Abstract
There is a pressing need to improve the efficiency of drug development, and nowhere is that need more clear than in the case of neglected diseases like malaria. The peculiarities of pyrimidine metabolism in Plasmodium species make inhibition of dihydroorotate dehydrogenase (DHODH) an attractive target for antimalarial drug design. By applying a pair of complementary quantitative structure–activity relationships derived for inhibition of a truncated, soluble form of the enzyme from Plasmodium falciparum (s-PfDHODH) to data from a large-scale phenotypic screen against cultured parasites, we were able to identify a class of antimalarial leads that inhibit the enzyme and abolish parasite growth in blood culture. Novel analogs extending that class were designed and synthesized with a goal of improving potency as well as the general pharmacokinetic and toxicological profiles. Their synthesis also represented an opportunity to prospectively validate our in silico property predictions. The seven analogs synthesized exhibited physicochemical properties in good agreement with prediction, and five of them were more active against P. falciparum growing in blood culture than any of the compounds in the published lead series. The particular analogs prepared did not inhibit s-PfDHODH in vitro, but advanced biological assays indicated that other examples from the class did inhibit intact PfDHODH bound to the mitochondrial membrane. The new analogs, however, killed the parasites by acting through some other, unidentified mechanism 24–48 h before PfDHODH inhibition would be expected to do so.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There were approximately 228 million malaria cases in 2018, with 405,000 deaths attributed to malaria [1]. Infection by Plasmodium falciparum and P. vivax are the most common causes of the disease in humans, with the former being responsible for the greatest mortality. Growing resistance to currently available antimalarial drugs makes identification of new compounds with novel modes of action and activity against resistant parasites a matter of great urgency [2,3,4].
Finding new molecular targets is one way to reduce the risk of cross-resistance developing after introduction of a new antimalarial. Fortunately, biochemical idiosyncrasies of the parasite provide several distinct targets that reduce the risk of undesirable “off-target” effects. Unfortunately, the expected return on investment for “neglected diseases” like malaria is too low to motivate large-scale commercial development of compounds directed at those disease targets. A few pharmaceutical companies have made substantial contributions to public-sector drug discovery efforts (particularly in the form of screening results), but most work in the area has been done by academic groups and nonprofit organizations.
Funding constraints and limited synthesis resources make in silico and collaborative approaches particularly attractive for such noncommercial applications. Computational methods being employed range from constructing quantitative models of the relationship of molecular structure to the specific biological activity of interest (QSARs) or to more general molecular properties related to in vivo absorption, distribution, metabolism, excretion, and toxicity (ADMET QSPRs). Physiologically based pharmacokinetic (PBPK) simulations are also potentially useful, in that they are able to combine individual system-specific properties with drug-specific information in order to anticipate how a compound will behave in vivo. Such simulations are needed to take into account complex interactions between physicochemical properties—e.g., lipophilicity, solubility, and permeability. The goal here is to kill the parasites, of course, which makes such modeling applications somewhat different from those in which one is trying to adjust some physiological imbalance. The situation is similar to oncology, though the adversary in the case of malaria is—fortunately—more consistent in its presentation. We were targeting the blood-stage parasite, so the goal was to get compounds to red blood cells in the systemic circulation and keep them there; distribution to peripheral tissues was not particularly desirable.
Efforts to use virtual screening to identify new antimalarials have had limited success. Zhang et al. [5], for example, built QSAR models using training data from a 3,133-compound library that contained 158 confirmed actives. The models obtained were used to screen a commercial database (ChemBridge, San Diego, CA) in order to identify new potential leads. From this virtual screen, 176 compounds representing 22 unique scaffolds were identified and tested for growth inhibition. Of these, 7 were active against P. falciparum chloroquine-susceptible (3D7) and chloroquine-resistant (K1) strains ex vivo with a concentration required to kill half of the organisms (XC50) of 1 μM or less [5].
Here, we used QSAR models and PBPK simulations to pick an attractive lead series from “hits” in a phenotypic assay, then extended that series by generating a virtual library of novel analogs. An array of predicted properties—PfDHODH inhibition, physiochemical properties, and pharmacokinetic profiles—were used to select a handful of candidates for further evaluation. The full range of predicted properties taken into consideration are spelled out in Supplementary Data Table S2. The ADMET Risks [6] they highlight can reduce oral bioavailability or pose toxicity problems, which makes them relevant in any drug design program. PBPK simulations using predicted ADMET properties as input parameters were carried out to determine whether virtual compounds were likely to have good enough oral absorption and slow enough systemic clearance to sustain a blood concentrations high enough to kill the parasite given a dosing regimen acceptable for existing antimalarial drugs (vide infra). Surviving virtual compounds were synthesized and characterized experimentally.
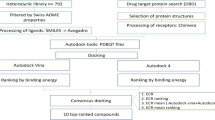
The methodology described herein focuses on the use of in silico tools for identifying novel lead compounds for synthesis in a way that exploits QSAR and PBPK modeling to minimize the bench resources needed. The workflow can be characterized as follows:
-
1.
Use enzymatic screening data to generate a predictive QSAR model for an attractive target.
-
2.
Apply the QSAR obtained to a large and diverse library of structures and associated phenotypic activity to identify a lead series predicted to exhibit good ADMET properties in aggregate.
-
3.
Create unique combinations of active substituents from discovered scaffolds.
-
4.
Predict activity, ADMET properties, and PK profiles based on QSAR and PBPK models.
-
5.
Choose and synthesize attractive candidates from the analogs identified.
-
6.
Measure biological activities and ADMET properties of the synthesized compounds.
-
7.
Iterate as needed.
The work described here covers a single iteration of this workflow that provided us with an opportunity to assess the ability of our models to make good activity and ADMET property predictions for novel compounds. It also represents a proof-of-concept exercise of our lead discovery and optimization tools and analytical methods developed over the past several years. Our hope was that we might, in the process, also contribute something of value to the search for novel antimalarials.
As a proof-of-concept project, the scale of our work was necessarily limited. The scope was nonetheless broad and deep enough that we encountered several challenges typical of larger drug development projects. Those challenges included needing to revise our original design to incorporate more practical synthesis schemes, to revise synthesis targets to work around failed reactions, and to encountering an off-target mode of action. Fortunately, in our case the off-target activity was one that results in more rapid death of the parasite than inhibiting our intended target does.
Lead selection and compound design
Plasmodium species cannot salvage preformed pyrimidine bases for nucleic acid synthesis as its human hosts can. Dihydroorotate dehydrogenase (DHODH) is a critical enzyme in the de novo pyrimidine synthesis pathway in the parasite and, thus, a potential target for antimalarial drug therapies [7, 8]. The enzyme from Plasmodium species is located in the mitochondrion and utilizes ubiquinone (also known as coenzyme Q) as a cosubstrate.
Predicting DHODH inhibition
Data on PfDHODH inhibition were collected from the literature for a structurally diverse group of compounds. These included triazolopyrimidines [7, 9], diphenylureas [10], and 2-cyano-3-hydroxy acrylanilides [11], among others [12,13,14,15]. Quantitative data were not available for inhibition of the intact enzyme, which is bound to the mitochondrial membrane by its hydrophobic N-terminus in vivo [16]. A total of 89 of those compounds had been assayed against the soluble recombinant form of PfDHODH (s-PfDHODH) from which the hydrophobic “tail” has been removed. The very low solubility of CoQ-10 limits its utility in vitro, so an artificial ubiquinone (dodecylubiquinone [DQ]) is usually used as substrate instead.
The IC50 data for these 89 compounds were converted to inhibition constants (Ki’s) by assuming competitive inhibition against DQ and applying the method described by Cheng and Prusoff [17]. Two artificial neural network ensemble (ANNE) regression models were constructed (henceforth referred to as Model A and Model B) using ADMET Modeler™ module of ADMET Predictor® [18]. The models were built on distinct 20% hold-out test sets (see Supplementary Methods for details). That and the use of different random number seeds led to somewhat different descriptor sets, complexities and performance statistics (Table 1). The two models complemented each other well, with Model A yielding more conservative activity estimates (fewer false positives) and model B being more sensitive (more prone to generating false negatives). Hence, using them together to identify potent compounds worked better than using either model alone. Plotting the observed versus predicted values from the models illustrated the ability of both models to adequately predict the Ki values from both the training set that was used to build the models and the test set used to validate the models (Fig. 1).
Performance plots for the ANNE regression models developed for in vitro PfDHODH inhibition built from literature data. The plot for Model A is on the left and the plot for Model B is on the right. Labeled points correspond to data from Supplementary Table S1: (a) Tz18; (b) P05; (c) G47; (d) D10; and (e) Tz11
Selecting a lead series
The developed QSAR models were used to analyze structures for 13,533 antimalarials identified in a phenotypic screen of nearly 2 million compounds that was carried out by GlaxoSmithKline (GSK). The published data included P. falciparum growth inhibition for the chloroquine-sensitive 3D7 (PubChem AID 2306) as well as for the multidrug resistant strain Dd2 (PubChem AID 2302). Data from a counter screen for mammalian cytotoxicity was available from PubChem AID 2303 [19].
There were some important limitations in working with this data set. First, evaluating efficacy against parasites cultured in human erythrocytes provides no direct information on a compound’s mode of action. Secondly, only results for the active compounds (those that inhibited parasite growth by at least 80% at a concentration of 2 μM) were reported. The lack of data on inactives posed a considerable risk of synthesis resources being wasted on seemingly “novel” analogs that in fact had already been synthesized but not reported due to their lack of activity. Lastly, many of the compounds in the screening data set exhibited undesirable physical properties and/or undesirable substructures (e.g., multiple halogenated thiophene rings) that would cause them to be avoided by many pharmaceutical companies [20].
Candidate lead series from the active compounds in the phenotypic screening data set were identified and structural classes were generated in MedChem Studio™ [21] based on shared substructures. Unlike most other approaches [22], the method used in MedChem Studio does not cluster compounds (in this case, actives) into exclusive subsets based on the degree of pairwise overlap in their substructural fingerprints. Rather, it generates nonexclusive classes of molecules that share a maximal common substructure. In most cases, this shared substructure is, itself, a scaffold in the medicinal chemistry sense. If not, it can readily be modified to become one. The nonexclusivity is advantageous for lead series identification because it is typically easier to see cases where overlapping classes can profitably be combined by merging the scaffolds that define them.
Classes were evaluated as potential lead series based on the QSAR-predicted inhibitory potencies against PfDHODH and the degree of parasite growth inhibition observed in the phenotypic screen for chloroquine-sensitive P. falciparum 3D7 (PubChem AID 2306). ADMET Risk™, which indicates how many of an array of ADMET property rules are violated by a compound [6], was also taken into consideration. The rule thresholds are calibrated against a subset of the World Drug Index (WDI) that is enriched in orally delivered commercial drugs similar to that used by Lipinski, et al., for anticipating solubility and oral absorption problems [23]. Ninety percent of the WDI reference set have an ADMET Risk score less than seven.
Low group averages were preferred for both predicted Ki and ADMET Risk, but the spread in each profile was also important. Lack of variation in potency indicates a “lack of SAR”, which makes it difficult to enhance activity against the target by modifying structure. Similarly, a lack of variation in ADMET Risk suggests that it will be difficult to engineer out potential liabilities by changing structures. The exception would be a class wherein all ADMET Risk scores are very low, and that was never the case in this dataset.
Results for three classes at an intermediate stage of the series definition process are shown in Fig. 2. Two of the classes—diphenyl ureas (DPUs) and triazolopyrimidines (Tzs)—represent PfDHODH inhibitor lead series that had already been explored or were being actively explored by other research groups [24, 27].
Comparing property distributions across three representative classes: triazolopyrimidines (Tzs), aminopropylaminoquinolones (APAQs) and diphenylureas (DPUs). The portion of each representative structure highlighted in blue corresponds to the class scaffold. Growth inhibition is shown as %inhibition vs. P. falciparum strains 3D7 and DD2, which are chloroquine-sensitive and -resistant, respectively. pKi_pred values are the negative log of the predicted Ki in µM, so a larger number indicates higher potency; their distribution for Models A and B are shown. ADMET Risk is a measure of likely development liabilities that can range from 0 to 24 [6]
The 2-(3-aminopropylamino)-4-quinolone (APAQ) class was particularly promising because its scaffold bore a distinct resemblance to ubiquinone, which is a cosubstrate for PfDHODH in vivo (Fig. 3). They also resembled N-hydroxy-2-dodecyl-4-quinolone (HDQ) (Fig. 3) which Dong et al. had determined to have good antimalarial activity, with PfDHODH as its primary target [25]. Although they reported that HDQ inhibited parasite growth in culture and were able to show that PfDHODH was the site of action, they did not have access to soluble enzyme. Hence the XC50 obtained in culture could not be translated into an in vitro Ki for the enzyme. HDQ was therefore not included in the data set used to build the PfDHODH inhibition models. Two quinolones found among the “hits” reported by Patel et al. [8] were initially grouped with some of the APAQs. These lack the defining 2-amino substitution and so did not appear in the final lead series, which was comprised of 113 actives.
None of the APAQs were reported to be cytotoxic to the mammalian HepG2 cells used in the companion AID 2303 screen, which suggests that the intrinsic risk of mammalian toxicity is small. That said, many of them had discouragingly high ADMET Risk scores. In particular, the two most active examples (Fig. 3) violated 7 and 9 of the standard ADMET Risk rules, respectively. Both are large and were predicted to be excessively hydrophobic, to be highly bound in plasma, to exhibit low water solubility, and to be acutely toxic to rats. In addition, CID 44535189 has an excessively large number of rotatable bonds and was predicted to exhibit chronic toxicity in mice. Besides the five likely liabilities shared with CID 44535189, CID 44534046 was predicted to be metabolized by CYP2D6 and CYP3A4, to inhibit CYP3A4, and to bind to the human ether-a-go-go (hERG) gene product.
Analog generation and selection
The predictions of high lipophilicity, low water solubility, high susceptibility to metabolism by cytochrome P450s (especially CYP2D6), and hepatotoxicity were common among the actives in the APAQ class. Substituents from the less risky APAQs were combinatorially reshuffled in silico to generate lead candidates with more favorable activity and property profiles. Substituents from each R-group were drawn from the known actives in the class. Doing so increases the likelihood that the structures produced will be synthetically accessible, since at least one example already exists where that substituent was placed at that position. In addition: only single substitutions were allowed at the distal nitrogen; only simple substituents (hydrogen, halogen, methyl, or trifluoromethyl) were allowed on the quinolone ring; and substitution was restricted on the distal aromatic rings. The resulting constrained combinatorial R-group explosion produced a virtual library of ~ 99,000 novel analogs.
Attractive lead analogs from the virtual library were selected based on their predicted PfDHODH inhibitory power in both QSAR models, their predicted ADMET properties, and their expected ease of synthesis. Compounds for which the predicted PfDHODH Ki for Model A was greater than 0.1 µM and with ADMET Risk > 4 were set aside, as were compounds that were out-of-scopeFootnote 1 with respect to Model A. Though Model B was useful for identifying good lead series, it failed to provide much further discrimination between APAQ analogs, hence whether its prediction for a particular molecule was out-of-scope or not was disregarded.
As is often the case, descriptors for some of the compounds ultimately selected for synthesis based on activity predictions fell well outside the range of descriptors for compounds used to train one or more of the metabolism models available at the time. Predictions for such compounds were considered “out-of-scope” extrapolations and, therefore, not necessarily reliable. Compounds with in-scope predictions were favored in selecting candidates for further analysis but ignoring those for which any prediction was out-of-scope was impractical. Such limitations in coverage are common for any novel chemistry, but they are also a major motivation for continuously refining and expanding ADMET models to improve their predictive performance.
Once analogs predicted to have relatively low potency or discouraging ADMET properties had been filtered out, approximately 34,001 analogs still remained. Of those, 17,334 bore symmetrical central diamine moieties, a feature expected to greatly simplify synthesis. ADMET Predictor now includes a synthetic difficulty score analogous to that described to the synthetic accessibility score described by Ertl and Schuffenhauer [26]. That functionality was not available when this project began, however, so assessment was carried out manually by visual inspection of analogs displayed in a tile view, i.e., in a gridded rather than a row format. In retrospect, the manual process was reasonably effective. The synthetic difficulty scores for the symmetrical diamine analogs ranged from 1.95 to 5.0 (median 3.62, where 10 is most difficult) and the final dozen candidates (vide infra) ranged from 2.5 to 4.15 (median 3.35), indicating that the final candidates were fairly representative of the initial pool in terms of synthetic accessibility.
Marginal aqueous solubility was a central concern that had to be balanced against model predictions that substitution at the central carbon of a 1,3-diaminopropyl bridge should improve activity and reduce CYP metabolism. This and other ADMET Risks prompted us to manually expand the library by introducing small point changes to some marginal analogs. One such change was “mutating” a gem-cyclohexyl ring at the central carbon of the aminopropylamino bridge to a gem-cyclopentyl ring. No such compound existed among the GSK actives, though there were several derived from trans 1-amino-2-aminomethylcyclopentane [19, 27]. A 1,2-ring is expected to adopt a quite different 3D conformation from that for a 2,2-ring, thereby placing the terminal group at quite different positions in space.
After the initial set of lead compounds were selected, their expected pharmacokinetic profiles were evaluated by applying the in silico predictions and standard human physiologies in GastroPlus® [28] to predict plasma concentration-versus-time profiles. Property estimates were taken from ADMET Predictor and dosing regimens were similar to those used for existing antimalarial drugs, e.g., chloroquine. Target plasma concentrations were based on predicted Ki values.
An initial set of diverse synthesis candidates (Supplementary Table S2) was selected from among those predicted to have good potency as well as an acceptable ADMET Risk score and favorable simulated in vivo PK profiles. The twelve synthesis targets produced by this process had structural variations at three different positions: the R1 substituent on the quinolone end of the scaffold; the R2 and R3 substituents off of the distal nitrogen of the aminopropylamino bridge; and the R4 and R5 substituents off of the central carbon in the bridge. The beginning of the reduction-to-practice phase was announced in a press release in September 2011 [29] and bids were solicited from several contract synthesis companies. Four of the targets distributed are shown in Fig. 4.
The individual compounds proposed were synthetically feasible, but the number of separate intermediates and variations in reaction conditions required would have been prohibitively expensive. Instead, variations were restricted to either end of the molecule and we focused on a single kind of aminopropylamino bridge. Doing so made it likely that similar reaction conditions and intermediates could be used to make multiple analogs. The convergent synthesis plan greatly reduced the potential for complications and budget overruns. Though not often dwelt on in the literature, such considerations are often a critical practical consideration for any molecular design project; hence their explicit inclusion here. In the end, we settled on seven analogs built around a simplified scaffold bearing the novel gem-cyclopentyl group at the central carbon of the bridge. Diversity was provided by placing three different substituents on the distal nitrogen (i.e., R2) (Fig. 5).
Kalexsyn, Inc. (Kalamazoo, MI) synthesized a revised list of seven target analogs. Carbonyl precursors suitable for reductive amination were available or accessible for some of the desired R2 substituents, considerably simplifying their synthesis.
Experimental results and discussion
The candidates chosen under the plan outlined above struck a reasonable balance between structural diversity and being accessible via reasonably convergent and parallel synthetic methodologies. Despite our hopes and best intentions, however, the particular substituents targeted for the distal nitrogen still needed some tweaking as regards how the coupling was carried out. Similarly, although the 6-methoxy analog moiety is very similar in structure to that in the 6-chloro and unsubstituted quinolones, its synthesis required a distinct approach. Fortunately, both approaches were relatively straightforward. What is provided here is an overview for illustrative purposes. That said, it is complete enough to give a sense of how much effort was involved in generating the seven targeted analogs. More details are provided in the Supplementary Materials.
Synthesis
The general synthesis procedure used is outlined in Scheme 1. gem-Dicyanocyclopentane was prepared by condensation of malononitrile with dibromobutane and reduced to the diamine 5 using lithium hydride in tetrahydrofuran. Subsequent reaction with 2-chloro-4-methoxyquinolines yielded 7a–c after hydrolysis in HCl.
2-Chloro-4,6-dimethoxyquinoline (6c) was prepared by condensation of malonic acid with p-anisidine, yielding 2,4-dichloro-6-methoxy quinoline [30], which was converted to the desired dimethoxy compound by treatment with methoxide in methanol (Scheme 2).
Reductive alkylation of the distal amino group in 7a with thiophene carbaldehyde to yield 8 proceeded readily with cyanoborohydride in methanol at room temperature, whereas preliminary formation of the imine and heating were required to generate the tetrahydrocarbazole analog 9 and the tetrahydroquinolines 10a–c from the corresponding ketones (Scheme 3).
Protection of the indole nitrogen in the former case was not necessary. In fact, difficulties with deprotecting the product made it counterproductive. Blocking the amino group in the tetrahydroquinolone, on the other hand, was essential. Though reductive amination with the free amine failed, the t-butyloxycarbonyl(Boc)-protected dihydroquinolone reacted cleanly. Deprotection with ZnBr2 in dichloromethane (DCM) or transesterification in formic acid followed by hydrolysis in NaOH afforded the targeted analogs 11a–c (Scheme 3).
Analogs bearing an unsubstituted imidazolinone ring in place of the dimethylthiophene ring in 8 were attractive in terms of predicted activity and ADMET properties. Unfortunately, several attempts to prepare and purify them failed. Some N-protected intermediates were generated, but all broke down during deprotection or during purification. The 1,3-dimethylimidazolinones 12a and 12b were synthesized in their stead (Scheme 4). The added methyl groups were predicted to reduce solubility somewhat, but absorption was expected to be enhanced. Moreover, the methyl groups were predicted to be cleaved off in the liver to produce the original targets. This change in targets necessitated the use of N,N’-dimethylimidazolinone-4-carboxaldehyde, which was prepared from ethylene glycol and N,N’-dimethyl urea, then formylated with POCl3 in dimethylformamide. Condensation with the respective free amino precursor yielded the desired APAQs (Scheme 4).
ADMET properties: predicted vs. experimental
To become a drug candidate, a lead molecule must possess many favorable pharmacological properties, including good solubility, metabolic stability, and low toxicity as well as activity. Hence, ADMET QSARs and QSPRs provide a valuable complement to activity models in avoiding unfavorable ADMET profiles, provided the predictions they provide are accurate and reliable. Measured physicochemical properties of the most active synthesized compounds were determined experimentally and are compared with the predictions from ADMET Predictor 9.0 in Table 2. Most of the observed and predicted properties have a root mean squared error (RMSE) of 0.8 log units or less. Aqueous solubility was an exception, but not an unexpected one. That is because the compounds were all isolated as foams that subsequently solidified to glassy solids, the solubilities of which can be quite difficult to measure accurately.
Rates of in vitro clearance determined from a panel of recombinant microsomes containing human CYPs were measured, as was that by human liver microsomes (HLMs). The results obtained are shown in Table 3. The overall performance of the models in ADMET Predictor is good, but interpretation is more complicated than for the physicochemical properties. These are not predictions of simple intrinsic clearance, as several of the relevant concentrations at half-maximal metabolic velocity (Km’s) are predicted to be low enough for partial saturation to be an issue (Supplementary Table S3). This underscores the desirability of having affinity models (here, for Km) available to put data on microsomal stability obtained at a single concentration into proper context.
The results for CYP2C9 and CYP2C19 in Table 3 also illustrate why it is important to make regression models for enzyme activity contingent on classification models that distinguish substrates from nonsubstrates [6, 31]. Nonsubstrates are necessarily absent from the data used to construct the regression models, and the classification models confidently identify them [32] as being out-of-scope for the corresponding regression models. The clearance predictions provided represent the values expected if they were substrates, which the classification models confidently predict they are not.
The situation is somewhat different for CYP1A2. Here the substrate classification predictions are uncertain—the confidences are near toss-up (41–60%), but they are still consistent with the low to nonexistent experimental clearance values for this isoform.
CYP kinetics are complicated by the potential for autoinhibition [33]. The APAQs were not tested as inhibitors, but all were predicted to inhibit CYP2D6. That may explain the consistent overestimation of CYP2D6 clearance for CYP2D6, for which the predicted affinities are high—i.e., for which the Km’s are all less than 1 μM (Supplementary Table S3). All four analogs are also predicted to be CYP3A4 inhibitors, but the corresponding Km’s are 15 μM or above, making autoinhibition unlikely to be a problem at 1 μM.
Pharmacokinetic predictions
The intrinsic clearances for the most active APAQs would have posed a risk high enough to discourage further development of these compounds had they not exhibited acceptable pharmacokinetic profiles using the other property predictions. In fact, high predicted microsomal clearance was the reason several otherwise attractive candidates that failed to yield acceptable pharmacokinetic profiles were set aside during analog selection. The result for compound 9 is shown in Fig. 6. Here the predicted fraction unbound in plasma (fup < 3%) and high volume of distribution (Vd; estimated at 7 L/kg) offset the relatively high predicted intrinsic clearance. Basically, too little of the compound is likely to get into the liver for hepatic metabolism to be a problem. As a result, the bioavailability is predicted to be high enough to produce a favorable in vivo profile, given an appropriate dosing schedule—here, a schedule similar to that currently used for chloroquine.
Human concentration–time profile expected for 9 based on PBPK simulation using GastroPlus. Pharmacokinetic parameters were taken from experimental values where available and estimated using the QSAR models in ADMET Predictor otherwise. Conc concentration, Ki predicted inhibition constant for PfDHODH when plasma protein binding is taken into account
Both fup and Vd are in fact flagged as ADMET Risks by default, because they can make it harder for a drug to get from the plasma into peripheral tissues. That is not relevant to the blood phase of the Plasmodium life cycle, but would affect the ability of the compound to attack liver stages. The fact that they can offset potential first-pass metabolic liabilities is one good reason to examine pharmacokinetic simulation results for compounds of interest that raise piece-wise risk flags [31].
The QSAR predictions of Vd and fup for 9 are similar to the values predicted for chloroquine (10.6 L/kgFootnote 2 and 9%, respectively), which is widely used to treat malaria, and thus are not unrealistic pharmacokinetic parameters.
Biological results
All aminoquinolones synthesized for this project were assayed for ex vivo antimalarial activity in blood culture and for inhibition of s-PfDHODH. Characterization in terms of kill rate is very resource-intensive, as is determining whether or not parasites can be rescued from growth inhibition by transfection with Saccharamyces cerevisiae DHODH (ScDHODH) DNA (see the section below on “Inhibition of intact PfDHODH”). These assays were, therefore, only carried out for the two most active candidates for which sufficient material was available—8 and 11b. In addition, historical data for some of the APAQs in the PubChem dataset, including ScDHODH rescue and ex vivo activity, were made available by the Tres Cantos Medicines Development Campus-Diseases of the Developing World, GSK (Tres Cantos) [19].
XC50s for cultured parasites
Antimalarial activity in blood culture was determined as previously described [35, 36] at the University of California, Riverside, under conditions essentially identical to those used by GSK [19]. Table 4 shows the experimental results obtained in asynchronous cultures, expressed as the concentration required to reduce growth by 50% (XC50). With the exception of the imidazolinones 12a and 12b, the new compounds were more potent than the most active APAQs in the GSK data set.
With the exception of 12a, the novel analogs met the minimum potency threshold (better than 2 μM) for parasite proliferation called for in the Medicines for Malaria Venture (MMV) guidelines at the time [37], with 11b satisfying the ideal criteria with an XC50 below 100 nM. Similar antimalarial potencies were obtained in replicate assays carried out at UC Riverside and at Tres Cantos. The compounds were also tested in the chloroquine-resistant Dd2 strain, which contains an efflux pump [38]. It was gratifying to see that the ratio of XC50′s between the resistant and susceptible strains—i.e., the resistance ratio (Table 4)—was lower for all of the APAQs than the value of 15.5 reported for chloroquine [39].
Speed of killing time course assay
The parasite reduction ratio (PRR) is an indicator of how rapidly parasites are killed by antimalarial treatment [40]. Preliminary assays carried out at Tres Cantos for 8 and 11b yielded 48-h parasite in vitro reduction ratios (PPRs) of approximately 3 log units, i.e., 99.9% of the parasites were killed in the first 48 h of exposure (1 lifecycle). This was slightly better than pyrimethamine, which was included as a positive control. Such rapid killing of the parasite is very desirable for antimalarials [41]. Little or no lag time in the onset of killing was evident for either compound, compared to the 24–48 h time lag reported for triazolopyrimidine inhibitors of PfDHODH [42].
Inhibition of s-PfDHODH in vitro
Identification of a compound’s mode of action is important in drug discovery because it opens the door to structure-based drug design and because it helps alert researchers to undesirable off-target activities that might cause concern. In some cases, it highlights opportunities for synergistic inhibition of multiple targets by a single compound, i.e., for polypharmacology [43]. PfDHODH was the intended target for this proof-of-principle project, so we wanted to know whether the identified lead series (in general) and our new analogs (in particular) inhibited the enzyme in intact parasites.
The literature data set used to build our PfDHODH inhibition models was based on in vitro assays carried out using the truncated, soluble form of the enzyme introduced by Baldwin et al. [10]. Patel et al. subsequently used it to screen a library of 208,000 compounds and validated the 38 “hits” obtained by testing for inhibition of parasite growth in blood culture [8] The seven compounds we designed and synthesized were poor inhibitors of s-PfDHODH with in vitro IC50 > 100 μM when assayed under those conditions at the Tres Cantos laboratory.
This result was somewhat unexpected, because of the active quinolones in the QSAR model training set (compounds P01 and P02 in Supplemental Table S1) and the potency of HDQ mentioned above. The substituents at the C2 position in the APAQs are longer and bulkier than are the styryl and pentyl groups in P01 and P02, but they are still much smaller than the 30- to 50-carbon polyisoprenoid tail of the natural substrate, CoQ10.
The solubility of s-PfDHODH results from the removal of the hydrophobic tail that anchors the enzyme to the mitochondrial membrane, where its native cosubstrate resides in vivo. A compound that inhibits the truncated, soluble enzyme in vitro is likely to inhibit the full-length, membrane-bound form. The converse—that an inhibitor of the full-length enzyme will necessarily inhibit the truncated one—is not necessarily true, especially for inhibitors like the APAQs that resemble ubiquinone more than they resemble dihydroorotate [16]. Atovaquone, for example, inhibits the full-length enzyme from P. falciparum with a Ki of 27 µM after solubilization in Triton X-100 [44], whereas the corresponding value for the truncated enzyme is reportedly > 500 µM when assayed under similar conditions [45]. Binding to the s-RrDHODH from rat is much tighter (Ki ~ 0.08 µM) and intermediate for the corresponding human enzyme ((Ki ~ 2 µM). The organization of the N-terminal “stump” where competitive CoQ inhibitors bind differs substantially from species to species and changes upon inhibitor binding [45]. These observations suggest that the absence of inhibition by APAQs in the in vitro assay could be due to the loss of the hydrophobic N-terminal residues and may not reflect the activity in living parasites.
Inhibition of intact PfDHODH
Fortunately, there is a way to address this question, though the method is somewhat indirect. P. falciparum parasites engineered to express DHODH from the yeast Saccharamyces cerevisiae (ScDHODH) use fumarate as an oxidizing agent instead of ubiquinone, so their growth is not inhibited by (i.e., they are “rescued from”) inhibitors that compete with ubiquinone. Painter et al. used this technique to validate PfDHODH as an antimalarial drug target [46].
Unbeknownst to the Simulations Plus and U.C. Riverside groups, the Tres Cantos group had already tested several 2-aminoquinolone actives from the GSK antimalarial data set for rescue by ScDHODH transfection. Their previously unpublished work showed that parasites were rescued from growth inhibition by 10 of 102 APAQs assayed, with some additional compounds giving an equivocal result, i.e., partial rescue. An additional sixteen 2-amino-4-quinolones having different bridging groups were also assayed; three of these were clearly subject to rescue, whereas three yielded ambiguous results. Illustrative example structures are shown in Fig. 7.
The APAQs from which parasites were rescued by ScDHODH transfection did inhibit s-PfDHODH in vitro, but with higher in vitro IC50′s (i.e., with lower affinity) than expected given the phenotypic assay results—3.1 and 5.2 μM for the most potent examples. The most potent GSK analog (CID 44,537,350) had an in vitro IC50 of 0.86 μM. It bore a 4-(2-pyridyl)butylamino group in place of the 3-aminopropyl amino group in our series.
The limitation of this test is that the failure of transfection to rescue parasites from the other APAQs tested—including 8 and 11b—does not imply that those compounds do not inhibit the intact Plasmodium enzyme. It does imply, however, that PfDHODH inhibition is not their exclusive mode of antimalarial action. Basically, 8 and 11b start killing the parasite in culture immediately (see above), whereas specific PfDHODH inhibitors take 24–48 h to do so [47]. A somewhat analogous situation exists for atovaquone, which is much more potent in blocking respiratory oxidation of ubiquinol than in inhibiting PfDHODH directly [46]. In that case, the time frame and the end result—the parasite is starved of the CoQ it needs to synthesize pyrimidine nucleotides—are the same for both targets,only the target itself is different.
The bottom line is that APAQs as a class evidently do inhibit PfDHODH ex vivo but that, at least for our most potent analogs, some other, more rapid-onset mode of action is responsible for their antimalarial activity.
Other suggested modes of action
The malarial methionyl t-RNA synthetase (PfMRS) has also been suggested as a target. A patent for 2-aminoquinolones as antibiotics was filed in 1999 [27, 48] and the mode of action was characterized as being inhibition of bacterial MRS. However, the 26 compounds characterized as MRS inhibitors in that patent were not among the actives in the Gamo et al. data set [19], which argues against the idea that MRS might be the target in malaria. Subsequent to the work described here, two other APAQs characterized as bacterial MRS inhibitors (REP3123 and REP8839 in Fig. 8) were shown to inhibit the malarial enzyme in vitro and to be potent antimalarials ex vivo [49].
Those authors did not test for activity against PfDHODH, however, nor did they present conclusive evidence that blocking PfMRS was the compounds’ mode of action in the intact parasite.
In fact, the rapid knockdown and absence of a lag time in the PRR experiments argues against any kind of protein synthesis inhibition being involved, at least for 8 and 11b [50].
Conclusions
The 2-(3-aminopropylamino)-4-quinolones generated by the model-driven in silico design approach described here are potent antimalarial lead drug candidates. The methodology used to design compounds that target PfDHODH yielded lead candidates with excellent activity against P. falciparum parasites as well as acceptable ADMET properties and favorable PK profiles. Compounds from the first design iteration were predicted to be more potent in blood culture than structurally related literature compounds and their experimentally determined potencies were consistent with that expectation. It seems likely that further iterations and in vivo characterization would be productive.
The results presented here do not pin down the primary antimalarial mode of action for the APAQs, but do indicate that it is probably neither PfDHODH nor protein synthesis. Fortunately, the lack of cytotoxicity the lead series showed against mammalian cells in culture suggests that inhibition of the primary target, whatever it is, is not intrinsically problematic with respect to human beings. If that is indeed the case, having multiple activities should improve prospects for further development iterations centered around this class of chemistry, since hitting multiple targets is a good way to slow development of resistance.
This successful proof-of-principle study highlights how useful in silico tools can be in de novo design and for identifying lead compounds from phenotypic screens. The example chosen illustrates how including predictive PBPK simulations early in the workflow can reduce the risk of relying too heavily on any single ADMET property filter when selecting analogs for synthesis. Applied more generally, the overall approach has the potential to reduce the number of compounds that need to be synthesized to get from discovery through optimization, to make animal testing more efficient, and to reduce attrition in clinical trials. Beyond being useful as a proof-of-principle, the exercise provided prospective validation of several kinds of in silico property predictions as well as an example of the kind of unexpected off-target activity commonly encountered in drug development projects—in this case, a desirable one.
Data availability
Our supply of synthesized material is exhausted at present, but the experimental procedures required to synthesize more are provided. NMR spectra and any details not included in the supplementary materials are available from the authors upon request.
Code availability
ADMET Predictor® and GastroPlus® are commercially available from Simulations Plus, Inc., Lancaster CA 93534 (https://www.simulations-plus.com).
Notes
A prediction was flagged as “out of scope” when any of its descriptor values fell more than 10% outside the range of the corresponding values seen for the data set upon which the model was trained.
The actual Vd for chloroquine is difficult to determine experimentally, but is probably much higher [34].
References
World Health Organization (2019) World malaria report 2019. https://www.who.int/malaria/publications/world-malaria-report-2019/report/en/. Accessed 7 Apr 2020
Wells TNC, van Huijsduijnen RH, Van Voorhis WC (2015) Malaria medicines: a glass half full? Nat Rev Drug Discov 14:424–442
Cui L, Mharakurwa S, Ndiaye D, Rathod PK, Rosenthal PJ (2015) Antimalarial drug resistance: literature review and activities and findings of the ICEMR network. Am J Trop Med Hyg 93(3 Suppl):57–68
Bhattacharjee AK, Geyer JA, Woodard CL, Kathcart AK, Nichols DA, Prigge ST, Li Z, Mott BT, Waters NC (2004) A three-dimensional in silico pharmacophore model for inhibition of Plasmodium falciparum cyclin-dependent kinases and discovery of different classes of novel Pfmrk specific inhibitors. J Med Chem 47:5418–5426
Zhang L, Fourches D, Sedykh A, Zhu H, Golbraikh A, Ekins S, Clark J, Connelly MC, Sigal M, Hodges D, Guiguemde A (2013) Discovery of novel antimalarial compounds enabled by QSAR-based virtual screening. J Chem Inf Model 53:475–492
Lawless MS, Waldman M, Fraczkiewicz R, Clark RD (2016) Using cheminformatics in drug discovery. In: Nielsch U, Fuhrmann U, Jaroch S (eds) Handbook of experimental pharmacology: new approaches to drug discovery. Springer, Switzerland, pp 139–168
Phillips MA, Rathod PK (2010) Plasmodium dihydroorotate dehydrogenase: a promising target for novel anti-malarial chemotherapy. Infect Disord Drug Targets 10:226–239
Patel V, Booker M, Kramer M, Ross L, Celatka CA, Kennedy LM, Dvorin JD, Duraisingh MT, Sliz P, Wirth DF, Clardy J (2008) Identification and characterization of small molecule inhibitors of Plasmodium falciparum dihydroorotate dehydrogenase. J Biol Chem 283:35078–35085
Phillips MA, Gujjar R, Malmquist NA, White J, El Mazouni F, Baldwin J, Rathod PK (2008) Triazolopyrimidine-based dihydroorotate dehydrogenase inhibitors with potent and selective activity against the malaria parasite Plasmodium falciparum. J Med Chem 51:3649–3653
Baldwin J, Michnoff CH, Malmquist NA, White J, Roth MG, Rathod PK, Phillips MA (2005) High-throughput screening for potent and selective inhibitors of Plasmodium falciparum dihydroorotate dehydrogenase. J Biol Chem 280:21847–21853
Davies M, Heikkila T, McConkey GA, Fishwick CWG, Parsons MR, Johnson AP (2009) Structure-based design, synthesis, and characterization of inhibitors of human and Plasmodium falciparum dihydroorotate dehydrogenases. J Med Chem 52:2683–2693
Heikkilä T, Ramsey C, Davies M, Galtier C, Stead AM, Johnson AP, Fishwick CW, Boa AN, McConkey GA (2007) Design and synthesis of potent inhibitors of the malaria parasite dihydroorotate dehydrogenase. J Med Chem 50:186–191
Gujjar R, Marwaha A, El Mazouni F, White J, White KL, Creason S, Shackleford DM, Baldwin J, Charman WN, Buckner FS, Charman S (2009) Identification of a metabolically stable triazolopyrimidine-based dihydroorotate dehydrogenase inhibitor with antimalarial activity in mice. J Med Chem 52:1864–1872
Gujjar R, El Mazouni F, White KL, White J, Creason S, Shackleford DM, Deng X, Charman WN, Bathurst I, Burrows J, Floyd DM (2011) Lead-optimization of aryl and aralkyl amine based triazolopyrimidine inhibitors of Plasmodium falciparum dihydroorotate dehydrogenase with anti-malarial activity in mice. J Med Chem 54:3935–3949
The Binding Database (2012). https://www.bindingdb.org
Hansen M, Le Nours J, Johansson EV, Antal T, Ullrich A, Löffler M, Larsen S (2004) Inhibitor binding in a class 2 dihydroorotate dehydrogenase causes variations in the membrane-associated N-terminal domain. Protein Sci 13:1031–1042
Cheng YC, Prusoff WH (1973) Relationship between the inhibition constant (Ki) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem Pharmacol 22:3099–3108
ADMET Predictor (2020) Simulations Plus, Inc., Lancaster, CA. https://www.simulations-plus.com
Gamo FJ, Sanz LM, Vidal J, De Cozar C, Alvarez E, Lavandera JL, Vanderwall DE, Green DV, Kumar V, Hasan S, Brown JR (2010) Thousands of chemical starting points for antimalarial lead identification. Nature 465:305–310
Ekins S, Williams AJ (2010) When pharmaceutical companies publish large datasets: an abundance of riches or fool's gold? Drug Discov Today 15:812–815
MedChem Studio (2020) Simulations Plus, Inc., Lancaster, CA. https://www.simulations-plus.com; this functionality has since been subsumed into the MedChem Studio Module of ADMET Predictor
Plouffe D, Brinker A, McNamara C, Henson K, Kato N, Kuhen K, Nagle A, Adrián F, Matzen JT, Anderson P, Nam TG (2008) In silico activity profiling reveals the mechanism of action of antimalarials discovered in a high-throughput screen. Proc Natl Acad Sci 105:9059–9064
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26
Coteron JM, Marco M, Esquivias J, Deng X, White KL, White J, Koltun M, El Mazouni F, Kokkonda S, Katneni K, Bhamidipati R (2011) Structure-guided lead optimization of triazolopyrimidine-ring substituents identifies potent Plasmodium falciparum dihydroorotate dehydrogenase inhibitors with clinical candidate potential. J Med Chem 54:5540–5561
Dong CK, Patel V, Yang JC, Dvorin JD, Duraisingh MT, Clardy J, Wirth DF (2009) Type II NADH dehydrogenase of the respiratory chain of Plasmodium falciparum and its inhibitors. Bioorg Med Chem Lett 19:972–975
Ertl P, Schuffenhauer A (2009) Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J Cheminform 1:8
Berge JM, Brown P, Elder JS, Forrest AK, Hamprecht DW, Jarvest RL, McNair DJ, Sheppard RJ (2001) Quinolones used as MRS inhibitors and bactericides. US Patent 6320051
GastroPlus (2020) Simulations Plus, Inc., Lancaster, CA. https://www.simulations-plus.com
Simulations Plus, Inc., Press release (9/13/2011) https://www.irdirect.net/prviewer/release/id/2168302
Okubo T, Kumagi T, Nakamura T, Abe K, Amada Y, Ishizaka T, Sun X-M, Sekiguchi Y, Sasako S, Shimizu T, Nagatsuka T (2011) Aminopyrrolidine compound. US Patent US8044068B2
Clark RD (2018) Predicting mammalian metabolism and toxicity of pesticides in silico. Pest Manage Sci 74:1992–2003
Clark RD, Liang W, Lee AC, Lawless MS, Fraczkiewicz R, Waldman M (2014) Using beta binomials to estimate classification uncertainty for ensemble models. J Cheminform 6:34
Houston JB, Kenworthy KE (2000) In vitro-in vivo scaling of CYP kinetic data not consistent with the classical Michaelis–Menten model. Drug Metab Dispos 28:246–254
Looareesuwan S, White NJ, Chanthavanich PO, Edwards G, Nicholl DD, Bunch C, Warrell DA (1986) Cardiovascular toxicity and distribution kinetics of intravenous chloroquine. Br J Clin Pharmacol 22:31–36
Daub ME, Prudhomme J, Le Roch K, Vanderwal CD (2015) Synthesis and potent antimalarial activity of kalihinol B. J Am Chem Soc 137:4912–4915
Prudhomme J, McDaniel E, Ponts N, Bertani S, Fenical W, Jensen P, Le Roch K (2008) Marine actinomycetes: a new source of compounds against the human malaria parasite. PLoS ONE 3:e2335
Ray S, Madrid PB, Catz P, LeValley SE, Furniss MJ, Rausch LL, Guy RK, DeRisi JL, Iyer LV, Green CE, Mirsalis JC (2010) Development of a new generation of 4-aminoquinoline antimalarial compounds using predictive pharmacokinetic and toxicology models. J Med Chem 53:3685–3695
Martin RE, Marchetti RV, Cowan AI, Howitt SM, Bröer KK (2009) Chloroquine transport via the malaria parasite’s chloroquine resistance transporter. Science 325:1680–1682
Kumar R, Duffy S, Avery VM, Davis RA (2017) Synthesis of antimalarial amide analogues based on the plant serrulatane diterpenoid 3,7,8-trihydroxyserrulat-14-en-19-oic acid. Biorg Med Chem Lett 27:4091–4095
White NJ (2011) The parasite clearance curve. Malar J 10:278
Burrows JN, Duparc S, Gutteridge WE, van Huijsduijnen RH, Kaszubska W, Macintyre F, Mazzuri S, Möhrle JJ, Wells TN (2017) New developments in anti-malarial target candidate and product profiles. Malar J 16:26
Phillips MA, Lotharius J, Marsh K, White J, Dayan A, White KL, Njoroge JW, El Mazouni F, Lao Y, Kokkonda S, Tomchick DR (2015) A long-duration dihydroorotate dehydrogenase inhibitor (DSM265) for prevention and treatment of malaria. Sci Transl Med 7:296ra111
Chaudhari R, Tan Z, Zhang S (2017) Overview of drug polypharmacology and multitargeted molecular design. In: Wilson AGE (ed) Comprehensive medicinal chemistry III. Elsevier, Oxford, pp 259–275
Seymour KK, Lyons SD, Phillips L, Rieckmann KH, Christopherson RI (1994) Cytotoxic effects of inhibitors of de novo pyrimidine biosynthesis upon Plasmodium falciparum. Biochemistry 33:5268–5274
Hurt DE, Widom J, Clardy J (2006) Structure of Plasmodium falciparum dihydroorotate dehydrogenase with a bound inhibitor. Acta Cryst D62:312–323
Painter HJ, Morrisey JM, Mather MW, Vaidya AB (2007) Specific role of mitochondrial electron transport in blood-stage Plasmodium falciparum. Nature 446:88–91
Phillips MA, Gujjar R, Malmquist NA, White J, El Mazouni F, Baldwin J, Rathod PK (2016) A triazolopyrimidine-based dihydroorotate dehydrogenase inhibitor with improved drug-like properties for treatment and prevention of malaria. ACS Infect Dis 2:945–957
Jarvest RL, Berge JM, Berry V, Boyd HF, Brown MJ, Elder JS, Forrest AK, Fosberry AP, Gentry DR, Hibbs MJ, Jaworski DD, O’Hanlon PJ, Pope AJ, Rittenhouse S, Sheppard RJ, Slater-Radosti C, Worby A (2002) Nanomolar Inhibitors of Staphylococcus aureus methionyl tRNA synthetase with Potent antibacterial activity against gram-positive pathogens. J Med Chem 45:1959–1962
Hussain T, Yogavel M, Sharma A (2015) Inhibition of protein synthesis and malaria parasite development by drug targeting of methionyl-tRNA synthetases. Antimicrob Agents Chemother 59:1856–1867
Baragaña B, Hallyburton I, Lee MC, Norcross NR, Grimaldi R, Otto TD, Proto WR, Blagborough AM, Meister S, Wirjanata G, Ruecker A (2015) A novel multiple-stage antimalarial agent that inhibits protein synthesis. Nature 522:315–320
Acknowledgements
The authors are indebted to the Medicines for Malaria Venture for logistical support and to Jeremy Burrows and Xavier Ding for orchestrating and facilitating the transfer of materials to Tres Cantos for assay; to Richard Lloyd of GlaxoSmithKline for his support and encouragement throughout the antimalarial project; to Mike Melnick for his synthesis support; and to Kayla Andrews for her careful reading of an earlier version of the manuscript and editorial suggestions. We would also like to thank our reviewers for their careful reading of the manuscript and thoughtful comments.
Funding
No external funding was used for the research described herein.
Author information
Authors and Affiliations
Contributions
The Simulations Plus and Cognigen groups did the modeling, analog design, and pharmacokinetic simulations work described herein and drafted the manuscript. The Kalexsyn group helped refine the list of synthesis targets, then planned and carried out the synthesis of the final set of cyclopentyl-1,1-dimethylene analogs. The UC Riverside and Tres Cantos groups carried out XC50 determinations in blood culture. The Tres Cantos GSK group also conducted the kill rate and ScDHODH rescue assays. Synthesis and academic work were underwritten by Simulations Plus, Inc.
Corresponding author
Ethics declarations
Conflict of interest
Robert Clark, Michael Lawless, and Walter Woltosz are employees of Simulations Plus, Inc. and own stock and/or options in the company. The authors declare no other conflicts of interest or competing interests beyond those implicit in their employment by their respective institutions.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10822_2020_333_MOESM1_ESM.xlsx
Supplementary file1 (XLSX 362 kb)—Supplemental Information: Data.Supplementary Table S1 contains structures and Ki values used to build the PfDHODH inhibition models. Structuresof the original 12 synthesis candidates are provided in Supplementary Table S2, along with their predictedPfDHODH Ki’s and critical ADME property predictions. Data on the four analogs that were synthesized and fullycharacterized (8, 9, 11a and 11b) are given in Supplementary Tables S3 and S4. Supplementary Table S3 comparesmetabolism predictions from ADMET Predictor v9.0 with those from v6.0 and with the corresponding experimental values. Predictions of the corresponding predicted and experimentally determined physicochemical properties areprovided in Table S4.
10822_2020_333_MOESM2_ESM.docx
Supplementary file2 (DOCX 207 kb)—Supplemental Information: MethodsMethodological details concerning computations, assays, synthesis and chemical characterization of products areprovided in the supporting materials.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Clark, R.D., Morris, D.N., Chinigo, G. et al. Design and tests of prospective property predictions for novel antimalarial 2-aminopropylaminoquinolones. J Comput Aided Mol Des 34, 1117–1132 (2020). https://doi.org/10.1007/s10822-020-00333-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-020-00333-x