. https://doi.org/10.1016/j.abb.2015.03.007
Pant A, Kumar R, Wani NA et al (2018) Sci Rep 8:16193. https://doi.org/10.1038/s41598-018-34564-8
Hernandez Gonzalez JE, Hernandez Alvarez L, Pascutti PG et al (2019) J Phys Chem B 123:7327–7342. https://doi.org/10.1021/acs.jpcb.9b05021
Novinec M, Korenc M, Caflisch A et al (2014) Nat Commun 5:3287. https://doi.org/10.1038/ncomms4287
Novinec M, Lenarcic B, Baici A (2014) PLoS ONE 9:e106642. https://doi.org/10.1371/journal.pone.0106642
Novinec M, Rebernik M, Lenarcic B (2016) FEBS Lett 590:4507–4518. https://doi.org/10.1002/1873-3468.12495
Costa TF, dos Reis FC, Lima AP (2012) Biochim Biophys Acta 1824:493–501. https://doi.org/10.1016/j.bbapap.2011.12.006
Lima AP et al (2002) J Biol Chem 277:5875–5881. https://doi.org/10.1074/jbc.M108518200
Li Z, Kienetz M, Cherney MM et al (2008) J Mol Biol 383:78–91. https://doi.org/10.1016/j.jmb.2008.07.038
Li Z, Yasuda Y, Li W et al (2004) J Biol Chem 279:5470–5479. https://doi.org/10.1074/jbc.M310349200
Judice WA et al (2013) PLoS ONE 8:e80153. https://doi.org/10.1371/journal.pone.0080153
Almeida PC, Nantes IL, Chagas JR et al (2001) J Biol Chem 276:944–951. https://doi.org/10.1074/jbc.M003820200
Almeida PC, Nantes IL, Rizzi CC et al (1999) J Biol Chem 274:30433–30438. https://doi.org/10.1074/jbc.274.43.30433
Irwin JJ, Shoichet BK (2005) J Chem Inf Model 45:177–182. https://doi.org/10.1021/ci049714+
Hernandez Alvarez L, Barreto Gomes DE, Hernandez Gonzalez JE et al (2019) PLoS ONE 14:e0211227. https://doi.org/10.1371/journal.pone.0211227
Molecular Operating Environment (MOE) (2016) 2013.08 Chemical Computing Group Inc., Montreal
Morris GM, Huey R, Lindstrom W et al (2009) J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Trott O, Olson AJ (2010) J Comput Chem 31:455–461. https://doi.org/10.1002/jcc.21334
Rastelli G, Del Rio A, Degliesposti G et al (2010) J Comput Chem 31:797–810. https://doi.org/10.1002/jcc.21372
Hou T, Wang J, Li Y et al (2011) J Chem Inf Model 51:69–82. https://doi.org/10.1021/ci100275a
Genheden S, Ryde U (2015) Expert Opin Drug Discov 10:449–461. https://doi.org/10.1517/17460441.2015.1032936
Rastelli G, Pinzi L (2019) Front Chem 7:498. https://doi.org/10.3389/fchem.2019.00498
Nguyen H, Roe DR, Simmerling C (2013) J Chem Theory Comput 9:2020–2034. https://doi.org/10.1021/ct3010485
Miller BR 3rd, McGee TD Jr, Swails JM et al (2012) J Chem Theory Comput 8:3314–3321. https://doi.org/10.1021/ct300418h
Case DA, Ben-Shalom IY, Brozell SR et al (2018) AMBER 2018. University of California, San Francisco
Case DA, Babin V, Berryman JT et al (2014) AMBER14. University of California, San Francisco
Wang J, Wolf RM, Caldwell JW et al (2004) J Comput Chem 25:1157–1174. https://doi.org/10.1002/jcc.20035
Frisch MJ, Trucks GW, Schlegel HB et al (2009) Gaussian 09. Gaussian, Inc., Wallingford
Hanwell MD, Curtis DE, Lonie DC et al (2012) J Cheminform 4:17. https://doi.org/10.1186/1758-2946-4-17
Rappe AK, Casewit CJ, Colwell KS et al (1992) J Am Chem Soc 114:10024–10035. https://doi.org/10.1021/ja00051a040
Maier JA, Martinez C, Kasavajhala K et al (2015) J Chem Theory Comput 11:3696–3713. https://doi.org/10.1021/acs.jctc.5b00255
Hernandez Gonzalez JE, Hernandez Alvarez L, Pascutti PG et al (2017) Proteins 85:1666–1683. https://doi.org/10.1002/prot.25322
Reif MM, Hunenberger PH, Oostenbrink C (2012) J Chem Theory Comput 8:3705–3723. https://doi.org/10.1021/ct300156h
Abraham MJ, Murtola T, Schulz R et al (2015) SoftwareX 1:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Hess B, Bekker H, Berendsen HJ et al (1997) J Comput Chem 18:1463–1472. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12%3c1463::AID-JCC4%3e3.0.CO;2-H
Berendsen HJC, Postma JPM, van Gunsteren WF et al (1984) J Chem Phys 81:3684–3690. https://doi.org/10.1063/1.448118
Bussi G, Donadio D, Parrinello M (2007) J Chem Phys 126:014101. https://doi.org/10.1063/1.2408420
Parrinello M, Rahman A (1981) J Appl Phys 52:7182–7190. https://doi.org/10.1063/1.328693
Aldeghi M, Heifetz A, Bodkin MJ et al (2016) Chem Sci 7:207–218. https://doi.org/10.1039/c5sc02678d
Hernandez Gonzalez JE, Hernandez Alvarez L, Leite VBP et al (2020) J Chem Inf Model 60:5499–5512. https://doi.org/10.1021/acs.jcim.0c00294
Klimovich PV, Shirts MR, Mobley DL (2015) J Comput Aided Mol Des 29:397–411. https://doi.org/10.1007/s10822-015-9840-9
Boresch S, Tettinger F, Leitgeb M et al (2003) J Phys Chem B 107:9535–9551. https://doi.org/10.1021/jp0217839
Case DA, Betz RM, Cerutti DS et al (2016) AMBER 2016. University of California, San Francisco
DeLano WL (2002) PyMOL. 2.1.0. DeLano Scientific, San Carlos
Lerbret A, Mason PE, Venable RM et al (2009) Carbohydr Res 344:2229–2235. https://doi.org/10.1016/j.carres.2009.08.003
Sarduy ES, Munoz AC, Trejo SA et al (2012) Protein Expr Purif 83:59–69. https://doi.org/10.1016/j.pep.2012.03.008
Copeland RA (2013). In: Copeland RA (ed) Evaluation of enzyme inhibitors in drug discovery: a guide for medicinal chemists and pharmacologists. Wiley, Hoboken, pp 123–168
Selwyn MJ (1965) Biochim Biophys Acta 105:193–195. https://doi.org/10.1016/S0926-6593(65)80190-4
One-way ANOVA followed by Dunnett’s multiple comparisons test was performed using GraphPad Prism version 5.03 for Windows, GraphPad Software: San Diego, California USA, www.graphpad.com
(2004) Assay Guidance Manual. Eli Lilly & Company and the National Center for Advancing Translational Sciences, Bethesda
Copeland RA (2000). In: Copeland RA (ed) Enzymes: a practical introduction to structure, mechanism, and data analysis. Wiley, New York, pp 266–304
Machin JM, Kantsadi AL, Vakonakis I (2019) Malar J 18:388. https://doi.org/10.1186/s12936-019-3043-0
Cournia Z, Allen B, Sherman W (2017) J Chem Inf Model 57:2911–2937. https://doi.org/10.1021/acs.jcim.7b00564
Jiang W (2019) J Chem Theory Comput 15:2179–2186. https://doi.org/10.1021/acs.jctc.8b01147
Vajda S, Beglov D, Wakefield AE et al (2018) Curr Opin Chem Biol 44:1–8. https://doi.org/10.1016/j.cbpa.2018.05.003
Nussinov R, Tsai CJ (2013) Cell 153:293–305. https://doi.org/10.1016/j.cell.2013.03.034
Funding
JEHG and CO gratefully acknowledge financial support in the context of an Ernst Mach Scholarship (OeAD), financed by the Austrian Federal Ministry for Education, Science and Research. JEHG and LHA thank the Sao Paulo Research Foundation (FAPESP), Grant Numbers 2016/24587–9, 2018/03911–8 and 2020/10214-1 for financial support. ESS is grateful to the National Research Council (CONICET, Argentina). VBPL was funded by VBPL was funded by FAPESP Grant 2019/22540–3 and 2016/19766–1, and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq). PGP thanks CNPq e Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) for general financial support (Finance Code 001).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hernández González, J.E., Salas-Sarduy, E., Hernández Alvarez, L. et al. In silico identification of noncompetitive inhibitors targeting an uncharacterized allosteric site of falcipain-2. J Comput Aided Mol Des 35, 1067–1079 (2021). https://doi.org/10.1007/s10822-021-00420-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-021-00420-7