Abstract
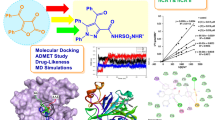
Human acrosin is an attractive target for the discovery of male contraceptive drugs. For the first time, structure-based drug design was applied to discover structurally diverse human acrosin inhibitors. A parallel virtual screening strategy in combination with pharmacophore-based and docking-based techniques was used to screen the SPECS database. From 16 compounds selected by virtual screening, a total of 10 compounds were found to be human acrosin inhibitors. Compound 2 was found to be the most potent hit (IC50 = 14 μM) and its binding mode was investigated by molecular dynamics simulations. The hit interacted with human acrosin mainly through hydrophobic and hydrogen-bonding interactions, which provided a good starting structure for further optimization studies.










Similar content being viewed by others
References
Klemm U, Muller-Esterl W, Engel W (1991) Acrosin, the peculiar sperm-specific serine protease. Hum Genet 87:635–641
Howes L, Jones R (2002) Interactions between zona pellucida glycoproteins and sperm proacrosin/acrosin during fertilization. J Reprod Immunol 53:181–192
Baskova IP, Zavalova LL (2001) Proteinase inhibitors from the medicinal leech Hirudo medicinalis. Biochemistry (Mosc) 66:703–714
Pakzad R (1989) The effect of the trypsin inhibitor aprotinin (Trasylol) and TLCK on the gelatinolytic activity of acrosin and the motility of rabbit sperm in vitro. Z Mikrosk Anat Forsch 103:8–13
Jones R, Parry R, Lo Leggio L, Nickel P (1996) Inhibition of sperm-zona binding by suramin, a potential ‘lead’ compound for design of new anti-fertility agents. Mol Hum Reprod 2:597–605
Fraser LR (1982) p-Aminobenzamidine, an acrosin inhibitor, inhibits mouse sperm penetration of the zona pellucida but not the acrosome reaction. J Reprod Fertil 65:185–194
Gupta G, Jain RK, Maikhuri JP, Shukla PK, Kumar M, Roy AK, Patra A, Singh V, Batra S (2005) Discovery of substituted isoxazolecarbaldehydes as potent spermicides, acrosin inhibitors and mild anti-fungal agents. Hum Reprod 20:2301–2308
Bourinbaiar AS, Lee-Huang S (1995) Acrosin inhibitor, 4′-acetamidophenyl 4-guanidinobenzoate, an experimental vaginal contraceptive with anti-HIV activity. Contraception 51:319–322
Tranter R, Read JA, Jones R, Brady RL (2000) Effector sites in the three-dimensional structure of mammalian sperm beta-acrosin. Structure 8:1179–1188
Lu JG, Sheng CQ, Zhang M, Ji HT, Zhang WN, Zhou YJ, Zhu J, Jiang JH (2006) Homology modeling of human acrosin and its molecular docking study with KF950. Acta Chimica Sinica 64:1073–1078
Zhang J, Zheng C, Sheng C, Zhou Y, Zhu J, Lv J (2009) Characterization of human acrosin active site and binding modes with its inhibitors. Chem J Chinese U 30:2409–2414
Kolb P, Rosenbaum DM, Irwin JJ, Fung JJ, Kobilka BK, Shoichet BK (2009) Structure-based discovery of beta2-adrenergic receptor ligands. Proc Natl Acad Sci U S A 106:6843–6848
Li H, Huang J, Chen L, Liu X, Chen T, Zhu J, Lu W, Shen X, Li J, Hilgenfeld R, Jiang H (2009) Identification of novel falcipain-2 inhibitors as potential antimalarial agents through structure-based virtual screening. J Med Chem 52:4936–4940
Shoichet BK (2004) Virtual screening of chemical libraries. Nature 432:862–865
Leach AR, Gillet VJ, Lewis RA, Taylor R (2010) Three-dimensional pharmacophore methods in drug discovery. J Med Chem 53:539–558
Steindl T, Langer T (2004) Influenza virus neuraminidase inhibitors: generation and comparison of structure-based and common feature pharmacophore hypotheses and their application in virtual screening. J Chem Inf Comput Sci 44:1849–1856
Disocovery studio 2.5 software package AI, San Diego, CA (USA): http://www.accelrys.com
La Motta C, Sartini S, Tuccinardi T, Nerini E, Da Settimo F, Martinelli A (2009) Computational studies of epidermal growth factor receptor: docking reliability, three-dimensional quantitative structure-activity relationship analysis, and virtual screening studies. J Med Chem 52:964–975
Rao SN, Head MS, Kulkarni A, LaLonde JM (2007) Validation studies of the site-directed docking program LibDock. J Chem Inf Model 47:2159–2171
Jones G, Willett P, Glen RC, Leach AR, Taylor R (1997) Development and validation of a genetic algorithm for flexible docking. J Mol Biol 267:727–748
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) Automated docking using a lamarckian genetic algorithm and empirical binding free energy function. J Comput Chem 19:1639–1662
Lineweaver H, Burk D (1934) The determination of enzyme dissociation constants. J Am Chem Soc 56:658–666
Burk D (1984) Enzyme kinetic constants: the double reciprocal plot. Trends Biochem Sci 9:202–204
Greenidge PA, Carlsson B, Bladh LG, Gillner M (1998) Pharmacophores incorporating numerous excluded volumes defined by X-ray crystallographic structure in three-dimensional database searching: application to the thyroid hormone receptor. J Med Chem 41:2503–2512
Norinder U (2000) Refinement of catalyst hypotheses using simplex optimisation. J Comput Aided Mol Des 14:545–557
Palomer A, Cabre F, Pascual J, Campos J, Trujillo MA, Entrena A, Gallo MA, Garcia L, Mauleon D, Espinosa A (2002) Identification of novel cyclooxygenase-2 selective inhibitors using pharmacophore models. J Med Chem 45:1402–1411
Shaw DE (2005) A fast, scalable method for the parallel evaluation of distance-limited pairwise particle interactions. J Comput Chem 26:1318–1328
Bowers KJ, Chow E, Xu H, Dror RO, Eastwood MP, Gregersen BA, Klepeis JL, Kolossváry I, Moraes MA, Sacerdoti JK, Shan Y, Shaw DE (2006) Scalable algorithms for molecular dynamics simulations on commodity clusters. In: Proceedings of the ACM/IEEE conference on supercomputing (SC06), Tampa, Florida, November 11–17
Bowers KJ, Dror RO, Shaw DE (2006) The midpoint method for parallelization of particle simulations. J Chem Phys 124:184109
Bowers KJ, Dror RO, Shaw DE (2007) Zonal methods for the parallel execution of range-limited N-body simulations. J Comput Phys 221:303–329
Chillemi G, D’Annessa I, Fiorani P, Losasso C, Benedetti P, Desideri A (2008) Thr729 in human topoisomerase I modulates anti-cancer drug resistance by altering protein domain communications as suggested by molecular dynamics simulations. Nucleic Acids Res 36:5645–5651
Marco E, Laine W, Tardy C, Lansiaux A, Iwao M, Ishibashi F, Bailly C, Gago F (2005) Molecular determinants of topoisomerase I poisoning by lamellarins: comparison with camptothecin and structure-activity relationships. J Med Chem 48:3796–3807
Kennedy WP, Kaminski JM, Van der Ven HH, Jeyendran RS, Reid DS, Blackwell J, Bielfeld P, Zaneveld LJ (1989) A simple, clinical assay to evaluate the acrosin activity of human spermatozoa. J Androl 10:221–231
Hortin GL, Warshawsky I, Laude-Sharp M (2001) Macromolecular chromogenic substrates for measuring proteinase activity. Clin Chem 47:215–222
Yang TC, Liu YR (1994) Kinetics of inhibitory effects of Phaseolus calcaratus inhibitor on acrosin of human body. J Fujian Normal University (Natural Science) 10:8–12
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
10822_2011_9476_MOESM1_ESM.pdf
Dataset used in virtual screening test; Binding mode of hit 2 with ram and boar acrosin. Supplementary material 1 (PDF 401 kb)
Rights and permissions
About this article
Cite this article
Liu, X., Dong, G., Zhang, J. et al. Discovery of novel human acrosin inhibitors by virtual screening. J Comput Aided Mol Des 25, 977–985 (2011). https://doi.org/10.1007/s10822-011-9476-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-011-9476-3