Abstract
Purpose
To reveal the underlying roles that pyroptosis-related genes (PRGs) played in human spermatogenic dysfunction.
Methods
One discovery set and three validation sets were employed to inspect the previously reported 33 PRGs in the human testis with different status of spermatogenesis. PRGs that differentially expressed in all sets were considered as key differentially expressed pyroptosis-related genes (PR-DEGs). The relationships between key PR-DEGs and samples’ clinicopathological, therapeutic, and immune patterns were respectively studied. Single-cell RNA sequencing (scRNS-seq) analyses were conducted to show the expression changes and related mechanisms of key PR-DEGs at a single-cell resolution.
Results
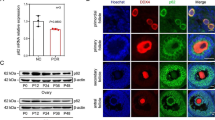
CASP4 and GPX4 were identified as two key PR-DEGs. These two genes were significantly dysregulated in spermatogenic dysfunctional samples, but with opposite tendency. CASP4 was negatively correlated with Johnsen scores but positively correlated with follicle-stimulating hormone (FSH) levels (all p < 0.05), while GPX4 exhibited significant positive correlations with Johnsen scores and negative relevance with FSH. For treatments, both molecules showed a prospective value of being predictors for sperm retrieval surgeries. Moreover, CASP4 and GPX4 were potential immunoregulators in the testicular immune microenvironment and showed significant correlations to testicular macrophages and mast cell infiltration. In scRNA-seq analyses, GPX4 was highly expressed in germ cells, which therefore suffered a sharp reduction with the loss of germ cells in spermatogenic dysfunction. On the other hand, CASP4 were basically somatic cell–derived, and the proportion of CASP4-positive Leydig cells significantly increased in disease testes (p = 0.0001).
Conclusion
In all, we revealed two key PRGs of human testes that might be functional in spermatogenic dysfunction.






Similar content being viewed by others
Data availability
The data that support the findings of this study are all publicly available or have been included in the manuscript.
Abbreviations
- PRGS:
-
pyroptosis-related genes
- PR-DEGs:
-
differentially expressed pyroptosis-related genes
- ScRNS-seq:
-
single-cell RNA sequencing
- FSH:
-
follicle-stimulating hormone
- ROC:
-
receiver operating characteristic
- NOA:
-
non-obstructive azoospermia
- PCD:
-
programmed cell death
- GEO:
-
Gene Expression Omnibus
- KNN:
-
k-nearest neighbor
- SNN:
-
shared nearest neighbor
- UMAP:
-
Uniform Manifold Approximation and Projection
- DEGs:
-
differentially expressed genes
- LogFC:
-
log2 fold change
- IHC:
-
immunohistochemical staining
- ssGSEA:
-
single sample Gene Set Enrichment Analysis
- tiPPI:
-
testicular immune protein-protein interaction
- GSEA:
-
gene set enrichment analysis
- MSigDB:
-
Molecular Signatures Database
- NES:
-
normalized enrichment score
- SD:
-
standard deviation
- SSCs&SPGs:
-
spermatogonial stem cells and spermatogonia
- SPCs:
-
spermatocytes
- PTM cells:
-
peritubular myoid cells
- HPA:
-
Human Protein Atlas
References
Kang C, Punjani N, Schlegel PN. Reproductive chances of men with azoospermia due to spermatogenic dysfunction. J Clin Med. 2021;10(7). https://doi.org/10.3390/jcm10071400.
Hu Z, Li Z, Yu J, Tong C, Lin Y, Guo X, Lu F, Dong J, Xia Y, Wen Y, et al. Association analysis identifies new risk loci for non-obstructive azoospermia in Chinese men. Nat Commun. 2014;5:3857. https://doi.org/10.1038/ncomms4857.
Johnsen SG. Testicular biopsy score count--a method for registration of spermatogenesis in human testes: normal values and results in 335 hypogonadal males. Hormones. 1970;1(1):2–25. https://doi.org/10.1159/000178170.
De Kretser DM, Holstein AF. Testicular biopsy and abnormal germ cells. In: ESE H, editor. Human semen and fertility regulation in men. St. Louis: Mosby; 1976. p. 332–43.
Chen X, Ma Y, Zou S, Wang S, Qiu J, Xiao Q, Zhou L, Ping P. Comparison and outcomes of nonobstructive azoospermia patients with different etiology undergoing MicroTESE and ICSI treatments. Transl Androl Urol. 2019;8(4):366–73. https://doi.org/10.21037/tau.2019.04.08.
Shukla KK, Mahdi AA, Rajender S. Apoptosis, spermatogenesis and male infertility. Front Biosci (Elite Ed). 2012;4(2):746–54. https://doi.org/10.2741/415.
Callard GV, Jorgensen JC, Redding JM. Biochemical analysis of programmed cell death during premeiotic stages of spermatogenesis in vivo and in vitro. Dev Genet. 1995;16(2):140–7. https://doi.org/10.1002/dvg.1020160207.
D'Souza CA, Heitman J. Dismantling the Cryptococcus coat. Trends Microbiol. 2001;9(3):112–3. https://doi.org/10.1016/s0966-842x(00)01945-4.
Yu P, Zhang X, Liu N, Tang L, Peng C, Chen X. Pyroptosis: mechanisms and diseases. Signal Transduct Target Ther. 2021;6(1):128. https://doi.org/10.1038/s41392-021-00507-5.
Wang B, Yin Q. AIM2 inflammasome activation and regulation: a structural perspective. J Struct Biol. 2017;200(3):279–82. https://doi.org/10.1016/j.jsb.2017.08.001.
Kang R, Zeng L, Zhu S, Xie Y, Liu J, Wen Q, Cao L, Xie M, Ran Q, Kroemer G, et al. Lipid peroxidation drives gasdermin D-mediated pyroptosis in lethal polymicrobial sepsis. Cell Host Microbe. 2018;24(1):97–108 e104. https://doi.org/10.1016/j.chom.2018.05.009.
Ye Y, Dai Q, Qi H. A novel defined pyroptosis-related gene signature for predicting the prognosis of ovarian cancer. Cell Death Discov. 2021;7(1):71. https://doi.org/10.1038/s41420-021-00451-x.
Al Mamun A, Mimi AA, Aziz MA, Zaeem M, Ahmed T, Munir F, Xiao J. Role of pyroptosis in cancer and its therapeutic regulation. Eur J Pharmacol. 2021;910:174444. https://doi.org/10.1016/j.ejphar.2021.174444.
Wang D, Weng Y, Zhang Y, Wang R, Wang T, Zhou J, Shen S, Wang H, Wang Y. Exposure to hyperandrogen drives ovarian dysfunction and fibrosis by activating the NLRP3 inflammasome in mice. Sci Total Environ. 2020;745:141049. https://doi.org/10.1016/j.scitotenv.2020.141049.
Cai Z, He S, Liu R, Zhou L, Zhao L. Plumbagin rescues the granulosa cell's pyroptosis by reducing WTAP-mediated N6-methylation in polycystic ovary syndrome. J Ovarian Res. 2022;15(1):126. https://doi.org/10.1186/s13048-022-01058-1.
Li MY, Zhu XL, Zhao BX, Shi L, Wang W, Hu W, Qin SL, Chen BH, Zhou PH, Qiu B, et al. Adrenomedullin alleviates the pyroptosis of Leydig cells by promoting autophagy via the ROS-AMPK-mTOR axis. Cell Death Dis. 2019;10(7):489. https://doi.org/10.1038/s41419-019-1728-5.
Hong Y, Zhou Y, Shen L, Wei Y, Long C, Fu Y, Wu H, Wang J, Wu Y, Wu S, Wei G. Exposure to DEHP induces testis toxicity and injury through the ROS/mTOR/NLRP3 signaling pathway in immature rats. Ecotoxicol Environ Saf. 2021;227:112889. https://doi.org/10.1016/j.ecoenv.2021.112889.
Liang F, Zhang F, Zhang L, Wei W. The advances in pyroptosis initiated by inflammasome in inflammatory and immune diseases. Inflamm Res. 2020;69(2):159–66. https://doi.org/10.1007/s00011-020-01315-3.
Zhang Y, Xi F, Yu Q, Lou W, Zeng Z, Su N, Gao J, Duan S, Deng Y, Guo S, et al. Identification of a novel pyroptosis-related gene signature correlated with the prognosis of diffuse glioma patients. Ann Transl Med. 2021;9(24):1766. https://doi.org/10.21037/atm-21-6011.
Li XY, Zhang LY, Li XY, Yang XT, Su LX. A pyroptosis-related gene signature for predicting survival in glioblastoma. Front Oncol. 2021;11:697198. https://doi.org/10.3389/fonc.2021.697198.
Chu L, Yi Q, Yan Y, Peng J, Li Z, Jiang F, He Q, Ouyang L, Wu S, Fu C, et al. A prognostic signature consisting of pyroptosis-related genes and SCAF11 for predicting immune response in breast cancer. Front Med (Lausanne). 2022;9:882763. https://doi.org/10.3389/fmed.2022.882763.
Wang J, Huang Z, Lu H, Zhang R, Feng Q, He A. A pyroptosis-related gene signature to predict patients' prognosis and immune landscape in liver hepatocellular carcinoma. Comput Math Methods Med. 2022;2022:1258480. https://doi.org/10.1155/2022/1258480.
Wu J, Zhu Y, Luo M, Li L. Comprehensive analysis of pyroptosis-related genes and tumor microenvironment infiltration characterization in breast cancer. Front Immunol. 2021;12:748221. https://doi.org/10.3389/fimmu.2021.748221.
Lin W, Chen Y, Wu B, Chen Y, Li Z. Identification of the pyroptosis-related prognostic gene signature and the associated regulation axis in lung adenocarcinoma. Cell Death Discov. 2021;7(1):161. https://doi.org/10.1038/s41420-021-00557-2.
Luo J, Lai J. Pyroptosis-related molecular classification and immune microenvironment infiltration in breast cancer: a novel therapeutic target. J Cell Mol Med. 2022;26(8):2259–72. https://doi.org/10.1111/jcmm.17247.
Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30(1):207–10. https://doi.org/10.1093/nar/30.1.207.
Parkinson H, Kapushesky M, Shojatalab M, Abeygunawardena N, Coulson R, Farne A, Holloway E, Kolesnykov N, Lilja P, Lukk M, et al. ArrayExpress--a public database of microarray experiments and gene expression profiles. Nucleic Acids Res. 2007;35(Database issue):D747–50. https://doi.org/10.1093/nar/gkl995.
Diez D, Alvarez R, Dopazo A. Codelink: an R package for analysis of GE healthcare gene expression bioarrays. Bioinformatics. 2007;23(9):1168–9. https://doi.org/10.1093/bioinformatics/btm072.
Dong F, Ping P, Wang S-Q, Ma Y, Chen X-F. Identification and validation of CCL2 as a potential biomarker relevant to mast cell infiltration in the testicular immune microenvironment of spermatogenic dysfunction. Cell Biosci. 2023;13(1):94. https://doi.org/10.1186/s13578-023-01034-2.
Troyanskaya O, Cantor M, Sherlock G, Brown P, Hastie T, Tibshirani R, Botstein D, Altman RB. Missing value estimation methods for DNA microarrays. Bioinformatics. 2001;17(6):520–5. https://doi.org/10.1093/bioinformatics/17.6.520.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47. https://doi.org/10.1093/nar/gkv007.
Zhao L, Yao C, Xing X, Jing T, Li P, Zhu Z, Yang C, Zhai J, Tian R, Chen H, et al. Single-cell analysis of developing and azoospermia human testicles reveals central role of Sertoli cells. Nat Commun. 2020;11(1):5683. https://doi.org/10.1038/s41467-020-19414-4.
Nie X, Munyoki SK, Sukhwani M, Schmid N, Missel A, Emery BR, DonorConnect SJB, Mayerhofer A, Orwig KE, et al. Single-cell analysis of human testis aging and correlation with elevated body mass index. Dev Cell. 2022;57(9):1160–1176 e1165. https://doi.org/10.1016/j.devcel.2022.04.004.
Wang M, Liu X, Chang G, Chen Y, An G, Yan L, Gao S, Xu Y, Cui Y, Dong J, et al. Single-cell RNA sequencing analysis reveals sequential cell fate transition during human spermatogenesis. Cell Stem Cell. 2018;23(4):599–614 e594. https://doi.org/10.1016/j.stem.2018.08.007.
Guo J, Grow EJ, Mlcochova H, Maher GJ, Lindskog C, Nie X, Guo Y, Takei Y, Yun J, Cai L, et al. The adult human testis transcriptional cell atlas. Cell Res. 2018;28(12):1141–57. https://doi.org/10.1038/s41422-018-0099-2.
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson A, Kampf C, Sjostedt E, Asplund A, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347(6220):1260419. https://doi.org/10.1126/science.1260419.
Feig C, Kirchhoff C, Ivell R, Naether O, Schulze W, Spiess AN. A new paradigm for profiling testicular gene expression during normal and disturbed human spermatogenesis. Mol Hum Reprod. 2007;13(1):33–43. https://doi.org/10.1093/molehr/gal097.
Schulze W, Thoms F, Knuth UA. Testicular sperm extraction: comprehensive analysis with simultaneously performed histology in 1418 biopsies from 766 subfertile men. Hum Reprod. 1999;14(Suppl 1):82–96. https://doi.org/10.1093/humrep/14.suppl_1.82.
Bindea G, Mlecnik B, Tosolini M, Kirilovsky A, Waldner M, Obenauf AC, Angell H, Fredriksen T, Lafontaine L, Berger A, et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity. 2013;39(4):782–95. https://doi.org/10.1016/j.immuni.2013.10.003.
Senbabaoglu Y, Gejman RS, Winer AG, Liu M, Van Allen EM, de Velasco G, Miao D, Ostrovnaya I, Drill E, Luna A, et al. Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol. 2016;17(1):231. https://doi.org/10.1186/s13059-016-1092-z.
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF, Schinzel AC, Sandy P, Meylan E, Scholl C, et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462(7269):108–12. https://doi.org/10.1038/nature08460.
Yamanaka K, Fujisawa M, Tanaka H, Okada H, Arakawa S, Kamidono S. Significance of human testicular mast cells and their subtypes in male infertility. Hum Reprod. 2000;15(7):1543–7. https://doi.org/10.1093/humrep/15.7.1543.
Zheng W, Zhang S, Jiang S, Huang Z, Chen X, Guo H, Li M, Zheng S. Evaluation of immune status in testis and macrophage polarization associated with testicular damage in patients with nonobstructive azoospermia. Am J Reprod Immunol. 2021;86(5):e13481. https://doi.org/10.1111/aji.13481.
Bhattacharya S, Dunn P, Thomas CG, Smith B, Schaefer H, Chen J, Hu Z, Zalocusky KA, Shankar RD, Shen-Orr SS, et al. ImmPort, toward repurposing of open access immunological assay data for translational and clinical research. Sci Data. 2018;5:180015. https://doi.org/10.1038/sdata.2018.15.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13. https://doi.org/10.1093/nar/gky1131.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–50. https://doi.org/10.1073/pnas.0506580102.
Malcher A, Rozwadowska N, Stokowy T, Kolanowski T, Jedrzejczak P, Zietkowiak W, Kurpisz M. Potential biomarkers of nonobstructive azoospermia identified in microarray gene expression analysis. Fertil Steril. 2013;100(6):1686–94 e1681-1687. https://doi.org/10.1016/j.fertnstert.2013.07.1999.
Zarezadeh R, Fattahi A, Nikanfar S, Oghbaei H, Ahmadi Y, Rastgar Rezaei Y, Nouri M, Dittrich R. Hormonal markers as noninvasive predictors of sperm retrieval in non-obstructive azoospermia. J Assist Reprod Genet. 2021;38(8):2049–59. https://doi.org/10.1007/s10815-021-02176-3.
Bhushan S, Theas MS, Guazzone VA, Jacobo P, Wang M, Fijak M, Meinhardt A, Lustig L. Immune cell subtypes and their function in the testis. Front Immunol. 2020;11:583304. https://doi.org/10.3389/fimmu.2020.583304.
Caroppo E, Colpi GM. Prediction models for successful sperm retrieval in patients with non-obstructive azoospermia undergoing microdissection testicular sperm extraction: is there any room for further studies? J Clin Med. 2021;10(23). https://doi.org/10.3390/jcm10235538.
Kim ED, Barqawi AZ, Seo JT, Meacham RB. Apoptosis: its importance in spermatogenic dysfunction. Urol Clin North Am. 2002;29(4):755–65, vii. https://doi.org/10.1016/s0094-0143(02)00093-9.
Guo H, Ouyang Y, Yin H, Cui H, Deng H, Liu H, Jian Z, Fang J, Zuo Z, Wang X, et al. Induction of autophagy via the ROS-dependent AMPK-mTOR pathway protects copper-induced spermatogenesis disorder. Redox Biol. 2022;49:102227. https://doi.org/10.1016/j.redox.2021.102227.
Li Y, Zhu Z, Cui H, Ding K, Zhao Y, Ma X, Adetunji AO, Min L. Effect of zearalenone-induced ferroptosis on mice spermatogenesis. Animals (Basel). 2022;12(21). https://doi.org/10.3390/ani12213026.
Shi J, Gao W, Shao F. Pyroptosis: gasdermin-mediated programmed necrotic cell death. Trends Biochem Sci. 2017;42(4):245–54. https://doi.org/10.1016/j.tibs.2016.10.004.
Hsu SK, Li CY, Lin IL, Syue WJ, Chen YF, Cheng KC, Teng YN, Lin YH, Yen CH, Chiu CC. Inflammation-related pyroptosis, a novel programmed cell death pathway, and its crosstalk with immune therapy in cancer treatment. Theranostics. 2021;11(18):8813–35. https://doi.org/10.7150/thno.62521.
Ma J, Dong S, Lu H, Chen Z, Yu H, Sun X, Peng R, Li W, Wang S, Jiang Q, et al. The hydrogen storage nanomaterial MgH(2) improves irradiation-induced male fertility impairment by suppressing oxidative stress. Biomater Res. 2022;26(1):20. https://doi.org/10.1186/s40824-022-00266-6.
Hasani A, Khosravi A, Behnam P, Ramezani F, Eslami Farsani B, Aliaghaei A, Pirani M, Akaberi-Nasrabadi S, Abdi S, Abdollahifar MA. Non-apoptotic cell death such as pyroptosis, autophagy, necroptosis and ferroptosis acts as partners to induce testicular cell death after scrotal hyperthermia in mice. Andrologia. 2022;54(2):e14320. https://doi.org/10.1111/and.14320.
Song JL, Zhang GL. Deoxynivalenol and zearalenone: different mycotoxins with different toxic effects in the Sertoli cells of Equus asinus. Cells. 2021;10(8). https://doi.org/10.3390/cells10081898.
Lu Y, Bhushan S, Tchatalbachev S, Marconi M, Bergmann M, Weidner W, Chakraborty T, Meinhardt A. Necrosis is the dominant cell death pathway in uropathogenic Escherichia coli elicited epididymo-orchitis and is responsible for damage of rat testis. PLoS One. 2013;8(1):e52919. https://doi.org/10.1371/journal.pone.0052919.
Qian Z, Zhao Y, Wan C, Deng Y, Zhuang Y, Xu Y, Zhu Y, Lu S, Bao Z. Pyroptosis in the initiation and progression of atherosclerosis. Front Pharmacol. 2021;12:652963. https://doi.org/10.3389/fphar.2021.652963.
Wu Y, Pan B, Zhang Z, Li X, Leng Y, Ji Y, Sun K, Chen AF. Caspase-4/11-mediated pulmonary artery endothelial cell pyroptosis contributes to pulmonary arterial hypertension. Hypertension. 2022;79(3):536–48. https://doi.org/10.1161/HYPERTENSIONAHA.121.17868.
Cavalcanti MC, Steilmann C, Failing K, Bergmann M, Kliesch S, Weidner W, Steger K. Apoptotic gene expression in potentially fertile and subfertile men. Mol Hum Reprod. 2011;17(7):415–20. https://doi.org/10.1093/molehr/gar011.
Wang K, Sun Q, Zhong X, Zeng M, Zeng H, Shi X, Li Z, Wang Y, Zhao Q, Shao F, Ding J. Structural mechanism for GSDMD targeting by autoprocessed caspases in pyroptosis. Cell. 2020;180(5):941–955 e920. https://doi.org/10.1016/j.cell.2020.02.002.
Seibt TM, Proneth B, Conrad M. Role of GPX4 in ferroptosis and its pharmacological implication. Free Radic Biol Med. 2019;133:144–52. https://doi.org/10.1016/j.freeradbiomed.2018.09.014.
Russo AJ, Rathinam VAK. Lipid peroxidation adds fuel to pyr(optosis). Cell Host Microbe. 2018;24(1):8–9. https://doi.org/10.1016/j.chom.2018.06.010.
Zhu H, Santo A, Jia Z, Robert LY. GPx4 in bacterial infection and polymicrobial sepsis: involvement of ferroptosis and pyroptosis. React Oxyg Species (Apex). 2019;7(21):154–60. https://doi.org/10.20455/ros.2019.835.
Gu X, Wang Y, He Y, Zhao B, Zhang Q, Li S. MiR-1656 targets GPX4 to trigger pyroptosis in broilers kidney tissues by activating NLRP3 inflammasome under Se deficiency. J Nutr Biochem. 2022;105:109001. https://doi.org/10.1016/j.jnutbio.2022.109001.
Fan R, Sui J, Dong X, Jing B, Gao Z. Wedelolactone alleviates acute pancreatitis and associated lung injury via GPX4 mediated suppression of pyroptosis and ferroptosis. Free Radic Biol Med. 2021;173:29–40. https://doi.org/10.1016/j.freeradbiomed.2021.07.009.
Canli O, Alankus YB, Grootjans S, Vegi N, Hultner L, Hoppe PS, Schroeder T, Vandenabeele P, Bornkamm GW, Greten FR. Glutathione peroxidase 4 prevents necroptosis in mouse erythroid precursors. Blood. 2016;127(1):139–48. https://doi.org/10.1182/blood-2015-06-654194.
Seiler A, Schneider M, Forster H, Roth S, Wirth EK, Culmsee C, Plesnila N, Kremmer E, Radmark O, Wurst W, et al. Glutathione peroxidase 4 senses and translates oxidative stress into 12/15-lipoxygenase dependent- and AIF-mediated cell death. Cell Metab. 2008;8(3):237–48. https://doi.org/10.1016/j.cmet.2008.07.005.
Liu A, Li F, Xu P, Chen Y, Liang X, Zheng S, Meng H, Zhu Y, Mo J, Gong C, Zhou JC. Gpx4, Selenov, and Txnrd3 are three most testis-abundant selenogenes resistant to dietary selenium concentrations and actively expressed during reproductive ages in rats. Biol Trace Elem Res. 2022; https://doi.org/10.1007/s12011-022-03118-5.
Khalid A, Khudhair N, He H, Peng Z, Yaguang T, Guixue Z. Effects of dietary selenium supplementation on seminiferous tubules and SelW, GPx4, LHCGR, and ACE expression in chicken testis. Biol Trace Elem Res. 2016;173(1):202–9. https://doi.org/10.1007/s12011-016-0646-y.
Gong Z, Li Q, Yang J, Zhang P, Sun W, Ren Q, Tang J, Wang W, Gong H, Li J. Identification of a pyroptosis-related gene signature for predicting the immune status and prognosis in lung adenocarcinoma. Front Bioeng Biotechnol. 2022;10:852734. https://doi.org/10.3389/fbioe.2022.852734.
Kharchenko PV, Silberstein L, Scadden DT. Bayesian approach to single-cell differential expression analysis. Nat Methods. 2014;11(7):740–2. https://doi.org/10.1038/nmeth.2967.
Funding
This work was funded by the National Natural Science Foundation of China (81871199).
Author information
Authors and Affiliations
Contributions
Y.M. and X.-F.C. designed the study. F.D. did data acquisition, data analysis, and figure preparation. F.D. completed the original draft. F.D., Y.M., and X.-F.C. discussed and amended the manuscript. All authors approved the version to be published.
Corresponding authors
Ethics declarations
Ethics approval
This study is based on published datasets, and no human participate/material was involved. Therefore, no ethics approval is needed.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information

ESM 1
Figure S1. Quality control information of scRNA-seq analyses. (A) Violin plots of number of features(genes) per cell (left), number of UMI counts per cell (middle), and percentage of mitochondrial genes (right). (B) Scatter plots of mitochondrial genes versus UMI counts (left), and features(genes) versus UMI counts (right). (PNG 770 kb)

ESM 2
Figure S2. Ratios of CASP4+/CASP4- cells and related GSEA results in different testicular cells. (A),(B),(C) Proportions of CAPS4 positive (>0) endothelial cells (A), macrophages (B) and PTM cells (C), respectively in control/disease groups. (D),(E),(F) Top three enriched pathways (according to |NES|) of GSEA based on the decreasingly ranked gene list according to avg_log2FC (calculated with Findmarkers) between CASP4+ and CASP4− endothelial cells (D), macrophages (E) and PTM cells (F), respectively. (PNG 378 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dong, F., Ma, Y. & Chen, XF. Identification of a novel pyroptosis-related gene signature in human spermatogenic dysfunction. J Assist Reprod Genet 40, 2251–2266 (2023). https://doi.org/10.1007/s10815-023-02892-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10815-023-02892-y