Abstract
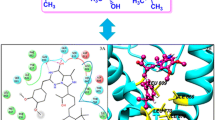
The hyperactivity of the cyclic dependent kinase 5 (CDK5) induced by the activator protein p25 has been linked to a number of pathologies of the brain. The CDK5–p25 complex has thus emerged as a major therapeutic target for Alzheimer’s disease (AD) and other neurodegenerative conditions. Experiments have shown that the peptide p5 reduces the CDK5–p25 activity without affecting the endogenous CDK5–p35 activity, whereas the peptide TFP5, obtained from p5, elicits similar inhibition, crosses the blood–brain barrier, and exhibits behavioral rescue of AD mice models with no toxic side effects. The molecular basis of the kinase inhibition is not currently known, and is here investigated by computer simulations. It is shown that p5 binds the kinase at the same CDK5/p25 and CDK5/p35 interfaces, and is thus a non-selective competitor of both activators, in agreement with available experimental data in vitro. Binding of p5 is enthalpically driven with an affinity estimated in the low µM range. A quantitative description of the binding site and pharmacophore is presented, and options are discussed to increase the binding affinity and selectivity in the design of drug-like compounds against AD.





Similar content being viewed by others
References
Nikolic M, Dudek H, Kwon YT, Ramos YF, Tsai LH (1996) The cdk5/p35 kinase is essential for neurite outgrowth during neuronal differentiation. Genes Dev 10:816–825
Ohshima T, Ward JM, Huh CG, Longenecker G, Veeranna Pant HC et al (1996) Targeted disruption of the cyclin-dependent kinase 5 gene results in abnormal corticogenesis, neuronal pathology and perinatal death. Proc Natl Acad Sci USA 93:11173–11178
Tan TC, Valova VA, Malladi CS, Graham ME, Berven LA, Jupp OJ et al (2003) Cdk5 is essential for synaptic vesicle endocytosis. Nat Cell Biol 5:701–710
Ahlijanian MK, Barrezueta NX, Williams RD, Jakowski A, Kowsz KP, McCarthy S et al (2000) Hyperphosphorylated tau and neurofilament and cytoskeletal disruptions in mice overexpressing human p25, an activator of cdk5. Proc Natl Acad Sci USA 97:2910–2915
de la Monte SM, Ganju N, Feroz N, Luong T, Banerjee K, Cannon J et al (2000) Oxygen free radical injury is sufficient to cause some Alzheimer type molecular abnormalities in human CNS neuronal cells. J Alzheimer’s Dis 2:261–281
Patrick GN, Zukerberg L, Nikolic M, de la Monte S, Dikkes P, Tsai LH (1999) Conversion of p35 to p25 deregulates CDK5 activity and promotes neurodegeneration. Nature 402:615–622
Shukla V, Zheng YL, Mishra SK, Amin ND, Steiner J, Grant P et al (2013) A truncated peptide from p35, a Cdk5 activator, prevents Alzheimer’s disease phenotypes in model mice. FASEB J 27:174–186
Lee MS, Kwon YT, Li M, Peng J, Friedlander RM, Tsai LH (2000) Neurotoxicity induces cleavage of p35 to p25 by calpain. Nature 405:360
Noble W, Olm V, Takata K, Casey E, Mary O, Meyerson J et al (2003) CDK5 is a key factor in tau aggregation and tangle formation in vivo. Neuron 38:555–565
Bard F, Cannon C, Barbour R, Burke RL, Games D, Grajeda H et al (2000) Peripherally administered antibodies against amyloid beta-peptide enter the central nervous system and reduce pathology in a mouse model of Alzheimer disease. Nat Med 6:916–919
Maier M, Seabrook TJ, Lazo ND, Jiang L, Das P, Janus C et al (2006) Short amyloid-beta (Abeta) immunogens reduce cerebral Abeta load and learning deficits in an Alzheimer’s disease mouse model in the absence of an Abeta-specific cellular immune response. J Neurosci 26:4717–4728
Ritchie CW, Ames D, Clayton T, Lai R (2004) Meta-analysis of randomized trials of the efficacy and safety of donepezil, galantamine, and rivastigmine for the treatment of Alzheimer disease. Am J Geriatr Psychiatry 12:358–369
Bonda DJ, Wang X, Perry G, Nunomura A, Tabaton M, Zhu X et al (2010) Oxidative stress in Alzheimer disease: a possibility for prevention. Neuropharmacology 59:290–294
Lee HP, Zhu X, Casadesus G, Castellani RJ, Nunomura AS, Smith MA, Lee HG et al (2010) Antioxidant approaches for the treatment of Alzheimer’s disease. Expert Rev Neurother 10:1201–1208
Helal CJ, Sanner MA, Cooper CB, Gant T, Adam M, Lucas JC et al (2004) Discovery and SAR of 2-aminothiazole inhibitors of cyclin-dependent kinase 5/p25 as a potential treatment for Alzheimer’s disease. Bioorg Med Chem Lett 14:5521–5525
Helal CJ, Kang Z, Lucas JC, Gant T, Ahlijanian MK, Schachter JB et al (2009) Potent and cellularly active 4 aminoimidazole inhibitors of cyclin-dependent kinase 5/p25 for the treatment of Alzheimer’s disease. Bioorg Med Chem Lett 19:5703–5707
Knockaert M, Wieking K, Schmitt S, Leost M, Grant KM, Mottram JC et al (2002) Intracellular targets of paullones. Identification following affinity purification on immobilized inhibitor. J Biol Chem 277:25493–25501
Zheng YL, Kesavapany S, Gravell M, Hamilton RS, Schubert M, Amin ND et al (2005) A Cdk5 inhibitory peptide reduces tau hyperphosphorylation and apoptosis in neurons. EMBO J 24:209–220
Zheng YL, Amin ND, Hu YF, Rudrabhatla P, Shukla VK, Kanungo J et al (2010) A 24-residue peptide (p5), derived from p35, the cdk5 neuronal activator, specifically inhibits CDK5–p25 hyperactivity and tau hyperphosphorylation. J Biol Chem 285:34202–34212
Binukumar BK, Zheng YL, Shukla V, Amin ND, Grant P, Pant HC (2014) TFP5, a peptide derived from p35, a CDK5 neuronal activator, rescues cortical neurons from glucose toxicity. J Alzheimer’s Dis 39:899–909
Cardone A, Pant H, Hassan SA (2013) Specific and non-specific protein association in solution: computation of solvent effects and prediction of first-encounter modes for efficient configurational bias Monte Carlo simulations. J Phys Chem B 117:12360–12374
Cardone A, Bornstein A, Pant HC, Brady M, Sriram R, Hassan SA (2015) Detection and characterization of nonspecific, sparsely-populated binding modes in the early stages of complexation. J Comput Chem 36:983–995
Hassan SA, Steinbach PJ (2011) Water-exclusion and liquid-structure forces in implicit solvation. J Phys Chem B. 115:14668
Hassan SA, Mehler EL (2012) In silico approaches to structure and function of cell components and their assemblies: molecular electrostatics and solvent effects. In: Egelman E (ed) Comprehensive biophysics. Academic Press, New York
Hassan SA (2014) Implicit treatment of solvent dispersion forces in protein simulations. J Comput Chem 35:1621–1629
Tang C, Ghirlando R, Clore GM (2008) Visualization of transient ultra-weak protein self-association in solution using paramagnetic relaxation enhancement. J Am Chem Soc 130:4048
Johansson H, Jensen MR, Gesmar H, Meier S, Vinther JM, Keeler C et al (2014) Specific and nonspecific interactions in ultraweak protein-protein association revealed by solvent paramagnetic relaxation enhancements. J Am Chem Soc 136:10277–10286
Tarricone C, Dhavan R, Peng J, Areces LB, Tsai LH, Musacchio A (2001) Structure and regulation of the CDK5–p25(nck5a) complex. Mol Cell 8:657
Steinbach PJ (2004) Exploring peptide energy lanscapes:a test of force fields and implicit solvent models. Proteins 57:665
Brooks BR, Brooks CL, Mackerell AD, Nilsson L, Petrella RJ, Roux B et al (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30:1545
Hassan SA, Mehler EL, Zhang D, Weinstein H (2003) Molecular dynamics simulations of peptides and proteins with a continuum electrostatic model based on screened Coulomb potentials. Proteins 51:109–125
Hassan SA, Mehler EL (2005) From quantum chemistry and the classical theory of polar liquids to continuum approximations in molecular mechanics calculations. Int J Quantum Chem 102:986
Szekely GJ, Rizzo ML (2005) Hierarchical clustering via joint between-within distances: extending Ward’s minimum variance method. J Classif 22:151–183
Mapelli M, Massimiliano L, Crovace C, Seeliger MA, Tsai LH, Meijer L et al (2005) Mechanism of CDK5/p25 binding by CDK inhibitors. J Med Chem 48:671
Cardone A, Hassan SA, Albers RW, Sriram RD, Pant HC (2010) Structural and dynamic determinants of ligand binding and regulation of cyclin-dependent kinase 5 by pathological activator p25 and inhibitory peptide CIP. J Mol Biol 401:478–492
Cardone A, Albers RW, Sriram RD, Pant HC (2010) Evaluation of the interaction of cyclin-dependent kinase 5 with activator p25 and with p25-derived inhibitor CIP. J Comput Biol 17:1–15
D’Aquino JA, Freire E, Amzel LM (2000) Binding of small organic molecules to macromolecular targets: evaluation of conformational entropy changes. Proteins Struct Funct Genet 41(S4): 93–107
Yu YB, Privalov PL, Hodges RS (2001) Contribution of translational and rotational motions to molecular association in aqueous solution. Biophys J 81:1632–1642
Andricioaei I, Karplus M (2001) On the calculation of entropy from covariance matrices of the atomic fluctuations. J Chem Phys 115:6289–6292
Acknowledgments
This study utilized the high-performance computer capabilities of the Biowulf PC/Linux cluster at the NIH. This work was supported by the NIH Intramural Research Program through the CIT and NINDS, and an internal NIST Research Fund.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclaimer Commercial products are identified in this document in order to specify the experimental procedure adequately. Such identification is not intended to imply recommendation or endorsement by the National Institute of Standards and Technology or the National Institutes of Health, nor is it intended to imply that the products identified are necessarily the best available for the purpose.
Rights and permissions
About this article
Cite this article
Cardone, A., Brady, M., Sriram, R. et al. Computational study of the inhibitory mechanism of the kinase CDK5 hyperactivity by peptide p5 and derivation of a pharmacophore. J Comput Aided Mol Des 30, 513–521 (2016). https://doi.org/10.1007/s10822-016-9922-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-016-9922-3