Abstract
The phosphodiesterase (PDE) family of proteins are important regulators of signal transduction, which they achieve by controlling the secondary messengers cyclic AMP (cAMP) and cyclic GMP (cGMP). cAMP and cGMP are involved in many critical intracellular processes such as gene transcription, kinase activation, signal transduction in learning and memory, and channel function as secondary messengers. The involvement of PDEs in neuronal communication has made them important therapeutic targets. Considering the recent discovery that PDE2A inhibition can improve cognitive functioning, a combined molecular dynamics simulation and scoring and docking study was carried out to identify selective inhibitors of PDE2A that specifically interact with the recently discovered hydrophobic groove in PDE2A. Using the X-ray crystal structure of PDE2A (from PDB ID: 4HTX), we investigated the binding modes of a range of promising inhibitors based on the known PDE2A inhibitor BAY60-7550 to PDE2A.

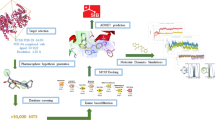
The lead molecule showing highest MMPBSA binding energy with 2D and 3D binding pose in hydrophobic groove.






Similar content being viewed by others
References
Mobashir M, Madhusudhan T, Isermann B, Beyer T, Schraven B (2014) Negative interactions and feedback regulations are required for transient cellular response. Sci Rep 4:3718
Umar T, Hoda N (2015) Selective inhibitors of phosphodiesterases: therapeutic promise for neurodegenerative disorders. Med Chem Commun 6:2063–2080
Fawcett L, Baxendale R, Stacey P, McGrouther C, Harrow I, Soderling S et al (2000) Molecular cloning and characterization of a distinct human phosphodiesterase gene family: PDE11A. Proc Natl Acad Sci USA 97:3702–3707
Beavo JA, Brunton LL (2002) Cyclic nucleotide research—still expanding after half a century. Nat Rev Mol Cell Biol 3:710–718
Manganiello VC, Degerman E (1999) Cyclic nucleotide phosphodiesterases (PDEs): diverse regulators of cyclic nucleotide signals and inviting molecular targets for novel therapeutic agents. Thromb Haemost 82:407–411
Laddha S, Wadodkar S, Meghal S (2009) cAMP-dependent phosphodiesterase inhibition and SAR studies on novel 6,8-disubstituted 2-phenyl-3-(substituted benzothiazole-2-yl)-4[3H]-quinazolinone. Med Chem Res 18:268–276
Francis SH, Blount MA, Corbin JD (2011) Mammalian cyclic nucleotide phosphodiesterases: molecular mechanisms and physiological functions. Physiol Rev 91:651–690
Conti M, Beavo J (2007) Biochemistry and physiology of cyclic nucleotide phosphodiesterases: essential components in cyclic nucleotide signaling. Annu Rev Biochem 76:481–511
Lugnier C (2006) Cyclic nucleotide phosphodiesterase (PDE) superfamily: a new target for the development of specific therapeutic agents. Pharmacol Ther 109:366–398
Iffland A, Kohls D, Low S, Luan J, Zhang Y, Kothe M et al (2005) Structural determinants for inhibitor specificity and selectivity in PDE2A using the wheat germ in vitro translation system. Biochemistry 44:8312–8325
Zhu J, Yang Q, Dai D, Huang Q (2013) X-ray crystal structure of phosphodiesterase 2 in complex with a highly selective, nanomolar inhibitor reveals a binding-induced pocket important for selectivity. J Am Chem Soc 135:11708–11711
Wang H, Robinson H, Ke H (2007) The molecular basis for different recognition of substrates by phosphodiesterase families 4 and 10. J Mol Biol 371:302–307
Wang H, Liu Y, Hou J, Zheng M, Robinson H, Ke H (2007) Structural insight into substrate specificity of phosphodiesterase 10. Proc Natl Acad Sci USA 104:5782–5787
Xu RX, Hassell AM, Vanderwall D, Lambert MH, Holmes WD, Luther MA et al (2000) Atomic structure of PDE4: insights into phosphodiesterase mechanism and specificity. Science 288:1822–1825
Liu S, Mansour MN, Dillman KS, Perez JR, Danley DE, Aeed PA et al (2008) Structural basis for the catalytic mechanism of human phosphodiesterase 9. Proc Natl Acad Sci USA 105:13309–13314
Gupta A, Gandhimathi A, Sharma P, Jayaram B (2007) ParDOCK: an all atom energy based Monte Carlo docking protocol for protein–ligand complexes. Protein Pept Lett 14:632–646
Pearlman DA, Case DA, Caldwell JW, Ross WS, Cheathem JE III et al (1995) AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Comput Phys Commun 91:1–41
van der Spoel D, Lindahl E, Hess B, Kutzner C, van Buuren AR, Apol E et al (2006) GROMACS user manual, version 4.0. GROMACS Development Team, Groningen
Schüttelkopf AW, van Aalten DMF (2004) PRODRG: a tool for high-throughput crystallography of protein–ligand complexes. Acta Crystallogr D 60:1355–1363
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Kumari R, Kumar R, Lynn A (2014) g_mmpbsa—a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962
He JY, Li C, Wu G (2014) Discovery of potential drugs for human-infecting H7N9 virus containing R294K mutation. Drug Des Devel Ther 8:2377–2390
Kastritis PL, Bonvin AM (2010) Are scoring function in protein–protein docking ready to predict interactomes? Clues from a novel binding affinity benchmark. J Proteome Res 9(5):2216–2225
Acknowledgements
JK thanks the CSIR for a doctoral research fellowship. TU thanks the UGC for a basic scientific research fellowship. We also thank Prof. B. Jayaram, IIT Delhi, for access to the supercomputing facility. We acknowledge the support provided by the BRAF at C-DAC, Pune, India, when we were carrying out MD simulations using computational resources at that facility.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 3426 kb)
Rights and permissions
About this article
Cite this article
Kumar, J., Umar, T., Kausar, T. et al. Identification of lead BAY60-7550 analogues as potential inhibitors that utilize the hydrophobic groove in PDE2A: a molecular dynamics simulation study. J Mol Model 23, 7 (2017). https://doi.org/10.1007/s00894-016-3171-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-016-3171-1