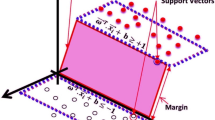
We investigated the feasibility of using surface-enhanced Raman scattering (SERS) technology combined with the AdaBoost algorithm to rapidly discriminate cervical cancer patients from hysteromyoma patients. Using Au colloids as the SERS active substrate, we recorded Raman signal measurements on serum RNA samples obtained from 35 patients diagnosed with cervical cancer and 30 patients diagnosed with hysteromyoma. Analysis of RNA SERS spectra using principal component analysis, then three principal components (PC2, PC11, and PC24) with significant differences were chosen using the independent samples t-test (p < 0.05). The distinctive peak intensities of the relevant substance, measured at 448, 519, 698, 1003, and 1076 cm–1, were found to be correlated with the substance’s alterations during the carcinogenesis process. The ideal AdaBoost classification model was developed by fi ne-tuning its parameters. The model showcased an impressive accuracy of 96.92%, exhibiting a high sensitivity of 94.28% and an exceptional specificity of 100%, as reported in the results. Compared to the linear discriminant analysis, support vector machine models, the effectiveness of classification greatly improved. The current findings indicate that serum SERS technology, combined with the AdaBoost algorithm, is anticipated to be developed into a potent screening tool for cervical cancer.
Similar content being viewed by others
References
H. Sung, J. Ferlay, R.L. Siegel, et al., Cancer J. Clin., 71, No. 3, 209–249 (2021); https://doi.org/10.3322/caac.21660.
A. Slomski, JAMA, 327, No. 9, 804 (2022); https://doi.org/10.1001/jama.2022.2188.
P. A. Cohen, A. Jhingran, A. Oaknin, et al., Lancet, 393, No. 10167, 169–182 (2019); https://doi.org/10.1016/S01406736(18)32470-X.
M. Vu, J. Yu, O. A. Awolude, et al., Curr. Probl. Cancer, 42, No. 5, 457–465 (2018); /https://doi.org/10.1016/j.currproblcancer.2018.06.003.
E. T. Fontham, A. M. Wolf, T. R. Church, et al., Cancer J. Clin., 70, No. 5, 321–346 (2020); https://doi.org/10.3322/caac.21628.
R. Catarino, P. Petignat, G. Dongui, et al., World J. Clin. Oncol., 6, No. 6, 281 (2015); https://doi.org/10.5306/wjco.v6.i6.281.
P. Rajaraman, B. O. Anderson, P. Basu, et al., Lancet Oncol., 16, No. 7, e352–e361 (2015); https://doi.org/10.1016/S1470-2045(15)00078-9.
R. Sankaranarayanan, P. O. Esmy, R. Rajkumar, et al., Lancet, 370, No. 9585, 398–406 (2007); org/https://doi.org/10.1016/S0140-6736(07)61195-7.
R. R. Jones, D. C. Hooper, L. Zhang, et al., Nanoscale Res. Lett., 14, No. 1, 1–34 (2019); https://doi.org/10.1186/s11671-019-3039-2.
X. Zhang, C. R. Yonzon, M. A. Young, et al., IEE Proc. Nanobiotechnol., 152, No. 6, 195–206 (2005); https://doi.org/10.1049/ip-nbt:20050009.
L. A. Lane, X. Qian, and S. Nie, Chem. Rev., 115, No. 19, 10489–10529 (2015); https://doi.org/10.1021/acs.chemrev.5b00265.
Y. Chen, G. Chen, X. Zheng, et al., Med. Phys., 39, No. 9, 5664–5668 (2012); https://doi.org/10.1118/1.4747269.
Y. Chen, G. Chen, S. Feng, et al., J. Biomed. Opt., 17, No. 6, Article ID 067003 (2012); https://doi.org/10.1117/1.JBO.17.6.067003.
S. Nasir, M. I. Majeed, H. Nawaz, et al., Photodiagnosis Photodyn. Ther., 33, Article ID 102152 (2021); https://doi.org/10.1016/j.pdpdt.2020.102152.
X. Wang, H. K. Wang, Y. Li, et al., Proc. NAS, 111, No. 11, 4262–4267 (2014); https://doi.org/10.1073/pnas.1401430111.
M. L. Tornesello, R. Faraonio, L. Buonaguro, et al., Front. Oncolo., 10, Article ID 150 (2020); https://doi.org/10.3389/fonc.2020.00150.
Y. He, J. Lin, Y. Ding, et al., Int. J. Cancer, 138, No. 6, 1312–1327 (2016); https://doi.org/10.1002/ijc.29618.
J. D. Driskell, A. G. Seto, L. P. Jones, et al., Biosens. Bioelectron., 24, No. 4, 917–922 (2008); https://doi.org/10.1016/j.bios.2008.07.060.
J. D. Driskell, O. M. Primera-Pedrozo, R. A. Dluhy, et al., Appl. Spectrosc., 63, No. 10, 1107–1114 (2009); https://doi.org/10.1366/000370209789553183.
A. A. Bunaciu, S. Fleschin, V. D. Hoang, et al., Crit. Rev. Anal. Chem., 47, No. 1, 67–75 (2017); https://doi.org/10.1080/10408347.2016.1209104.
X. D. Zhang, J. F. Li, Q. Q. Zhao, et al., Laser & Infrared, 38, 267–269 (2008); https://doi.org/10.1016/j.jpba.2007.11.019.
B. B. Tang, S. Y. Liu, Y. U. Zhan, et al., Exp. Therap. Med., 10, No. 1, 269–274 (2015); https://doi.org/10.3892/etm.2015.2455.
X. Dong, Z. Yu, W. Cao, et al., Front. Comp. Sci., 14, No. 2, 241–258 (2020); https://doi.org/10.1007/s11704-019-8208-z.
A. Savitzky and M. J. Golay, Anal. Chem., 36, No. 8, 1627–1639 (1964), https://doi.org/https://doi.org/10.1021/ac60319a045.
Z. M. Zhang, S. Chen, and Y. Z. Liang, Analyst, 135, No. 5, 1138–1146 (2010); https://doi.org/10.1039/b922045c.
J. D. Rodriguez, B. J. Westenberger, L. F. Buhse, et al., Analyst, 136, No. 20, 32–40 (2011); https://doi.org/10.1039/c1an15636e.
J. Hatwell, M. M. Gaber, and R. M. Atif Azad, BMC Med. Inform. Dec. Mak., 20, No. 1, 250 (2020); https://doi.org/10.1186/s12911-020-01201-2.
S. A. Sánchez-Rojo, B. E. Martínez-Zérega, E. F. Velázquez-Pedroza, et al., Rev. Mexicana de Física, 62, No. 3, 213–218 (2016).
X. Zheng, G. Wu, J. Wang, et al., Biomed. Opt. Express, 13, No. 4, 1912–1923 (2022); https://doi.org/10.1364/BOE.448121.
K . Mühlenbruch, A. Heraclides, E. W. Steyerberg, et al., Eur. J. Epidemiol., 28, No. 1, 25–33 (2013); https://doi.org/10.1007/s10654-012-9744-0.
A. C. S. Talari, Z. Movasaghi, S. Rehman, et al., Appl. Spectr. Rev., 50, No. 1, 46–111 (2015); https://doi.org/10.1080/05704928.2014.923902.
S. Qiu, Y. Xu, L. Huang, et al., Oncol. Lett., 11, No. 1, 884–890 (2016); https://doi.org/10.3892/ol.2015.3969.
D. Puchowicz and M. Cieslak, Raman Spectroscopy in the Analysis of Textile Structures, Recent Developments in Atomic Force Microscopy and Raman Spectroscopy for Materials Characterization, pp. 1–21 (2021); https://doi.org/10.5772/intechopen.99731.
Y. Li, R. Shen, H. Wu, et al., Spectrochim. Acta A: Mol. Biomol. Spectrosc., 225, Article ID 117483 (2020); https://doi.org/10.1016/j.saa.2019.117483.
Z. Huang, A. McWilliams, H. Lui, et al., Int. J. Cancer, 107, No. 6, 1047–1052 (2003); https://doi.org/10.1002/ijc.11500.
J. I. Githaiga, H. K. Angeyo, K. A. Kaduki, et al., J. Spectrosc. (2020); https://doi.org/10.1155/2020/8879985.
S. Chaichian, R. Shafabakhsh, S. M. Mirhashemi, et al., J. Cell. Physiol., 235, No. 2, 718–724 (2020); https://doi.org/10.1002/jcp.29009.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Abstract of article is published in Zhurnal Prikladnoi Spektroskopii, Vol. 91, No. 1, p. 166, January–February, 2024.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiao, Z., Wu, G., Wang, J. et al. Rapid Discrimination of Cervical Cancer from Hysteromyoma Using Label-Free Serum RNA Based on Surface-Enhanced Raman Spectroscopy and AdaBoost Algorithm. J Appl Spectrosc 91, 200–208 (2024). https://doi.org/10.1007/s10812-024-01707-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10812-024-01707-x