Abstract
With the International Maritime Organization’s (IMO) International Convention for the Control and Management of Ships’ Ballast Water and Sediments now in force, determining abundance and distribution of phytoplankton inside ballast tanks is critical for successful ballast water management, particularly when assessing compliance. The relationship between the abundance and distribution of cells was examined to obtain the best representative sample of the entire phytoplankton community in ballast tanks, comparing three ballast water sampling techniques including in-line, in-tank, and Van Dorn bottle methods. Lloyd’s index, Dy, and Gini index were applied to compare methods of sample collection and determine representativeness of samples and performance of sampling methods. Phytoplankton abundance trends from live microscopy counts using fluorescein diacetate (FDA) were also compared to those using a FlowCAM on preserved samples. The phytoplankton community showed a patchy distribution inside the ballast tank and this trend was observed across all voyages. The estimated marginal mean analysis showed that in hypothetical conditions (e.g., 702 m3 of water in ballast tank and phytoplankton whole-tank abundance of 19,522 cells), the difference among the three methods was small. Conversely, statistical analysis performed on empiric abundances using a negative binomial regression model determined that the volume discharged during sampling of ballast water has an effect on the number of cells collected on a given voyage. Results of this study also confirmed that the in-line method may be a better method at collecting phytoplankton samples from ballast tanks than the in-tank or Van Dorn method, regardless of the time at which samples are collected. Finally, the number of living cells and the number of preserved cells showed similar trends for most of the voyages, despite fewer samples analyzed using FDA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The non-native species transported in ships’ ballast water are one of the major threats to marine and freshwater biodiversity globally (Hayes and Sliwa 2003). Ballast water is pumped into or discharged from ballast tanks during cargo operations at ports. When cargo is offloaded, its weight is offset with harbor or sea water that contains local aquatic organisms, which is stored in ballast tanks providing balance and stability during the subsequent ship voyage. Once at the next port, cargo is loaded and ballast water is pumped out including any aquatic organisms that survived the voyage. These non-native species can disturb local ecology through competition for resources (food, space, spawning areas, etc.), threatening fisheries and endangered species, changing habitat and food web structure, and even impacting human health (Hallegraeff 1997).
To prevent unintentional introductions, and reduce impacts in marine and freshwater ecosystems, the International Maritime Organization (IMO) developed the International Convention for the Control and Management of Ships’ Ballast Water and Sediments (hereafter known as the Convention), which entered into force in September 2017. Under the Convention, ships are required to meet the D-2 standard no later than September 2024, limiting the number of viable organisms in discharged ballast water as follows: (1) < 10 viable zooplankton individuals·m−3 ≥ 50 μm in minimum dimension; (2) < 10 viable phytoplankton cells·mL−1 ≥ 10 and < 50 μm in minimum dimension; and (3) limits on indicator microbes including < 1 colony forming unit·(cfu) 100 mL−1 of Vibrio cholera, < 250 cfu·(100 mL)−1 of Escherichia coli, and < 100 cfu·(100 mL)−1 of intestinal Enterococci (IMO 2004). Therefore, determining reliable sampling protocols to verify ships’ compliance with the D-2 standard and improving the accuracy of phytoplankton abundance estimation in ballast water is critical for mitigating the risk of introducing harmful and potentially invasive aquatic species via ballast water.
Determining the abundance and the distribution of cells inside ballast tanks is critical for successful ballast water management. Numerous studies have attempted to determine whether there is a relationship between abundance and distribution of cells (Carney et al. 2013; Rajakaruna et al. 2018) while others have measured the proportion of the distribution occupied by phytoplankton cells and/or the aggregation of cells. The number of cells can be estimated directly and related to abundance by volume, when the individuals are detected and counted without error. However, conducting a complete census is impossible for ballast water research and compliance monitoring due to the large volumes involved. Instead, the distribution of cells can be inferred from few samples collected using different tools or methods (e.g., Niskin or Van Dorn bottles, plankton nets, pumps, or the in-line method) from a limited or specified area of one or more ballast tanks. As a result, the number of individuals detected varies based on both random variation associated with sampling and differences in local abundance inside the tanks.
Variation in sampling may be due to different access points within a ballast tank as well as different methods used to collect samples. In general, sampling approaches can be summarized into two types: in-tank and in-line. The in-tank approach was used for many years to collect water samples directly from the ballast tank through opened manholes or sounding pipes (Leppäkoski et al. 2002; Roy et al. 2012). A benefit to this approach is that sampling is conducted before ballast water is discharged, allowing corrective action to be taken should the ballast water not meet D-2 standards. However, ballast water held in tanks may be only partially treated, as some treatment may occur during discharge, and access to the ballast tank through manhole openings is sometimes limited by cargo or structures inside the tanks, which can make this approach impractical (Moser et al. 2018). In addition, because aggregation is a natural process that persists in natural environments as in tanks (Murphy et al. 2002; First et al. 2013), collection of ballast water samples using the in-tank approach from different depths may not be recommended since sample volumes are not representative with respect to the total ballast tank capacity and structural volume (David and Perkovič 2004). The in-line sampling approach uses the ballast pipes and pumps within a ship’s machinery space. Using this approach, the ballast water can be sampled at the same time ballast tanks are being discharged, which may provide access to a broader proportion of the total ballast water volume and the resident phytoplankton community. Therefore, this approach is recommended to assess the D-2 standard compliance (IMO 2018).
It is well known that spatial distribution of phytoplankton is highly dependent upon smaller scale interactions between the individual organism and its environment, including spatial and behavioral responses to predators and the physical environment (e.g., aggregation in subsurface layers with light) (McManus and Woodson 2012). At the scale of a ballast tank, multiple physical and behavioral processes can define the spatial distribution of phytoplankton, making it hard to collect a representative sample. Therefore, understanding and improving the accuracy of estimates of phytoplankton abundance and describing the distribution of cells inside ballast tanks continue to be key issues in ballast water research and management.
Earlier sampling protocols have used Niskin or Van Dorn bottles (Simard et al. 2011; Roy et al. 2012) and pumps (Gollasch et al. 1998; David and Perkovič 2004) to obtain discrete and continuous (or integrated) samples, respectively. While the intent of these protocols was to maximize the information collected during ballast water sampling, the degree of sample representativeness (and ability to measure the abundance and distribution of the population) was dependent not only on the method used to obtain the sample, but also on how much volume is sampled, when and where the volume is sampled, and how many samples are collected with respect to how the organisms are distributed inside the ballast tank (Murphy et al. 2005). While both methods would ideally collect sufficient number/volume of samples in a spatially explicit manner to encompass the range of conditions present in ballast tanks, limitations are often realized due to physical tank structure, time available, and equipment capacity (Murphy et al. 2005).
More recently, the use of isokinetic facilities and connections to the ships’ main ballast discharge line (in-line method) through sampling points aim to minimize the mortality of organisms (Wier et al. 2015) and improve estimations of organisms in a representative portion of ballast water. Additionally, collecting in-line samples using an isokinetic sampling facility is expected to capture variation in organism abundance (e.g., due to stratification inside tanks) (Murphy et al. 2002; First et al. 2013) at a given time; theoretically, samples could be collected over the entire duration of ballast water discharge (time-integrated samples).
Most studies using the in-line method have focused on zooplankton research, while fewer studies have been conducted on the distribution of phytoplankton cells inside tanks (Rajakaruna et al. 2018) and the effect of different sampling methods on the accuracy and precision of measuring phytoplankton abundance. To maximize the information obtained from sampling, this study aimed to examine the relationship between the abundance and distribution of cells to obtain the best representative sample of the entire phytoplankton community in ballast tanks. Data were fit to Poisson and negative binomial distributions to describe whether the individuals are randomly or patchily distributed in space, where the size parameter k from the negative binomial distribution represents the aggregation of individuals (Lloyd 1967; Taylor et al. 1988). Relevant indices (e.g., Lloyd’s index, Dy, and Gini index) were also applied to compare methods of collection and determine representativeness of samples and performance of these sampling methods, looking at trends of individuals by depth or by volume discharged. It was hypothesized that height (for in-tank sampling) and volume discharged (for in-line sampling) would have the same magnitude of effect on phytoplankton abundance. Finally, this study also examines differences across particle-counting methods, comparing live counts from microscopy using fluorescein diacetate (FDA) to FlowCAM analysis of preserved samples.
Methodology
Sample collection
Phytoplankton samples were collected from a single tank (the same tank for every experiment) on the bulk carrier M/V TIM S. DOOL (216.6 m), immediately before and during five separate ballast discharge events at Clarkson, Ontario (Lake Ontario), between August 2015 and September 2016 (Fig. 1 and Table 1) (see also Bailey and Rajakaruna 2017 for more details). The vessel has six pairs of double bottom tanks connected with side tanks. The #3 starboard-side ballast tank was selected for study and it has a maximum capacity of 830.6 m3.
Map of the ports visited by the bulk carrier M/V TIM S. DOOL during the five voyages. The vessel filled its tank with fresh water at ports of Montreal and Hamilton. Samplings were conducted at the port of Clarkson. Ballast water uptake locations are represented by stars while ballast water discharge location is represented by a dot
Three sampling methods (i.e., Van Dorn, in-tank and in-line) were used to measure the abundance of phytoplankton (cells sized ≥ 10 μm and < 50 μm in minimum dimension) in the side and bottom of the L-shaped ballast tank. Three vertical phytoplankton samples were collected from the top of the side tank through the manhole access using a 4.2 L-Van Dorn sampler (Halltech Environmental Inc., Guelph, Ontario, Canada) to achieve a composite sample of approximately 12 L; access was restricted to the depth of the horizontal stringer plate inside the ballast tank (three samples were collected spanning a depth of approximately 3 m). The composite sample was sieved using a 10-μm Nitex mesh, and the fraction retained on the mesh was rinsed into a graduated cylinder, brought to a volume of 240 mL, and transferred into a brown Nalgene bottle using 10 mL filtered rinse water (250 mL final concentration volume); this procedure was also used to process samples for in-tank and in-line methods.
In-tank samples were collected at fixed depths through the manhole access using a Jabsco 777 Nitrile impeller pump and 2.54-cm braided PVC tubing installed in the ballast tank. For the first four voyages, in-tank samples were collected at five depths in the side tank (0.5, 2.5, 4.0, 5.6, and 7.1 m above the bottom of the tank). For the fifth voyage, two sampling points were relocated from the side tank to the double bottom to increase the overlap between the in-tank and in-line methods. A 10-L carboy was filled with ballast water for each sampling depth point. The contents of each carboy were processed and concentrated to 250 mL as described above. The in-tank and Van Dorn samples were collected within 2 h of the ship’s arrival to port.
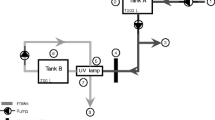
In-line, time-integrated samples were collected during ballast water discharge using a 2.54-cm isokinetic sample probe and sample collection system similar to that recommended by other studies (Cangelosi et al. 2011; Moser et al. 2018). Since ballast water was discharged by gravity during voyage 1, resulting in very low sampling flow rates compared to later voyages using ballast pumps, the in-line samples from voyage 1 were excluded from analysis. A General Electric TransPort PT878 Portable Ultrasonic liquid flow meter with clamp-on transducers was used to measure ballast water flow rate and volume in the ship’s piping upstream of the sample collection system; the sample flow rate was adjusted based on the measured flow rate in the ship’s ballast piping to maintain a sampling rate slightly lower than the calculated isokinetic sample flow rate. Samples were collected throughout the discharge event until the ballast pump lost suction or the flow rate markedly decreased (approximately 35 min after the start of ballast discharge). Sampled ballast water was collected continuously by filling multiple carboys in sequence, collecting five 10-L samples during each sampling event. Each 10-L sample was processed and concentrated to 250 mL as described above. While the in-line sample collection was standardized in terms of sample rate, sample volume, and duration, there were differences in the proportion (volume) of discharged ballast water through the main ballast pipe during sample collection, ranging from ~ 35 to 75% of the tank capacity.
Sample processing
Live counts
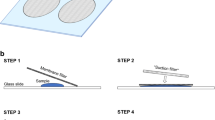
Samples were processed immediately following collection for enumeration of live phytoplankton using FDA and epifluorescence microscopy (see Adams et al. 2014; Casas-Monroy et al. 2016; Vanden Byllaardt et al. 2017). A 5-mL aliquot was removed from the 250 mL concentrated sample using a 5-mL pipette after mixing the sample by gentle inversion (5×). The subsample was placed in a 20-mL scintillation vial with 0.417 μL of FDA (working solution) and incubated for 10 min. Then 1 mL was transferred to a 1-mL Sedgewick-Rafter cell counting chamber (Wildlife Supply Company, USA). The counting chamber was systematically viewed from top to bottom with a Zeiss microscope (fitted with fluorescence excitation filter cube, based on LED modules; Carl Zeiss Canada, Ltd., Canada). The number of cells counted, in addition to the number of columns scanned, was carefully recorded to back-calculate the original cell abundance (cells·mL−1); FDA-labeled cells were counted across the whole chamber or to a maximum of 300 cells. Cell counts were completed within 20 min per sample. As all analyses were required to be completed within 3.5 h of sample collection to avoid cell death due to holding conditions, it was not possible to conduct live counts for all collected samples. Therefore, live counts were conducted on a stratified subset of samples across collection methods. All uncounted samples, as well as the remaining volume of samples after live counts, were preserved with 1% Lugol’s acid solution and transported off the ship for later laboratory analyses using a dynamic imaging particle analyzer (FlowCAM; Fluid Imaging Technologies, USA).
Phytoplankton abundance was estimated from preserved samples using the FlowCAM. Three 6- to 12-mL subsamples were removed from the 250-mL concentrated sample using a 1-mL Eppendorf pipette after mixing the sample by gentle inversion (5×). The subsamples were filtered through 53-μm Nitex mesh into a beaker and rinsed with 6 mL of deionized water for a total volume of 12–18 mL; these subsamples were called 10–53 μm fraction. The fraction retained on the 53-μm Nitex mesh was filtered through a 295-μm Nitex mesh into a scintillation vial, rinsed with deionized water and topped up to 10 mL. These subsamples were called 53–295 μm fraction. The 10–53 μm fraction was examined using an 80 μm flow cell with a 10× objective, while the ≥ 53–295 μm fraction was examined using a 300-μm flow cell with 4× objective. Both analyses were conducted using the auto-trigger mode, with approximately 30% efficiency (60-min runtime) for the 10–53 μm fraction and 68% efficiency (10-min runtime) for the ≥ 53–295 μm fraction. Imaged particles were counted using Fluid Imaging’s Visual Spreadsheet software. Visual Spreadsheet automatically enumerated ≥ 10 and < 50 μm single-celled entities and whole colonies (after manually deleting images of abiotic particles), whereas individuals ≥ 10 and < 50 μm within colonies were manually counted using NIS-Elements AR imaging software (Nikon Instruments Inc.). With all processing steps considered, approximately 1 ml of each preserved sample was analyzed, which is equivalent to 40 mL of the original 10-L ballast water sample.
Statistical analysis
Phytoplankton abundance trends from preserved samples (FlowCAM) were used to conduct all the statistical analysis. Preserved samples also were compared to live samples (epifluorescence microscopy) in order to account for differences between techniques analyzing cell abundance in ballast tanks.
Estimating tank-wide abundance
The average concentration of phytoplankton in the entire tank was calculated by combining the estimates produced by all sampling methods (Bailey and Rajakaruna 2017). This model converts measures of height (the in-tank samples collected from the vertical side tank) to measures of volume (the in-line samples collected from the ship’s main pipe during ballast discharge), for the whole tank using the following equation:
where a and b are scaling parameters, V is the total volume of the tank = 831 m3, X1 is the height of the tank (in meters) from bottom to the top, and X2 is the volume discharged (in cubic meters) by the midpoint of the in-line sample (Bailey and Rajakaruna 2017). Then, the average estimate for the entire phytoplankton population in the tank was calculated using a third degree polynomial function, obtaining the most accurate estimates based on tank height (X1) or volume discharged (X2).
Probability distributions
The variation in height and volume within each experiment were evaluated as two discrete probability distributions (negative binomial and Poisson), enabling the calculation of relevant indices. Suitable distributions were selected using Akaike’s information criterion (AIC), with parameters estimated using maximum likelihood (e.g., negative binomial: mean, μ; and dispersion, k). As the negative binomial distribution was more suitable to account for overdispersion, Generalized Linear Models (GLMs) were applied to assess correlations between phytoplankton abundance (cells mL−1) and W as height (m) and W as volume (m3) using a negative binomial regression model to observe differences among collection methods within each voyage and to compare factors such as depth and time of collection. The Akaike Information Criterion corrected for small sample size (AICc) was used to determine a set of plausible models; model averaging was used to obtain estimates of the effect of predictor variables on phytoplankton abundances.
Methods of sample collection were also compared using indices reflecting the aggregation of cells inside tanks. The Lorenz curve was applied to samples collected with different tools at various depths. The Lorenz curve—the cumulative distribution of the samples ordered by ascending size—describes the aggregation as the difference between the observed distribution and a distribution where every sample contains the same number of individuals. Indices based on the Lorenz curve include the Dy and the Gini Index (Rindorf and Lewy 2012). Both the Dy and the Gini index are derived from socioeconomics, but are also applied to natural communities (Rindorf and Lewy 2012). The Gini index (G) is defined as twice the area between the Lorenz curve and its diagonal, and has values ranging from 0 (samples are equal) to 1 (all individuals are recorded in a single sample) (Rindorf and Lewy 2012). Dy is the minimum area containing y% of the individuals and is calculated as D50, D75, and/or D95. Dy has values ranging from zero to y/100; individuals are concentrated in a few samples when Dy is zero or near zero, while all samples take equal values and only D95 is calculated when Dy is y/100. Since Dy is a function of the aggregation, high aggregation values result in a decrease in Dy.
Samples were scaled using the standardize function in R to ensure that the intercept represents the corrected mean (i.e., the predicted value for an observation which is average in every way, holding covariates at their mean values and averaging over group differences in factors). The Kendall rank coefficient was used for measuring the correlation between the response variable (preserved phytoplankton abundance across voyages) and the two explanatory variables (W, depending its special scales m and/or m3). Kendall’s rank correlation was used since its distribution under the null hypothesis is approximately normal even when the sample size is small (Brophy 1986).
A linear mixed effect model was fitted to each experiment to examine differences in empiric abundance (count variable from samples) and to create a reference-grid framework, which is an array of factor and predictor levels. The reference-grid framework was used to examine the average concentration of phytoplankton cells across the whole tank (predicted values). Subsequently, predicted values were calculated using the estimated marginal means or least squares means (Searle et al. 1980), from the linear mixed effect model for specified factors such as time, whole abundance of the tank, discharge volume and depth. The model accounted also for variability across methods as fixed effects and experiments as random effects.
The least squares means were used to examine trends of the two response variables (phytoplankton abundance from preserved samples (FlowCAM) and live samples (epifluorescence microscopy) across voyages and methods. Using the estimated marginal means from the mixed effect model, data was fitted by maximum likelihood method. Results were averaged over the levels of volume discharged and the confidence level used was 0.95. Pairwise comparisons were performed to compare slopes.
Lastly, a multi-model inference (Calcagno and de Mazancourt 2010) was run to generate all possible alternative model formulas that involved the main variables and fits them with the previously created negative binomial regression. The outcome for each voyage was the best model—selected among 108,600 models—with the relative importance of variables with main effect on the response variable. Variables that were retained in the model selection procedure were assessed for interaction. GLMs using the negative binomial function and model averaging were performed using the glmulti package. All the statistical analyses were performed using R software programming (3.6 version, R Development Core Team, 2019) and α = 0.05 was used to define statistical significance. Only variables that significantly influenced the response variable (number of cells mL−1) are presented in this study.
Results
Based on AIC values, negative binomial distributions were the most suitable distribution to represent the abundance of phytoplankton (cells mL−1) in ballast tanks across voyages, indicating that cells had an aggregate pattern inside the tank. Strong right skew was observed for voyages 2 and 3 since dispersion values of (k) were smaller than for the other voyages (Table 2). In general, parameter values for voyages 2 and 3 indicated overdispersion more than for other voyages (e.g., for voyage 2: parameters were k = 8.78 and u = 16.02, while for voyage 4 were k = 22.05 and u = 77.42; Table 2).
The negative binomial regression model revealed that factors such as height could influence the number of cells captured by each sampling method, for a given voyage. During voyage 1, the model confirmed that a higher number of cells were captured at the top of the tank by the Van Dorn method compared to the in-tank method (p < 0.001; Table 3). During voyage 2, results showed that the height was not a significant factor to explain the difference in the number of cells captured by the three methods, although the model showed that the in-line method was able to capture a significantly higher number of cells compared to the in-tank and the Van Dorn (Table 3). During voyage 3, the height was an important factor as the model revealed significant differences particularly between in-tank and in-line methods. Surprisingly, the model showed no difference between the Van Dorn and the in-line method, while significantly fewer cells were captured by the pump. During voyage 4, the in-tank method captured the highest number of cells followed by the in-line and Van Dorn methods (Table 3), while for voyage 5, it was followed by the Van Dorn and in-line methods (p < 0.0001; Table 3). During these two last voyages, the model showed that the number of cells was greatly influenced by height (p < 0.0001; Fig. 2; Table 3).
Estimates of phytoplankton abundance in the ballast tank for a in-tank and b in-line sampling methods, by voyage (voyage 1 = red; voyage 2 = green; voyage 3 = black; voyage 4 = blue; voyage 5 = orange). Phytoplankton abundance trends were analyzed using a linear regression with W = x1. The Van Dorn bottle was not plotted as there exists only 1 measurement for each voyage
Similarly, the model also showed that W (m3) during each voyage influenced the number of cells captured by each method, except in voyage 2 where the model did not show any statistical difference (Table 4). In voyage 3, the model showed that a significantly higher number of cells were captured with the pump, while comparing Van Dorn and in-line methods, the model did not show any significant effect. Surprisingly, for voyage 5, the model showed that the in-tank and Van Dorn methods collected significantly higher number of cells compared to the in-line method (p < 0.0001, p < 0.001, respectively; Table 4). When comparing within voyages, the model showed the last sample with the in-tank method captured significantly more cells than the other four measurements (estimate = 0.320, S.E. = 0.0641, p < 0.0001) in voyage 1. Similarly, for voyage 2, the last sample collected with the in-line method had a higher number of cells compared with the other measurements (estimate = 0.933, S.E. = 0.328, p < 0.001).
The Kendall rank correlation did not show any correlation between the response variable (phytoplankton abundance across voyages) and the two explanatory variables (W, as m and/or m3). For the in-tank method, the correlation between phytoplankton abundance and W (m) was not significant (Z = 0.31; Tau = − 0.047; p = 0.76), while similar results were found when comparing preserved phytoplankton abundance and W (m) using the in-line method (Z = − 066; Tau = − 0.11; p = 0.51). Similar results were found when comparing phytoplankton abundance with W (m3) for the same sampling methods (Z = − 014; Tau = − 0.02; p = 0.89 and Z = − 0.55; Tau = − 0.09; p = 0.58, respectively). Generally, for the in-line and in-tank methods, a Kendall’s coefficient was insignificant, negative and close to zero.
The Lorenz curve measures
The Lorenz curve was utilized to compare phytoplankton abundance (%) for each voyage against the proportion of the volume sampled by each sampling method (Fig. 3). Based on the Gini index, the samples with the most equal distribution of organisms were the ones collected using the Van Dorn method in voyage 1, whereas the samples with the most unequal distribution of organisms were from the Van Dorn method for voyage 2 and in-line method for voyage 3 (Table 5). Generally, samples with Gini index values closer to zero had higher mean phytoplankton abundances (e.g., Van Dorn bottle for voyage 1), while samples with values closer to one had lower mean abundances (e.g., in-line method for voyage 3).
Lorenz curve plots of relative phytoplankton abundance (%) for each voyage against the proportion of the volume sampled (%) of each sampling method during a voyage 1, b voyage 2, c voyage 3, d voyage 4, and e voyage 5. The Van Dorn bottle, in-tank, and in-line sampling methods are denoted by the red, green, and blue dashed lines, respectively
The Atkinson measure ranged from 0.002 to 0.69 for the whole dataset. The Van Dorn bottle had the lowest values out of all the sampling methods for voyages 1, 3, 4, and 5. Surprisingly, this method showed the highest values during voyage 2. On the other hand, the in-tank and in-line methods had the highest values for all voyages, except for voyage 2. Both methods followed a similar trend in Dy among voyages, but their highest values occurred in different voyages; voyage 1 for in-tank (0.047) and voyage 3 for in-line (0.069).
Trends for methods effectiveness
The sampling methods captured phytoplankton at different heights in the water column. Phytoplankton were only captured within the first two meters of the water column using the Van Dorn method, across the entire water column using the in-tank method, and at the bottom of the water column (during deballasting operations) using the in-line method. The estimated means for in-line and in-tank methods were 67.81 cell mL−1 and 65.01 cell mL−1, respectively, while the Van Dorn was higher than the other two methods (91.19 cell mL−1). A pairwise comparison showed that in-line and in-tank methods did not provide markedly different estimates of cell abundance, but were significantly lower than the Van Dorn method (Table 6 part A). When results were averaged over the entire tank volume using the model set at 702 m3 ballast water and 19,522 cells, the expected values were 68 cell mL−1 for in-line, 65 cell mL−1 for in-tank and 91 cell mL−1 for Van Dorn, and difference among methods was smaller (Table 6 part B).
Live vs. preserved counts
Counts measured by epifluorescence microscopy using FDA and FlowCAM were used to analyze trends on live and preserved phytoplankton abundance (cells mL−1) for the five voyages. Live cell concentrations in ballast ranged between 11 and 51 cells mL−1 (mean per voyage as estimated by microscopy). There was a highly significant difference between counting techniques for voyages 1 (estimate = 128 ± SE = 6.1, df = 194, p value < 0.0001), 4 (estimate = 28 ± 5.6, df = 194, p value < 0.0001), and 5 (estimate = 41 ± SE = 5.6, df = 194, p value < 0.0001). The linear mixed effect model showed that live counts follow the same trends as preserved counts, although the FlowCAM technique captured a higher number of cells for all voyages compared to the epifluorescence microscopy method (Fig. 4).
Phytoplankton abundances (cells·mL−1) estimated using epifluorescence microscopy with FDA (live samples) versus FlowCAM (preserved samples). Phytoplankton samples were obtained using a three sampling methods (Van Dorn bottle, in-tank, and in-line), b over five voyages. Linear trends were fitted to the data for both cell counting techniques. Purples bars indicate confidence intervals while red arrows indicate statistical significance (significant when not overlapping) with an α = 0.05
Phytoplankton community composition inside the tank was characterized under the low resolution given by the FlowCAM (when using natural preserved samples with Lugol’s acid instead of monoculture live samples). Voyage 1 was characterized by the presence of filaments of Oscillatoria, colonies of Chaetoceros and Fragilaria, while voyage 2 was characterized by the presence of Pediastrum and Staurastrum. Voyage 3 and 4 were characterized by the presence of Asterionella, Tabellaria, Synedra, Anabaena, and Coelastrum, while voyage 5 had predominantly Pediastrum and Stephanodiscus. For all voyages, cells of Cosmarium and Ceratium were common.
Model selection
The multi-model inference analysis resulted in selection of the best model that explained the variability of the data within each voyage. Variables retained for each voyage included time, method of collection, discharged volume (W (m), W (m3)), and sampling depth. Age of ballast water was evaluated across voyages. Although variables were retained with the highest relative importance, not all of them were retained for the same voyage since interactions between factors also accounted for some variability of data (Fig. 5). The best model for each voyage was selected among the 108,600 models that consisted of combinations of seven variables (see Table 7). Out of seven variables, only three (W(m3), water age, and sampling method) were retained in the final model set, based on their AIC, for a given voyage, for understanding of the system and predictions. Variables with the largest effect on the response variable are included in Fig. 5. Table 7 shows the best models for each voyage based on their AICc, accounting also for interactions between factors.
Model-averaged importance of variables retained for each model selection for each voyage. The importance value for a particular variable is equal to the sum of the weights/probabilities for the models in which the variable appears. Only variables with a high importance value (large weights) were plotted
Discussion
The present study was conducted in order to determine how, when and where sampling should be conducted to collect the most representative sample of phytoplankton from a ballast tank. The main findings of this research were (1) The negative binomial model fitted best to phytoplankton abundance data (number of cells mL−1), indicating that the cells had a patchy distribution inside the ballast tank; in other words, collecting a small instantaneous sample may not be representative. The lack of a significant correlation between abundance and W (m and/or m3) suggests that W did not influence the variability in the abundance of phytoplankton in terms of depth or volume, and patchiness is likely due to other physical factors beyond the scope of this paper. (2) The estimated marginal means analysis showed that under hypothetical conditions (e.g., with 702 m3 of water in ballast tank with phytoplankton whole-tank abundance of 19,522 cells), the difference between the three sampling methods is small. (3) The statistical analysis performed using the negative binomial regression model determined that the volume of discharged ballast water influenced significantly the number of cells collected on a given voyage. These results indicate that the in-line method may be a better method for collecting phytoplankton samples from ballast tanks than the in-tank or Van Dorn methods, regardless of the time at which samples are collected, since phytoplankton showed a patchy distribution. (4) The estimated marginal means analysis also showed that trends between the epifluorescence microscopy and FlowCAM counts are similar, although the FlowCAM technique can report a higher number of cells. Particularly, for voyages 1, 4, and 5, the number of cells were significantly higher.
Few studies have analyzed the distribution of particles between 10 and 50 μm in order to parameterize the performance of different mathematical models (Rajakaruna et al. 2018). Rather, the aim of this study was to address particular questions on how to optimize the collection of future phytoplankton samples based on available methods. Different factors influencing the distribution of phytoplankton cells inside tanks were also considered as well as limitations of each sampling method. Findings from the present study should be taken into account when evaluating whether or not a ship meets the IMO D-2 standard based on the number of viable cells mL−1.
Historically, ballast water sampling methods included Van Dorn bottles (Klein et al. 2009; Roy et al. 2012; Casas-Monroy et al. 2016) and non-submersible pumps (Murphy et al. 2002) with connected tubing at different depths. The Van Dorn bottle has been used to collect samples from a given location and depth(s) throughout the water column in the tank. However, for this study, the Van Dorn method was restricted to the uppermost portion of the water column, as a structural platform within the tank prevented the device from reaching the middle or bottom portions of the tank. Under such conditions, this method produced estimates of phytoplankton cells that were significantly higher than those observed by the in-tank or in-line methods. Furthermore, for voyages 1, 4, and 5, the Van Dorn indices seemed to be closer to the perfect line of the Lorenz curve, indicating the same number of cells across samples. Note that the number of cells collected using the Van Dorn method was obtained from a single measurement, while the number of cells collected using the in-tank and in-line methods was the average estimated concentration of at least five measurements. In general, the Van Dorn method had the highest dispersion and standard error; therefore, the Van Dorn bottle is the least suitable method to get a representative sample (Tables 4 and 5).
To our knowledge, there are few studies using a pump to sample phytoplankton at several depths in a ballast water tank using tubing installed into the double bottom tank (but see Rigby and Hallegraeff 1994). Similar sampling using a non-submersible pump with connected tubing at different depths has been used in several studies previously, mostly to sample zooplankton (e.g., Murphy et al. 2005; Ruiz and Smith 2005). Pump sampling enables collection of more accurate sample volumes compared to inserting samplers and tubing through the sounding pipes, which some studies have used to collect up to 10 L of ballast water for phytoplankton analysis (David and Perkovič 2004; David et al. 2007; Quilez-Badia et al. 2008). One of the main disadvantages with non-submersible pumps is the potential destruction of cells, resulting in difficult taxonomic identification and negative impacts on cell viability, which is particularly problematic for compliance assessments. Unexpectedly, for voyage 5, the in-tank method captured the highest number of cells compared to the Van Dorn and in-line methods (Table 6). Therefore, including the deepest part of the tank in the sampling procedure may increase the probability of obtaining an accurate representation of the biological community in the tank.
The in-line method (i.e., collection of time-integrated samples from ships’ ballast water discharge line) is a relatively new method for ballast water sampling (Cangelosi et al. 2011; Briski et al. 2014, 2015; Bailey and Rajakaruna 2017). The IMO Guidelines for Ballast Water Sampling (G2) recommend this method for monitoring ships’ compliance of Regulation D-2 (see Schillak and Stehouwer 2013; Drake et al. 2014). Additionally, the Convention requires ships to install a sampling port after their ballast water treatment system for collecting treated ballast water samples. When using the in-line method for monitoring compliance, samples should be collected over the duration of ballast discharge (i.e., time-integrated samples) to ensure that representative samples are obtained. For this study, it was not possible to collect samples during the entire discharge of the tank volume due to the tank structural design and loss of pump pressure (see Bailey and Rajakaruna 2017).
The multi-model inference analysis showed that time (volume discharged) was not a significant factor for explaining the variation in phytoplankton abundance (Fig. 5). On the other hand, this model determined that the age of ballast water was an important factor across voyages. Voyages with the age of ballast water of less than 3 days had the highest number of viable organisms. Similarly, Burkholder et al. (2007) concluded that ballast water that was less than 6 days old was positively correlated with high numbers of viable organisms.
Overall, the sampling method was the second-most significant factor for explaining phytoplankton abundance variation. In general, the in-tank and in-line methods reported the highest numbers of cells per mL. Interestingly, differences in cell numbers were important for voyage 1, when the in-line method was not taken into account; but particularly for voyages 4 and 5, which included the in-line method, this variable was considered only in fourth level by the multi-model analysis. Furthermore, under ideal conditions, for example with at least 702 m3 of water in the ballast tank and with an abundance of 19,500 cells, differences among the three methods are smaller at different time levels. In contrast, using empirical abundance and given the patchy distribution of phytoplankton, the in-line method may capture the same numbers of cells at the beginning, middle, and end of deballasting operations. Thus, a representative sample may be collected at any time between the beginning and end of deballasting operations in order to capture patchiness of the phytoplankton distribution.
Addressing the question on where sampling should be conducted, the multi-model inference analysis determined that variables such as volume and depth were moderately significant at explaining phytoplankton abundance variation. Thus, the sampling method has more relative importance than the location/timepoint of discharge where the water is collected. Additionally, the sampling location was highly dependent on the age of ballast water since there was a medium relative importance for the interaction between age of ballast water: W (m3).
In this study, the data used for most of the analyses was obtained from preserved samples, while the D-2 standard is based on the maximum number of viable cells per mL for the category of organisms ≥ 10 and < 50 μm. A principal component analysis (not included in this study) and the comparison between the number of living cells and the number of preserved cells showed similar trends for most of the voyages, despite differences in the number of samples. Previous studies have found that automated tools tend to underestimate particle concentration compared to microscopy counts (Kydd et al. 2018), while others have shown that the FlowCAM can generate higher number of particles per sample (Alvarez et al. 2014). Findings have shown that trends between both techniques are similar. However, discrepancy arose because (1) the number of subsamples analyzed was not the same. For the same measurement of volume discharged and/or height, one sample was analyzed using epifluorescence microscopy while three subsamples were analyzed using the FlowCAM. (2) Data acquisition, while Lugol’s acid preservative can improve cellular image capture efficiency (Graham et al. 2018), some heterotrophic and mixotrophic cells counted using FDA can have very faint green fluorescence, producing false negatives (Steinberg et al. 2011).
Conclusions
The goal of the present study was to identify sources and statistical challenges when sampling a non-random and low concentration phytoplankton population in ballast water. Both in-tank and in-line methods were closer to the true abundance of the tank, although the in-line method may better account for the patchiness distribution of phytoplankton during deballasting operations and across voyages because the method is less time-dependent. Therefore, it can be used as the recommended method to collect a sample that represents the whole ballast tank. There are also some logistic advantages to use the in-line method; for example, it does not require ships to be visited in advance to prepare the tank for sampling; it requires less equipment to be taken on board the ship and the equipment can be set up minutes before sampling and can be used to sample tanks with different shapes. One problem encountered in this investigation was accounting for the overlap at the bottom of the tank when installing the hoses for the in-tank method, in order to sample the same sector and compare samples between methods. Overlapping measurements was possible for only the final voyage, with the in-tank method capturing a greater number of cells than the in-line method; this was likely due to the patchy distribution of cells in the entire tank, increasing the chance to sample a higher number of cells at different depths or times.
References
Adams JK, Briski E, Ram JL, Bailey SA (2014) Evaluating the response of freshwater organisms to vital staining. Manag Biol Invas 5:197–208
Alvarez E, Moyano M, Lopez-Urrutia A, Nogueira E, Scharek R (2014) Routine determination of plankton community composition and size structure: a comparison between FlowCAM® and light microscopy. J Plankton Res 36:170–184
Bailey SA, Rajakaruna H (2017) Optimizing methods to estimate zooplankton concentration based on generalized patterns of patchiness inside ballast tanks and ballast water discharges. Ecol Evol 7:9689–9698
Briski E, Linley RD, Adams JK, Bailey SA (2014) Evaluating efficacy of a ballast water filtration system for reducing spread of aquatic species in freshwater ecosystems. Manag Biol Invas 5:245–253
Briski E, Gollasch S, David M, Linley RD, Casas-Monroy O, Rajakaruna H, Bailey SA (2015) Combining ballast water exchange and treatment to maximize prevention of species introductions to freshwater ecosystems. Environ Sci Technol 49:9566–9573
Brophy AL (1986) An algorithm and program for calculation of Kendall’s rank correlation coefficient. Behav Res Meth Ins C 18:45–46
Burkholder JM, Hallegraeff GM, Melia G, Cohen A, Bowers HA, Oldach DW, Parrow MW, Sullivan MJ, Zimba PV, Allen EH, Kinder CA, Mallin MA (2007) Phytoplankton and bacterial assemblages in ballast water of U.S. military ships as a function of port of origin, voyage time, and ocean exchange practices. Harmful Algae 6:486–518
Calcagno V, de Mazancourt C (2010) Glmulti: an R package for easy automated model selection with (generalized) linear models. J Stat Softw 34:1–29
Cangelosi A, Schwerdt T, Mangan T, Mays N, Prihoda K (2011) A ballast discharge monitoring system for Great Lakes relevant ships: a guidebook for researchers, ship owners, and agency officials. Washington, DC, USA: Great Ships Initiative, Northeast-Midwest Institute
Carney K, Basurko OC, Pazouki K, Marsham S, Delany JE, Desai DV, Anil AC, Mesbahi E (2013) Difficulties in obtaining representative samples for compliance with the ballast water management convention. Mar Pollut Bull 68:99–105
Casas-Monroy O, Chan PS, Linley RD, Vanden Byllaardt J, Kydd J, Bailey SA (2016) Comparison of three techniques to evaluate the number of viable phytoplankton cells in ballast water after ultraviolet irradiation treatment. J Appl Phycol 28:2821–2830
David M, Perkovič M (2004) Ballast water sampling as a critical component of biological invasions risk management. Mar Poll Bull 49:313–318
David M, Gollasch S, Cabirni M, Perkovič M, Bosnjak D, Virgilio D (2007) Results from the first ballast water sampling study in the Mediterranean Sea - the port of Koper study. Mar Poll Bull 54:53–65
Drake LA, Moser CS, Robbins-Wamsley SH, Riley SC, Wier TP, Grant JF, Herring PR, First MR (2014) Validation trials of a shipboard filter skid (p3SFS) demonstrate its utility for collecting living zooplankton. Mar Poll Bull 79:77–86
First MR, Robbins-Wamsley SH, Riley SC, Moser CS, Smith GE, Tamburri MN, Drake LA (2013) Stratification of living organisms in ballast tanks: how do organism concentrations vary as ballast water is discharged? Environ Sci Technol 47:4442–4448
Gollasch S, Dammer M, Lenz J, Andres HG (1998) Non-indigenous organisms introduced via ships into German waters. In Carlton, J. (Eds.), Ballast water: ecological and fisheries implications. ICES Coop Res Rep pp 50-64
Graham MD, Cook J, Graydon J, Kinniburgh D, Nelson H, Pilleci S, Vinebrooke RD (2018) High-resolution imaging particle analysis of freshwater cyanobacterial blooms. Limnol Oceanogr 16:669–679
Hallegraeff GM (1997) Transport of toxic dinoflagellates via ships' ballast water: bioeconomic risk assessment and efficacy of possible ballast water management strategies. Mar Ecol Prog Ser 168:297–309
Hayes KR, Sliwa C (2003) Identifying potential marine pests––a deductive approach applied to Australia. Mar Pollut Bull 46:91–98
International Maritime Organization. (2004). International convention for the control and management of ships ballast water and sediments. International Maritime Organization, IMO, London, p. 36 (Adopted on 13 February 2004)
International Maritime Organization (2018) Ballast water management convention and BWMS code with guidelines for implementation, 2018 edn. IMO Publishing, London
Klein G, Kaczmarska I, Ehrman JM (2009) The diatom Chaetoceros in ships’ ballast waters-survivorship of stowaways. Acta Bot Croat 68:325–338
Kydd J, Rajakaruna H, Briski E, Bailey SA (2018) Examination of a high resolution laser optical plankton counter and FlowCAM for measuring plankton concentration and size. J Sea Res 133:2–10
Leppäkoski E, Gollasch S, Olenin S (2002) Invasive aquatic species of Europe: distribution, Impacts and Managements. Kluwer Academic Publishers, Norwell MA, USA. 583p
Lloyd M (1967) Mean crowding. J Anim Ecol 36:1–30
McManus MA, Woodson CB (2012) Plankton distribution and ocean dispersal. J Exp Biol 215:1008–1016
Moser CS, First MR, Wier TP, Riley SC, Robbins-Wamsley SH, Molina V, Grant JF, Drake LA (2018) Design and validation of a ballast water compliance sampling device for shipboard use. Manag Biol Invasions 4:497–504
Murphy KR, Ritz D, Hewitt CL (2002) Heterogeneous zooplankton distribution in a ship's ballast tanks. J Plankton Res 24:729–734
Murphy K, Coble P, Boehme J, Field P, Cullen J, Perry E, Moore W, Ruiz G (2005) mid-ocean ballast water exchange: approach and methods for verification. Final Report to the US Coast Guard, Research and Development Center 124 pp + app
Quilez-Badia G, McCollin T, Josefsen KD, Vourdachas A, Gill ME, Mesbahi E, Frid CLJ (2008) On board short-time high temperature heat treatment of ballast water: a field trial under operational conditions. Mar Pollut Bull 56:127–135
R Development Core Team (2019). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org.
Rajakaruna H, Vanden Byllaardt J, Kydd J, Bailey SA (2018) Modeling the distribution of colonial species to improve estimation of plankton concentration in ballast water. J Sea Res 133:166–176
Ricciardi A, Hoopes MF, Marchetti MP, Lockwood JL (2013) Progress toward understanding the ecological impacts of non-native species. Ecol Monogr 83:263–282
Rigby GR, Hallegraeff GM (1994) The transfer and control of harmful marine organisms in shipping ballast water: behavior of marine plankton and ballast water exchange on the MV “Iron Whyalla”. J Marine Env 1:91–110
Rindorf A, Lewy P (2012) Estimating the relationship between abundance and distribution. Can J Fish Aquat Sci 69:382–397
Roy S, Parenteau M, Casas-Monroy O, Rochon A (2012) Coastal ship traffic: a significant introduction vector for potentially harmful dinoflagellates in eastern Canada. Can J Fish Aquat Sci 69:627–644
Ruiz, G. and Smith, G. (2005). Biological study of container ships arriving to the Port of Oakland. Final report. Port of Oakland
Schillak L, Stehouwer P-P (2013) Sampling and analysis of ballast water onboard ships for compliance analysis: report of an international research and development project; Global R & D Forum and Exhibition on Ballast Water Management, Conference Proceedings, (pp. 74-78) Busan, Korea
Searle SR, Speed FM, Milliken GA (1980) Population marginal means in the linear model: an alternative to least squares means. Am Stat 34:216–221
Simard N, Plourde S, Gilbert M, Gollasch S (2011) Net efficiency of open ocean ballast water exchange on plankton communities. J Plankton Res 33:1378–1395
Steinberg MK, Lemieux EJ, Drake LA (2011) Determining the viability of marine protists using a combination of vital, fluorescent stains. Mar Biol 158:1431–1437
Taylor LR, Perry JN, Woiwod IP, Taylor RAJ (1988) Specificity of the spatial power-law exponent in ecology and agriculture. Nature 332:721–722
Vanden Byllaardt J, Adams KA, Casas-Monroy O, Bailey SA (2017) Examination of an indicative tool for rapidly estimating viable organism abundance in ballast water. J Sea Res 133:29–35
Wier TP, Moser CS, Grant JF, First MR, Riley SC, Robbins-Wamsly SH, Drake LA (2015) Sample port design for ballast water sampling: refinement of guidance regarding the isokinetic diameter. Mar Poll Bull 98:148–155
Acknowledgments
The authors thank Robert D. Linley, Jocelyn Kydd, and Julie Vanden Byllaardt for conducting sample collection and analysis. Dawson Ogilvie gave constructive comments on an earlier draft and Celeste Remillard produced the map of ballast water operations. Finally, the authors thank reviewer’s comments that improved the manuscript.
Funding
This study was funded by Transport Canada and Fisheries and Oceans Canada. The authors gratefully acknowledge the ship crew, operators, and owners for their participation and support of this research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Casas-Monroy, O., Rajakaruna, H. & Bailey, S.A. Improving estimation of phytoplankton abundance and distribution in ballast water discharges. J Appl Phycol 32, 1185–1199 (2020). https://doi.org/10.1007/s10811-019-02034-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-019-02034-x