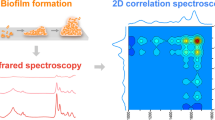
The objective of this work was to evaluate the ability of the functionally enhanced derivative spectroscopy (FEDS) algorithm to characterize the surface of microorganisms, namely, Candida albicans, by mid-IR spectroscopy and with the cellulose sensing surface technique. This work is a key stage in the study of cell–cell and cell–surface interactions between microorganisms, including the study of polymicrobial biofilms. Accordingly, C. albicans was selected as a microorganism model due to its importance in medical science and human health. Spectra were recorded in triplicate from 4000 to 500 cm–1 by the ATR technique. It was concluded that the FEDS transform of the mid-IR spectrum is a powerful analytical tool to improve spectral analysis by IR spectroscopy. In the particular case of C. albicans biofilms, it was observed that by FEDS, it is possible to deconvolute signals and achieve improved signal differentiation. For interpretation, serine, threonine, glycine, alanine, glutamic acid, proline, and N-acetyl-D-glucosamine units were taken as molecular models since these molecules have been described as the main components in the cell wall of C. albicans. In this way, it was found that the vibrational spectrum of C. albicans biofilms can be understood considering only the main components of the cell wall.
Similar content being viewed by others
References
J. M. Hornby, E. C. Jensen, A. D. Lisec, J. J. Tasto, B. Jahnke, R. Shoemaker, P. Dussault, and K. W. Nickerson, Appl. Environ. Microbiol., 67, 2982–2992 (2001).
H. H. Tuson and D. B. Weibel, Soft Matter., 9, 4368–4380 (2013).
C. R. Arciola, D. Campoccia, G. D. Ehrlich, and L. Montanaro, Adv. Exp. Med. Biol., 830, 29–46 (2015).
A. Kumar, A. Alam, M. Rani, N. Z. Ehtesham, and S. E. Hasnain, Int. J. Med. Microbiol., 307, 481–489 (2017).
A. Elbourne, J. Chapman, A. Gelmi, D. Cozzolino, R. J. Crawford, and V. K. Truong. J. Colloid Interf. Sci., 546, 192–210 (2019).
A. Alvarez-Ordoñez, D. Mouwen, M. Lopez, and M. Prieto. J. Microbiol. Methods, 84, 369–378 (2006).
J. Ojeda and M. Dittrich, Microbiol Systems Biology: Methods and Protocols, Methods in Molecular Biology, Springer Science (2012).
J. Prakash, S. Kar, C. Lin, C. Y. Chen, C. F. Chang, J. S. Jean, and T. R. Kulp, Spectrochim. Acta A: Mol. Biomol. Spectrosc., 116, 478–484 (2013).
M. Dadd, D. Sharp, A. Pettman, and C. Knowles, J. Microbiol. Methods., 41, 69–75 (2000).
W. Huang, D. Hopper, R. Goodacre, M. Beckmann, A. Singer, and J. Draper, J. Microbiol. Methods, 67, 273–280 (2006).
Z. Khatoon, C. McTiernan, E. Suuronen, T. F. Mah, and E. Alarcon, Heliyon, 4, e01067 (2018).
D. Singhalage, G. Seneviratne, S. Madawala, and I. Manawasinghe, Ceylon J. Sci., 47, 77–83 (2018).
W. Friesen and K. Michaelian, Appl. Spectrosc., 45, 50–56 (1991).
T. Vazhnova and D. Lukyanov, Anal. Chem., 85, 11291–11296 (2013).
M. Palencia, J. Adv. Res., 14, 53–62 (2018).
M. Palencia, T. Lerma, and N. Afanasjeva, Eur. Polym. J., 115, 212–220 (2019).
Y. Guan, C. J. Wurrey, and G. J. Thomas, Biophys. J., 66, 225–235 (1994).
Y. Guan and G. J. Thomas, Biopolymers, 39, 813–835 (1996).
W. Jiang, A. Saxena, B. Song, B. B. Ward, T. J. Beveridge, and S. Myneni, Langmuir, 20, 11433–11442 (2004).
A. Barth, Biochim. Biophys. Acta, 1767, 1073–1101 (2007).
S. Parker, Chem. Phys., 424, 75–79 (2013).
T. A. Lerma, S. Collazos, and A. Cordoba, J. Sci. Technol. Appl., 1, 30–38 (2016).
M. Palencia, T. A. Lerma, and A. Cordoba, J. Sci. Technol. Appl., 1, 39–52 (2016).
N. Arbelaez, T. A. Lerma, and A. Cordoba, J. Sci. Technol. Appl., 2, 75–83 (2017).
W. Volmer, D. Blanot, and M. A. De Pedro, FEMS Microbiol. Rev., 32, 149–167 (2008).
W. Lajean, J. L. López–Ribot, M. Casanova, D. Gozalbo, and J. P. Martínez, Microbiol. Mol. Biol. Rev., 62, 130–180 (1998).
E. Reyna-Beltran, C. I. Bazan, M. Iranzo, S. Mormeneo, and J. P. Luna-Arias, The Cell Wall of Candida albicans: A Proteomics View, (2019). https://doi.org/10.5772/intechopen.82348
J. Ruiz-Herrera, S. Mormeneo, P. Vanaclocha, J. Font-de-Mora, M. Iranzo, I. Puertes, and R. Sentandreu, Microbiol., 140, 1513–1523 (1994).
W. Nsangou, Comput. Theor. Chem., 966, 364–374 (2011).
E. Wiercigroch, E. Szafraniec, K. Czamara, M. Z. Pacia, K. Majzner, K. Kochan, A. Kaczor, K. Baranska, and K. Malek, Spectrochim. Acta A: Mol. Biomol. Spectrosc., 185, 317–335 (2017).
H. A. Wells and R. H. Atalla, J. Mol. Struct., 224, 385–424 (1990).
M. W. Ellzy, Computational and Experimental Studies Using Absorption Spectroscopy and Vibrational Circular Dischroism. Thesis, Drexel University, 1–333 (2006).
I. Nieduszynsky and R. H. Marchessault, Can. J. Chem., 50, 2130–2138 (1972).
A. Kovacs, B. Nyerges, and V. Izvekov, J. Phys. Chem., 112, 5728–5735 (2008).
M. E. Mohamed and A. M. A. Mohammed, Int. Lett. Chem., Phys. Astron., 10, 1–17 (2013).
L. E. Fernández, G. E. Delgado, L. V. Maturano, R. M. Tótaro, and E. L. Varetti, J. Mol. Struct., 1168, 84–91 (2018).
I. Adt, D. Toubas, J. M. Pinon, M. Manfait, and G. D. Sockalingum, Arch. Microbiol., 185, 277–285 (2006).
R. P. Hirschmann, R. N. Kniseley, and A. Fassel, Spectrochim. Acta, 21, 2125–2133 (1965).
C. V. Stephenson, W. C. Coburn, and W. S. Wilcox, Spectrochim. Acta, 17, 933–946 (1961).
F. O. Libnau, O. M. Kvalheim, A. A. Christy, and J. Toft, Vibr. Spectrosc., 7, 243–254 (1994).
Author information
Authors and Affiliations
Corresponding author
Additional information
Abstract of article is published in Zhurnal Prikladnoi Spektroskopii, Vol. 88, No. 1, p. 167, January–February, 2021.
Rights and permissions
About this article
Cite this article
Palencia, S.L., García, A. & Palencia, M. Mid-Infrared Vibrational Spectrum Characterization of the Outer Surface of Candida albicans by Functionally Enhanced Derivative Spectroscopy. J Appl Spectrosc 88, 166–180 (2021). https://doi.org/10.1007/s10812-021-01155-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10812-021-01155-x