Abstract
Mnemiopsis leidyi A. Agassiz, 1865 is an invasive ctenophore that has spread to many areas of the Eurasian seas in recent decades and is one of the 100 most dangerous species in the world. This species was first observed in the northern Adriatic Sea in 2005 and then disappeared until 2016, when its first bloom was recorded. After that, it bloomed every summer in the area, causing severe damage to artisanal fisheries. Given the lack of genetic data for the northern Adriatic, here we compare the genetic diversity and phylogenetic relationships of M. leidyi populations sampled in 2016, 2018 and 2021 on the north-eastern coast of Italy with native populations and those introduced in other basins using two molecular markers: CytB and ITS. The CytB haplotype found in the samples from the northern Adriatic was identical to that found in the Gulf of Mexico and the Black Sea/Mediterranean. ITS analysis revealed 11 alleles, including 8 novel ones. The presence of a panmictic population in the northern Adriatic and the lower genetic variability compared to the native populations suggest that M. leidyi has been introduced into the Adriatic more than once and the species recently expanded in this area.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Mediterranean Sea has an enormously rich native biodiversity (Coll et al., 2010) that is also among the most threatened in the world due to a number of human related pressures including overfishing, pollution, habitat fragmentation and loss, rising water temperature and acidity, global expansion and intensification of shipping routes and introduction of alien invasive species (Molnar et al., 2008; Coll et al., 2010; Templado, 2014; Corriero et al., 2016; Fernandes et al., 2017; Giangrande et al., 2020).
Non-indigenous species introduced in the Mediterranean Sea are nowadays more than 1000, of which 751 were recently recognized as established (Zenetos et al., 2022). Arrival of non-indigenous species in coastal marine environments can profoundly reshape local/native communities specially when some of them became invasive (Katsanevakis et al., 2014a, b). Among these latter, one of the most dangerous is the comb jelly Mnemiopsis leidyi A. Agassiz, 1865 (Fig. 1a), which is included in the list of the 100 of the world’s worst invasive species by the International Union for Conservation of Nature (IUCN) (Lowe et al., 2000). The danger posed by this species does not lie in its direct physical threat to humans, but in its negative impact on the functioning and services of the ecosystem in invaded areas. In particular, the ecological importance of M. leidyi has been recognised in recent decades following its arrival in the Black Sea and surrounding areas, where fishery collapses and ecosystem disturbances have been mainly associated with the introduction of this ctenophore (Kideys et al., 2005; Oguz et al., 2008). In the Mediterranean, the main recognised damage caused by this species to date is due to its negative impact on artisanal fisheries in lagoons, where massive aggregation of ctenophores can fill and clog fishermen’s nets (Fig. 1b) (Diciotti et al., 2016; Marchessaux et al., 2023; Piccardi et al., 2024).
The lobate ctenophore M. leidyi, commonly known as the “sea walnut”, is a marine holoplanktonic comb jellyfish native to the estuarine and coastal waters of the Atlantic coast of America (from Massachusetts in the USA down to the Gulf of Mexico and Argentina) (GESAMP, 1997). Mnemiopsis leidyi is not only capable of establishing over a wide range of temperatures and salinities (Costello et al., 2012; Shiganova, 2020), but it is also considered a highly successful invader (Costello et al., 2012), due in part to its extremely rapid growth rate combined with extremely high and abundant reproduction (Purcell et al., 2001; Malej et al., 2017; Jaspers et al., 2018), often resulting in large blooms. M. leidyi is a simultaneous hermaphrodite capable of self-fertilisation (Sasson & Ryan, 2016), which reproduces at a very small body size (~ 1 mm) and spawns continuously from this point onwards under suitable environmental conditions (Edgar et al., 2022) and can release up to 10,000 eggs per day as an adult (Baker & Reeve, 1974; Jaspers et al., 2015; Malej et al., 2017). It is a strict carnivore that feeds at high rates on zooplankton (Purcell et al., 2001; Finenko et al., 2006; Marchessaux et al., 2021) and can thus compete with planktivorous fish species and fish larvae (Oguz et al., 2008; Budiša et al., 2021), causing a cascade effect in planktonic food webs (Tiselius & Møller, 2017; Paliaga et al., 2021). Recently, it has been shown that M. leidyi can even resort to cannibalism when food is scarce (Javidpour et al., 2020).
The first sighting of M. leidyi in Eurasia dates back 40 years, when this species was observed in the Black Sea in the early 1980s, probably introduced with ballast water (Vinogradov et al., 1989). It was subsequently introduced into the adjacent Sea of Azov (Studenikina et al., 1991), the Caspian Sea (Ivanov et al., 2000), the Sea of Marmara (Shiganova, 1997), the northern Aegean Sea (Shiganova et al., 2004), the Levantine Sea (Galil et al., 2009), the western Mediterranean (Boero et al., 2009) and the Adriatic Sea (Shiganova & Malej, 2009; Malej et al., 2017). However, the spread of the species in northern Europe occurred more recently than in the Mediterranean: the first sighting dates back to 2005, when individuals were first recorded in Danish waters in summer/autumn (Tendal et al., 2007) and in the English Channel (port of Le Havre) in summer/autumn (Antajan et al., 2014). Subsequently, it was found in several locations in the Baltic Sea (Hansson, 2006; Javidpour et al., 2006) and the North Sea (Faasse & Bahya, 2006; Boersma et al., 2007) as well as in the Oslo Fjord (Norway) (Oliveira, 2007).
Although there was a consensus that transportation via ballast water was the main vector for the introduction of M. leidyi into European waters, genetic studies were needed to clarify the origin and routes of introduction (Reusch et al., 2010; Ghabooli et al., 2011, 2013; Bolte et al., 2013; Bayha et al., 2015). Studies in which various molecular approaches were applied (Reusch et al., 2010; Ghabooli et al., 2011; Jaspers et al., 2018, 2021) indicate that M. leidyi was introduced to Eurasia via at least two pathways: one in the 1980s/1990s from an area of origin in or near the Gulf of Mexico (e.g. Florida) into the Black Sea with subsequent spread across the Pontocaspian Basin and a second at the beginning of the current millennium from New England (possibly Narragansett Bay) into the Baltic and North Sea. Several introductions from a combination of Black Sea and native sources (Gulf of Mexico) have been proposed for the Mediterranean Sea (Bolte et al., 2013; Ghabooli et al., 2013).
Several specimens of this invasive ctenophore were first detected in the Gulf of Trieste (northernmost part of the Adriatic Sea) in October 2005, presumably via ballast water (Shiganova & Malej, 2009). As suggested by Malej et al. (2017), they probably failed to become established due to low propagule pressure (e.g. low number of individuals released, fragility of released individuals, etc.). In fact, M. leidyi has not been detected in the Adriatic Sea for more than 10 years, since the summer of 2016, when blooms occurred quasi-simultaneously in various locations (from lagoons to open waters), all restricted to the northern Adriatic Sea (Malej et al., 2017).
A molecular approach is essential to identify the pathway of introduction of non-indigenous species and thus develop efficient monitoring strategies. Considering the complete lack of genetic data of M. leidyi in the Adriatic Sea, in this study, we used Internal Transcribed Spacer (ITS) region and Cytochrome b (CytB) molecular markers to investigate for the first time the genetic diversity of M. leidyi in the northern Adriatic Sea and to explore the phylogenetic relationships of populations sampled from 2016 in the northern Italian coast (Marano and Grado Lagoon and Gulf of Trieste) with native and introduced populations in other basins.
Materials and methods
Study area
The areas of interest for this study are located in the northernmost part of the northern Adriatic Sea (NAS): the Gulf of Trieste (a shallow, semi-enclosed basin) and the nearby Marano and Grado Lagoon (a shallow transitional system considered one of the best-preserved wetlands in the entire Mediterranean; Bettoso et al., 2013) (Fig. 2). The NAS is one of the most productive areas of the Mediterranean (Morello & Arneri, 2009; Marini & Grilli, 2023) and at the same time affected by heavy maritime traffic. Zupančič et al. (2015) showed that the main traffic route extends from the entrance through the Strait of Otranto in the south to the main northern ports (Rijeka, Pula, Koper, Trieste, Venice, Ancona), where the cumulative ballast water discharge between 2012 and 2015 was estimated at almost 2 × 106 m3 per year (Penko et al., 2016). Most of the ballast water discharged in these ports originated from the Adriatic Sea (80.9%) and the rest from the Mediterranean Sea (16.4%), followed by the Black Sea (1.1%), the northern Atlantic Ocean (0.8%), the Red Sea (0.4%) and the Gulf of Mexico (0.2%) (Penko et al., 2016).
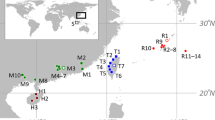
Mnemiopsis leidyi’s sampling sites with pie charts showing the distribution and frequency of the CytB haplotype (outer donut) and ITS alleles (inner circle) of Mnemiopsis leidyi in the study area. Only results obtained at sites with both molecular markers were shown. Blue dots indicate samples collected in the Gulf of Trieste, while green dots indicate samples collected in the Marano and Grado Lagoon. Each sampling site is represented with its own code (as shown in Table 1)
Sample collection, DNA extraction and sequencing
Samples of M. leidyi were collected in 2016, 2018 and 2021 in the Italian waters of the Gulf of Trieste and in the Marano and Grado Lagoon (Table 1 and Fig. 2) using WP2 and bongo nets in the sea and lagoon, respectively. Unfortunately, only a few individuals were immediately individually frozen at – 80 °C and all were analysed for the present study.
Genomic DNA was isolated from gelatinous lobe tissue of 61 ctenophores using a modified CTAB/Chloroform protocol from Ehrlich (2017). DNA amplification was carried out using two different types of molecular markers: a mitochondrial marker, Cytochrome b (CytB) using the KMBMT – 80 and KMBMT – 116 primers (Bayha, 2005), and a nuclear region (ITS-1, 5.8 S gene, and ITS-2) applying the universal ITS5F and ITS4R primers (White et al., 1990).
PCR amplification for both markers was performed using QIAGEN Multiplex—PCR Master Mix. The PCR protocol for CytB included an initial denaturing step at 94 °C for 2 min, followed by 36 amplification cycles (94 °C for 45 s, 50 °C for 1 min, 72 °C for 1:30 min), and a final elongation step at 72 °C for 10 min (Bayha, 2005). PCR for the nuclear region was performed with an initial denaturing step at 95 °C for 1 min, followed by 35 amplification cycles (95 °C for 30 s, 50 °C for 30 s, 72 °C for 50 s), and a final elongation step at 72 °C for 7 min (Ghabooli et al., 2011). PCR products were purified with ExoSAP-IT Express kit (Applied Biosystem®) and then sent to MACROGEN EUROPE (Milan, Italy) for sequencing.
Population genetic and phylogenetic analyses
All sequences for each of the two markers were checked for double peaks in the electropherograms and for stop-codons in the amino-acid translations (for the CytB) before running further analysis and then aligned using MEGA v11 software (Mega Evolutionary Genetics Analysis, Version 11) (Tamura et al., 2021).
Since for ITS sequences, double nucleotide calls (overlapping peaks) were observed, ITS alleles were estimated using PHASE 2.1 a new statistical method for haplotype reconstruction from population data (Stephens et al., 2001) on DNASP v. 5 (Librado & Rozas, 2009), which implements a coalescent-based Bayesian method to infer them. Both alleles of all individuals were included in the following analyses.
We assessed genetic diversity within populations using by means of number of alleles (A), allelic diversity (h) and nucleotide diversity (π) using DNAsp v.5.10.1 (Rozas & Rozas, 1999) and by calculating observed heterozygosity (Ho) and expected heterozygosity (He) using the GENEPOP software (accessible online at http://genepop.curtin.edu.au) and Arlequin (version 3.1) (Excoffier et al., 2005). The Markov chain method was used to estimate the probability of significant deviation from Hardy–Weinberg equilibrium (HWE) using GENEPOP.
The degree of genetic structure between populations of the north Adriatic Sea was determined from pairwise FST using Arlequin.
Then, M. leidyi’s sequences already present in NCBI GenBank were retrieved and then aligned with our sequences.
The function ModelTest in the Phangorn 2.5.5 package (Schliep, 2011; Schliep et al., 2016) in RStudio (RStudio Team, 2020) was used to find the best model of evolution, and the best-fitting model for the data was chosen based on the Akaike Information Criterion (Akaike, 1973). For ITS, the best evolution model was found to be the Symmetric Substitution Model + Gamma distribution with 4 categories + Invariant sites (SYM + G(4) + I). To further confirm the choice of model, the difference between likelihoods resulting from the use of the best-fitting model and the one having one parameter less was tested by computing an Analysis of Variance (ANOVA).
Phylogenetic relationship among alleles was reconstructed using the maximum likelihood (ML) method (Fukami & Tateno, 1989) performed with the function optim.pml in the Phangorn packages and then the tree was edited using MEGA (Tamura et al., 2021). An ITS sequence of the ctenophore Bolinopsis sp. (GenBank Accession no. U65480) was used as outgroup.
A parsimony network for the CytB was generated using Haplotype Viewer (Center for Integrative Bioinformatics Vienna), while a Median Joining network (Bandelt et al., 1999) for the ITS was generated using PopART (Leigh and Bryant, 2015).
Results
315 base pairs (bp) of partial CytB sequences were obtained for a total of 39 individuals from 12 sampling sites (Fig. 2; Table 1) (Accession numbers: OR290138-OR290176). All individuals have the same haplotype, regardless of whether they were sampled in the lagoon or at sea. Comparison of our sequences with those in NCBI GenBank (Accession numbers: KM035326 – KM035399) shows that the northern Adriatic haplotype corresponds to the globally distributed haplotype (Accession number KM035341), which was also observed in the Gulf of Mexico and the Black Sea/Mediterranean (Accession number KM035396) (Fig. SI 1).
619 base pairs (bp) including ITS1, 5.8S and ITS2 were obtained for 20 individuals of M. leidyi from 5 sampling sites (Fig. 2; Table 1). One of 20 amplified individuals presents 1 INDEL at position 37, resulting in an alignment of 620 bp (234 bp for ITS1, 158 bp for 5.8S and 228 bp for ITS2). Compared with the 18 ITS sequences in GenBank (Accession nos. GU062750-GU062762 obtained by Ghabooli et al. 2011; Accession nos. KF435100-KF435104 obtained by Ghabooli et al. 2013), we identified in the northern Adriatic 8 new alleles (designated S to Z to continue the nomenclature previously used by Ghabooli et al., 2013) (GenBank Accession numbers: OR724030-OR724036 and OR731403) defined by eight variable sites and one INDEL (five variable sites and one INDEL in the ITS1 region and three variable sites in the ITS2 region). No variable sites were observed for 5.8S region (Table SI 1). In addition, we found 3 alleles previously identified by Ghabooli et al. (2011): alleles A, B and D (Fig. 2, 3; Table SI 1). Allele U, one of the newly identified alleles, was the most prevalent among the studied individuals (27.5%), followed by allele B (22.5%) and allele A (17.5%) (Fig. 2), while all other alleles were present at lower percentages (2.5–7.5%) (Fig. 2). Overall, in the study area, the total frequency of alleles identified by Ghabooli et al. (2011, 2013) (A, B, D) (47.5%) was comparable to the total frequency of the “new” alleles (S, T, U, V, W, X, Y, Z) (52.5%). All samples contained allele A, whereas alleles B and U were only present in the marine samples (GRI, OGS and TRIO), while allele X was only found in OGS (sea) and MAR2 (lagoon) (Fig. 2).
Alleles distribution and frequency global map for Mnemiopsis leidyi, modified after Ghabooli et al. (2013). ITS alleles represented in this map were obtained by this study and by Ghabooli et al. (2011) and (2013). Each shade indicates a different allele. Following the colour used by Ghabooli et al. (2013), private alleles for Peninsula Valdes coast (G), North Caspian Sea (L, M) and Haifa (O) were highlighted in grey. Population sources were indicated as in Table S3 (Online Resource 1): AZ Sea of Azov, Yasenskaya Bay; BL Black Sea, transect from Blue Bay; BLA Black Sea, near Gelendzhik; NC North Caspian Sea, Makhachkala coast; SC South Caspian Sea, Sari and Noor coasts; BA Baltic Sea, Kiel–Kiel FJord; LD Limfjorden Fjord, Denmark; SP Dénia, Spain; FR Berre Lagoon, Marseille, France; IT Ligurian Sea, Italy; in red ADR Northern Adriatic Sea, Gulf of Trieste and Lagoon of Marano and Grado, Italy; HF Haifa, Israel; NB Narragansett Bay, Rhode Island; YR York River, Virginia; MH Morehead, North Carolina; FL Tampa Bay, Florida; PV Peninsula Valdes coast, Argentina. Modified after Ghabooli et al. (2013).
The 11 alleles resulted in 11 genotypes (Table SI 1). The number of alleles ranged from 2 in GRA4 – MAR2 to 7 in GRI-TRIO, while the allelic diversity (h) ranged from 0.615 in TRIO to 1 in GRA4 – MAR2 and the nucleotide diversity (π) was relatively low (Table 2). All the samples were in equilibrium of Hardy–Weinberg except GRI that showed a significant excess of heterozygosity (Table 2). No significant genetic difference was found between the samples from the northern Adriatic Sea (p > 0.01, Table SI 2), and therefore, all individuals were considered as belonging to a panmictic population and integrated into the allele distribution and frequency map created by Ghabooli et al. (2013) (Fig. 3; Table SI 3). Alleles A, B and D were most common in the Adriatic (ADR) (47.5%), as well as in the native areas of the Gulf of Mexico (MH, FL) and Mediterranean (SP, FR, IT, HF), Black Sea (BLA, BL)/Azov Sea (AZ) and Caspian Sea (NC, SC), where these alleles were even more frequent (≥ 67%). However, the Adriatic Sea presents very peculiar features as none of the other alleles identified in the Mediterranean by Ghabooli et al. (2013) were found in this area. In fact, all other populations in the Mediterranean (sites in France, Spain, Italy, Israel in Ghabooli et al., 2013) showed alleles F and N: allele F was also found in the populations of Florida and Morehead as well as in those of all other basins in southern Europe (Caspian Sea, Black Sea/Azov), while allele N was only detected in Morehead (Gulf of Mexico). Although allele F did not occur in the Adriatic Sea, the allelic network (Fig. 4) showed that it was very similar to allele V, one of the private alleles identified in the Adriatic Sea. Another allele that occured in almost all populations in the Mediterranean (with the exception of Haifa, Israel), the Sea of Azov and the Black Sea was allele H, which was also found at the native site of Morehead (MH). Allele U (the predominant allele in the Adriatic) is possibly descended from allele H, which in turn is genetically closely related to allele B (Fig. 4). In the allelic network (Fig. 4), 6 of the 8 private Adriatic alleles (S, T, U, W, X, Z) form a cluster with different mutation steps between the different alleles: for example, there are 8 mutation steps between allele Z and allele T, while only 2 mutation steps separate allele T from alleles U and P and allele Z from alleles S and C. Similar results were observed on phylogenetic tree that did not show any distinct phylogeographic structure confirming the results of Ghabooli et al. (2011) and a close relationship among the new founded alleles (Fig. SI 2).
Network relationships among ITS alleles for native and invasive populations, inferred by statistical parsimony. Circles in the network (A–Z) correspond to sampled alleles described in Table S3 and Fig. 3. The size of the circles corresponds to the frequency of the allele among all samples. Shades are showing different sampling locations: AZ Sea of Azov; BL Black Sea; CS Caspian Sea; BA Baltic Sea; NA North America; SA South America; SP Dénia, Spain; HF Haifa, Israel; LD Limfjorden Fjord, IT Ligurian Sea, Italy, FR Berre Lagoon, Marseille, France; ADR Northern Adriatic Sea, Gulf of Trieste, and Marano and Grado Lagoon, Italy
Discussion
Despite the recent attention paid to Mnemiopsis leidyi in the northern Adriatic (Malej et al., 2017; Fiori et al., 2019; Budiša et al., 2021; Tirelli et al., 2021; Paliaga et al., 2021; Schroeder et al., 2023; Fadeev et al., 2023; Piccardi et al., 2024), the results obtained in this study represent the first preliminary molecular characterization of M. leidyi individuals in this area. The aim of this work was to evaluate the genetic diversity and phylogenetic relationships between individuals sampled at different sites on the coasts of the Friuli Venezia Giulia region (Marano and Grado Lagoon and Gulf of Trieste, Italy) with those from native populations and those introduced in other basins, using two molecular markers: a mitochondrial marker, Cytochrome B (CytB), and a nuclear marker, Internal Transcribed Spacer (ITS) region.
The analysis of M. leidyi in the northern Adriatic Sea using the CytB marker has shown the presence of a unique haplotype that has already been observed in native (Gulf of Mexico) and invaded areas (Black Sea/Caspian Sea/Mediterranean Sea) (Bayha et al., 2015). Bayha et al. (2015) found 74 CytB haplotypes when monitoring 32 geographic locations (21 and 11 in the native and invasive range, respectively), of which only 8 occurred in southern Europe (Spain, France, 2 locations in Greece, Turkey, Black Sea, Russia, and Azerbaijan): Haplotype 16 was the most common (84% of southern European samples) and was the only one also found in the Adriatic Sea.
Despite the low number of available specimens, the analyses performed with the ITS marker revealed the presence of 11 alleles in our study area, 3 of which (A, B and D) were already found by Ghabooli et al. (2011 and 2013), while the other 8 (from S to Z) are alleles that have never been identified before. Six of them (S, T, V, W, Y and Z) were identified in only one individual each, while the other two (U and X) were found in at least two individuals. Comparing the results obtained for the Adriatic Sea with those of Ghabooli et al. (2011; 2013), similarities between ctenophores from the Gulf of Mexico (Florida, Morehead) and southern Europe (Mediterranean, Caspian Sea, Black/Azovian Sea) can be observed, while the samples from the Adriatic Sea did not show the allele C typical for North America (York River, Narragansett Bay) and northern Europe (Limfjorden, Baltic Sea). A total of 26 alleles have currently been found, 23 of them in the Gulf of Mexico and/or southern Europe (of which only 12 alleles have been found in more than one location), while only 6 alleles have been identified in northern Europe and in the original areas of North America. Overall, the Adriatic Sea is the site with the highest number of alleles (11) followed by the Morehead site (MH), where 9 alleles were identified. Moreover, in the Adriatic the majority of the alleles are singletons (6 out 8 alleles) with a very low nucleotide diversity. This result, although obtained on the basis of a relatively small number of individuals analysed (61 ctenophores), supports the hypothesis that M. leidyi may have been recently introduced into this area. Previous works have shown that individuals from the Gulf of Mexico spread in the Mediterranean Sea after their introduction into the Black Sea probably with ballast water (Ghabooli et al. 2011, 2013; Bayha et al., 2015; Shiganova et al., 2019; Jaspers et al., 2021) and it is therefore plausible that it has route also reached the northern Adriatic via this route, probably more than once in the last decades (Shiganova & Malej, 2009; Malej et al., 2017).
Lagoons have been identified as refugia for this ctenophore in native and invaded areas (Costello et al., 2012; Marchessaux et al., 2020), where this species can maintain a population in low numbers when environmental conditions become unfavourable for reproduction, and then spread again when temperature and food conditions improve. The connection between the lagoon and the sea could allow specimens that seem to have disappeared in the marine waters (as generally happens in the coastal waters of the Mediterranean during the cold season) to spread again and form the typical large summer aggregations. This could also be the case for M. leidyi in the Gulf of Trieste, where the species is almost absent in winter, while it blooms in summer (Tirelli et al., 2021). The refugia mechanism could also be supported by our results, as no significant genetic differences were found between lagoon and marine samples. In general, M. leidyi in the northern Adriatic showed low genetic diversity and no significant genetic structure among populations, suggesting that in this area individuals in this area belong to a single panmictic population, as also observed for another gelatinous zooplankton (Velella velella (Linnaeus, 1758)) in the central Mediterranean (Ligurian Sea–Tyrrhenian Sea) (Morello et al., 2023).
In conclusion, this work, although preliminary, lays the foundation for future studies to understand the patterns of introduction and spread of M. leidyi in the northern Adriatic, possibly also using others molecular markers such as microsatellites or more powerful analysis techniques such as Whole Genome Sequencing (e.g. Jaspers et al., 2021). As Pierson et al. (2020) note, without targeted and consistent monitoring of coastal ecosystems, the impact of invasive species such as M. leidyi on biodiversity is likely to go unnoticed until widespread changes have already occurred in the invaded ecosystems and management becomes more difficult. This was the case in the Black Sea, where studies on M. leidyi were intensified when strong impacts on small pelagic fisheries occurred. The Adriatic Sea is one of the most productive areas of the Mediterranean and an important nursery area for small pelagic fish, where the impact of M. leidyi is far from being known. The high diversity highlighted by the analysis of the specimens collected in the relatively small area of the Gulf of Trieste and the nearby Marano and Grado Lagoon shows how important it is to continue monitoring this species in this region by increasing the number of specimens and the temporal pattern of observations. Furthermore, it would be very important to carry out targeted transboundary projects on M. leidyi at the basin scale, involving the western and eastern Adriatic, in order to characterize the species as well as possible, both from a genetic and ecological point of view.
Data availability
All relevant data are within the paper and its Supporting Information files.
References
Akaike, H., 1973. Maximum likelihood identification of gaussian autoregressive moving average models. Biometrika 60: 255–265. https://doi.org/10.1093/biomet/60.2.255.
Antajan, E., T. Bastian, T. Raud, J. M. Brylinski, S. Hoffman, G. Breton, V. Cornille, A. Delegrange & D. Vincent, 2014. The invasive ctenophore Mnemiopsis leidyi A. Agassiz, 1865 along the english channel and the north sea French coasts: another introduction pathway in northern European waters? Aquatic Invasions 9: 167–173. https://doi.org/10.3391/ai.2014.9.2.05.
Baker, L. D. & M. R. Reeve, 1974. Laboratory culture of the lobate ctenophore Mnemiopsis mccradyi with notes on feeding and fecundity. Marine Biology 26: 57–62. https://doi.org/10.1007/BF00389086.
Bandelt, H. J., P. Forster & A. Rohl, 1999. Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution 16: 37–48. https://doi.org/10.1093/oxfordjournals.molbev.a026036.
Bayha, K. M., 2005. The molecular systematics and population genetics of four coastal ctenophores and scyphozoan jellyfish of the U.S. Atlantic and Gulf of Mexico. PhD. Dissertation, The University of Delaware, Newark.
Bayha, K. M., M. H. Chang, C. L. Mariani, J. L. Richardson, D. L. Edwards, T. S. DeBoer, C. Moseley, E. Aksoy, M. B. Decker, P. M. Gaffney, G. R. Harbison, J. H. McDonald & A. Caccone, 2015. Worldwide phylogeography of the invasive ctenophore Mnemiopsis leidyi (Ctenophora) based on nuclear and mitochondrial DNA data. Biological Invasions 17: 827–850. https://doi.org/10.1007/s10530-014-0770-6.
Bettoso, N., A. Acquavita, A. D’Aietti & G. Mattassi, 2013. The Marano and Grado Lagoon: A brief synopsis on the aquatic fauna and fisheries resources. Annales 23: 135–142.
Boero, F., M. Putti, E. Trainito, E. Prontera, S. Piraino & T. Shiganova, 2009. First records of Mnemiopsis leidyi (Ctenophora) from the Ligurian, Tyrrhenian and ionian seas (Western Mediterranean) and first record of Phyllorhiza punctata (Cnidaria) from the western Mediterranean. Aquatic Invasions 4: 675–680. https://doi.org/10.3391/ai.2009.4.4.13.
Boersma, M., A. M. Malzahn, W. Greve & J. Javidpour, 2007. The first occurrence of the ctenophore Mnemiopsis leidyi in the North Sea. Helgoland Marine Research 61: 153–155. https://doi.org/10.1007/s10152-006-0055-2.
Bolte, S., V. Fuentes, H. Haslob, B. Huwer, D. Thibault-Botha, D. Angel, B. Galil, J. Javidpour, A. Moss & T. Reusch, 2013. Population genetics of the invasive ctenophore Mnemiopsis leidyi in Europe reveal source–sink dynamics and secondary dispersal to the Mediterranean sea. Marine Ecology Progress Series 485: 25–36. https://doi.org/10.3354/meps10321.
Budiša, A., P. Paliaga, T. Juretić, D. Lučić, N. Supić, Z. Pasarić, T. Djakovac, M. Mladinić, V. Dadić & V. Tičina, 2021. Distribution, diet and relationships of the invasive ctenophore Mnemiopsis leidyi with anchovies and zooplankton, in the northeastern Adriatic sea. Mediterranean Marine Science 22: 827. https://doi.org/10.12681/mms.23305.
Corriero, G., C. Pierri, S. Accoroni, G. Alabiso, G. Bavestrello, E. Barbone, M. Bastianini, A. M. Bazzoni, F. Bernardi Aubry, F. Boero, M. C. Buia, M. Cabrini, E. Camatti, F. Cardone, B. Cataletto, R. Cattaneo Vietti, E. Cecere, T. Cibic, P. Colangelo, A. de Olazabal, G. D’Onghia, S. Finotto, N. Fiore, D. Fornasaro, S. Fraschetti, M. C. Gambi, A. Giangrande, C. Gravili, R. Guglielmo, C. Longo, M. Lorenti, A. Lugliè, P. Maiorano, M. G. Mazzocchi, M. Mercurio, F. Mastrototaro, M. Mistri, M. Monti, C. Munari, L. Musco, C. Nonnis-Marzano, B. M. Padedda, F. P. Patti, A. Petroncelli, S. Piraino, G. Portacci, A. Pugnetti, S. Pulina, T. Romagnoli, I. Rosati, D. Sarno, C. T. Satta, N. Sechi, S. Schiaparelli, B. Scopione, L. Sion, A. Terlizzi, V. Tirelli, C. Totti, A. Tursi, N. Ungaro, A. Zingone, V. Zupo & A. Basset, 2016. Ecosystem vulnerability to alien and invasive species: a case study on marine habitats along the Italian coast. Aquatic Conservation: Marine and Freshwater Ecosystems 26: 392–409. https://doi.org/10.1002/aqc.2550.
Costello, J. H., K. M. Bayha, H. W. Mianzan, T. A. Shiganova & J. E. Purcell, 2012. Transitions of Mnemiopsis leidyi (Ctenophora: Lobata) from a native to an exotic species: a review. Hydrobiologia 690: 21–46. https://doi.org/10.1007/s10750-012-1037-9.
Diciotti, R., J. Culurgioni, S. Serra, M. Trentadue, G. Chessa, C. T. Satta, T. Caddeo, A. Lugliè, N. Sechi & N. Fois, 2016. First record of Mnemiopsis leidyi (Ctenophora, Bolinopsidae) in Sardinia (S’Ena Arrubia Lagoon, Western Mediterranean): a threat to local fishery? Mediterranean Marine Science 17: 714–719. https://doi.org/10.12681/mms.1719.
Edgar, A., J. M. Ponciano & M. Q. Martindale, 2022. Ctenophores are direct developers that reproduce continuously beginning very early after hatching. Proceedings of the National Academy of Sciences 119(18): e2122052119. https://doi.org/10.1073/pnas.2122052119.
Ehrlich, M., 2017. Invasion Genomics: population structure and diversity patterns in the invasive ctenophore Mnemiopsis leidyi based on whole-genome re-sequencing. Master thesis, Christian-Albrechts-Universität Kiel, Kiel, Germany, p. 93
Excoffier, L., L. Guillaume & S. Schneider, 2005. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evolutionary Bioinformatics 1: 47–50. https://doi.org/10.1177/117693430500100003.
Faasse, M. & K. Bayha, 2006. The ctenophore Mnemiopsis leidyi A. Agassiz 1865 in coastal waters of the Netherlands: an unrecognized invasion? Aquatic Invasions 1: 270–277. https://doi.org/10.3391/ai.2006.1.4.9.
Fadeev, E., J. H. Hennenfeind, C. Amano, Z. Zhao, K. Klun, G. J. Herndl & T. Tinta, 2023. Bacterial degradation of ctenophore Mnemiopsis leidyi organic matter. bioRxiv. https://doi.org/10.1101/2023.08.14.553244.
Fernandes, P. G., G. M. Ralph, A. Nieto, M. García Criado, P. Vasilakopoulos, C. D. Maravelias, R. M. Cook, R. A. Pollom, M. Kovačić, D. Pollard, E. D. Farrell, A. B. Florin, B. A. Polidoro, J. M. Lawson, P. Lorance, F. Uiblein, M. Craig, D. J. Allen, S. L. Fowler, R. H. L. Walls, M. T. Comeros-Raynal, M. S. Harvey, M. Dureuil, M. Biscoito, C. Pollock, S. R. McCully Phillips, J. R. Ellis, C. Papaconstantinou, A. Soldo, Ç. Keskin, S. W. Knudsen, L. Gil De Sola, F. Serena, B. B. Collette, K. Nedreaas, E. Stump, B. C. Russell, S. Garcia, P. Afonso, A. B. J. Jung, H. Alvarez, J. Delgado, N. K. Dulvy & K. E. Carpenter, 2017. Coherent assessments of Europe’s marine fishes show regional divergence and megafauna loss. Nature Ecology & Evolution 1: 0170. https://doi.org/10.1038/s41559-017-0170.
Finenko, G., A. Kideys, B. Anninsky, T. Shiganova, A. Roohi, M. Tabari, H. Rostami & S. Bagheri, 2006. Invasive ctenophore Mnemiopsis leidyi in the Caspian Sea: feeding, respiration, reproduction and predatory impact on zooplankton community. Marine Ecology Progress Series 314: 171–185.
Fiori, E., M. Benzi, C. R. Ferrari & C. Mazziotti, 2019. Zooplankton community structure before and after Mnemiopsis leidyi arrival. Journal of Plankton Research 41: 803–820. https://doi.org/10.1093/plankt/fbz060.
Fukami, K. & Y. Tateno, 1989. On the maximum likelihood method for estimating molecular trees: uniqueness of the likelihood point. Journal of Molecular Evolution 28: 460–464. https://doi.org/10.1007/BF02603081.
Galil, B., N. Kress & T. Shiganova, 2009. First record of Mnemiopsis leidyi A. Agassiz, 1865 (Ctenophora; Lobata; Mnemiidae) off the Mediterranean coast of Israel. Aquatic Invasions 4: 357–360. https://doi.org/10.3391/ai2009.4.2.8.
GESAMP, 1997. Opportunistic settlers and the problem of the ctenophore Mnemiopsis leidyi invasion in the Black Sea, vol 58. GESAMP Reports and Studies. International Maritime Organization, London
Ghabooli, S., T. A. Shiganova, A. Zhan, M. E. Cristescu, P. Eghtesadi-Araghi & H. J. MacIsaac, 2011. Multiple introductions and invasion pathways for the invasive ctenophore Mnemiopsis leidyi in Eurasia. Biological Invasions 13: 679–690. https://doi.org/10.1007/s10530-010-9859-8.
Ghabooli, S., T. A. Shiganova, E. Briski, S. Piraino, V. Fuentes, D. Thibault-Botha, D. L. Angel, M. E. Cristescu & H. J. MacIsaac, 2013. Invasion pathway of the Ctenophore Mnemiopsis leidyi in the Mediterranean sea. PLoS ONE 8: e81067. https://doi.org/10.1371/journal.pone.0081067.
Giangrande, A., C. Pierri, M. Del Pasqua, C. Gravili, M. C. Gambi & M. F. Gravina, 2020. The Mediterranean in check: biological invasions in a changing sea. Marine Ecology 41: e12583. https://doi.org/10.1111/maec.12583.
Hansson, H. G., 2006. Ctenophores of the baltic and adjacent seas—the invader Mnemiopsis is here! Aquatic Invasions 1: 295–298. https://doi.org/10.3391/ai.2006.1.4.16.
Ivanov, V. P., A. M. Kamakin, V. B. Ushivtzev, T. Shiganova, O. Zhukova, N. Aladin, S. I. Wilson, G. R. Harbison & H. J. Dumont, 2000. Invasion of the Caspian sea by the comb jellyfish Mnemiopsis leidyi (Ctenophora). Biological Invasions 2: 255–258. https://doi.org/10.1023/A:1010098624728.
Jaspers, C., L. F. Møller & T. Kiørboe, 2015. Reproduction rates under variable food conditions and starvation in Mnemiopsis leidyi: significance for the invasion success of a ctenophore. Journal of Plankton Research 37: 1011–1018. https://doi.org/10.1093/plankt/fbv017.
Jaspers, C., B. Huwer, E. Antajan, A. Hosia, H. H. Hinrichsen, A. Biastoch, D. Angel, R. Asmus, C. Augustin, S. Bagheri, S. E. Beggs, T. J. S. Balsby, M. Boersma, D. Bonnet, J. T. Christensen, A. Dänhardt, F. Delpy, T. Falkenhaug, G. Finenko, N. E. C. Fleming, V. Fuentes, B. Galil, A. Gittenberger, D. C. Griffin, H. Haslob, J. Javidpour, L. Kamburska, S. Kube, V. T. Langenberg, M. Lehtiniemi, F. Lombard, A. Malzahn, M. Marambio, V. Mihneva, L. F. Møller, U. Niermann, M. I. Okyar, Z. B. Özdemir, S. Pitois, T. B. H. Reusch, J. Robbens, K. Stefanova, D. Thibault, H. W. Van Der Veer, L. Vansteenbrugge, L. Van Walraven & A. Woźniczka, 2018. Ocean current connectivity propelling the secondary spread of a marine invasive comb jelly across western Eurasia. Global Ecology and Biogeography 27: 814–827. https://doi.org/10.1111/geb.12742.
Jaspers, C., M. Ehrlich, J. M. Pujolar, S. Künzel, T. Bayer, M. T. Limborg, F. Lombard, W. E. Browne, K. Stefanova & T. B. H. Reusch, 2021. Invasion genomics uncover contrasting scenarios of genetic diversity in a widespread marine invader. Proceedings of the National Academy of Sciences 118: e2116211118. https://doi.org/10.1073/pnas.2116211118.
Javidpour, J., U. Sommer & T. Shiganova, 2006. First record of Mnemiopsis leidyi A. Agassiz 1865 in the Baltic sea. Aquatic Invasions 1: 299–302. https://doi.org/10.3391/ai.2006.1.4.17.
Javidpour, J., J. C. Molinero, E. Ramírez-Romero, P. Roberts & T. Larsen, 2020. Cannibalism makes invasive comb jelly, Mnemiopsis leidyi, resilient to unfavorable conditions. Communications Biology 3: 212. https://doi.org/10.1038/s42003-020-0940-2.
Katsanevakis, S., I. Wallentinus, A. Zenetos, E. Leppäkoski, M. E. Çinar, B. Oztürk, M. Grabowski, D. Golani & A. C. Cardoso, 2014a. Impacts of invasive alien marine species on ecosystem services and biodiversity: a pan-European review. Aquatic Invasions 9: 391–423. https://doi.org/10.3391/ai.2014.9.4.01.
Katsanevakis, S., M. Coll, C. Piroddi, J. Steenbeek & F. Ben Rais Lasram, A. Zenetos & A. C. Cardoso, 2014b. Invading the Mediterranean sea: biodiversity patterns shaped by human activities. Frontiers in Marine Science 1: 32. https://doi.org/10.3389/fmars.2014.00032.
Kideys, A., A. Roohi, S. Bagheri, G. Finenko & L. Kamburska, 2005. Impacts of invasive ctenophores on the fisheries of the Black sea and Caspian sea. Oceanography 18: 76–85. https://doi.org/10.5670/oceanog.2005.43.
Leigh, J. W. & D. Bryant, 2015. POPART: full-feature software for haplotype network construction. Methods in Ecology and Evolution 6: 1110–1116. https://doi.org/10.1111/2041-210X.12410.
Librado, P. & J. Rozas, 2009. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25: 1451–1452. https://doi.org/10.1093/bioinformatics/btp187.
Lowe, S., M. Browne, S. Boudjelas, & M. De Poorter, 2000. 100 of the world’s worst invasive alien species. A selection from the global invasive species database. Published by the Invasive Species Specialist Group (ISSG), Auckland—New Zealand, a specialist group of the Species Survival Commission (SSC) of the World Conservation Union (IUCN). Updated and reprinted version: November 2004.
Malej, A., V. Tirelli, D. Lučić, P. Paliaga, M. Vodopivec, A. Goruppi, S. Ancona, M. Benzi, N. Bettoso, E. Camatti, M. Ercolessi, C. R. Ferrari & T. Shiganova, 2017. Mnemiopsis leidyi in the northern Adriatic: here to stay? Journal of Sea Research 124: 10–16. https://doi.org/10.1016/j.seares.2017.04.010.
Marchessaux, G., V. Faure, C. Chevalier & D. Thibault, 2020. Refugia area for the ctenophore Mnemiopsis leidyi A. Agassiz 1865 in the berre lagoon (southeast France): the key to its persistence. Regional Studies in Marine Science 39: 101409. https://doi.org/10.1016/j.rsma.2020.101409.
Marchessaux, G., B. Belloni, J. Gadreaud & D. Thibault, 2021. Predation assessment of the invasive ctenophore Mnemiopsis leidyi in a French Mediterranean lagoon. Journal of Plankton Research 43: 161–179. https://doi.org/10.1093/plankt/fbab002.
Marchessaux, G., D. Thibault & C. Claeys, 2023. An interdisciplinary assessment of the impact of invasive gelatinous zooplankton in a French Mediterranean lagoon. Biological Invasions 25: 499–518. https://doi.org/10.1007/s10530-022-02930-3.
Marini, M. & F. Grilli, 2023. The role of nitrogen and phosphorus in eutrophication of the northern Adriatic sea: history and future scenarios. Applied Sciences 13: 9267. https://doi.org/10.3390/app12169267.
Molnar, J. L., R. L. Gamboa, C. Revenga & M. D. Spalding, 2008. Assessing the global threat of invasive species to marine biodiversity. Frontiers in Ecology and the Environment 6: 485–492. https://doi.org/10.1890/070064.
Morello, E.B. & E. Arneri, 2009. Anchovy and sardine in the Adriatic Sea—an ecological review. In: gibson RN, 375 Atkinson RJA, Gordon JDM (eds) Oceanography and Marine Biology. An Annual Review 47:209–256
Morello, B., M. Bo, F. Betti, G. Bavestrello, M. Abbiati & F. Costantini, 2023. Spatial and temporal genetic structure of Velella velella (Hydrozoa, Porpitidae) and its predator Janthina pallida (Gastropoda, Epitoniidae). Hydrobiologia 850: 1751–1762. https://doi.org/10.1007/s10750-022-05089-z.
Oguz, T., B. Fach & B. Salihoglu, 2008. Invasion dynamics of the alien ctenophore Mnemiopsis leidyi and its impact on anchovy collapse in the Black sea. Journal of Plankton Research 30: 1385–1397. https://doi.org/10.1093/plankt/fbn094.
Oliveira, O. M. P., 2007. The presence of the ctenophore Mnemiopsis leidyi in the oslofjorden and considerations on the initial invasion pathways to the north and Baltic seas. Aquatic Invasions 2: 185–189. https://doi.org/10.3391/ai.2007.2.3.5.
Paliaga, P., A. Budiša, J. Dautović, T. Djakovac, M. A. Dutour-Sikirić, H. Mihanović, N. Supić, I. Celić, N. Iveša, M. Buršić, I. Balković, L. Jurković & I. Ciglenečki, 2021. Microbial response to the presence of invasive ctenophore Mnemiopsis leidyi in the coastal waters of the northeastern Adriatic. Estuarine, Coastal and Shelf Science 259: 107459. https://doi.org/10.1016/j.ecss.2021.107459.
Penko, L., G. Zupančič, U. Kocijančič, A. Popit, M. Gorišek, A. Aubreht, L. Gosar, C. Forte, E. Magaletti, M. Marini, M. Bastianini, Ž. N. Gladan, D. Zec, D. Joksimović, M. Markovčić Kostelac, S. Vidas, M. David, G. De Vendictis, A. Scarpato, L. Spath, A. Petri, G. Prskalo, M. Kustura, J. Kolitari & M. Cara, 2016. Integrated operational plan for 380 ballast water management in the Adriatic. Adriatic IPA. Institute for Water of the R Slovenia, p. 76
Piccardi, F., F. Poli, C. Sguotti, V. Tirelli, D. Borme, C. Mazzoldi & A. Barausse, 2024. Assessing the impact of the invasive ctenophore Mnemiopsis leidyi on artisanal fisheries in the venice lagoon: an interdisciplinary approach. Hydrobiologia. https://doi.org/10.1007/s10750-024-05505-6.
Pierson, J., E. Camatti, R. Hood, T. Kogovšek, D. Lučić, V. Tirelli & A. Malej, 2020. Mesozooplankton and gelatinous zooplankton in the face of environmental stressors. In Malone, T. C., A. Malej & J. Faganeli (eds), Geophysical monograph series Wiley, Hoboken: 105–127.
Piroddi, C. M. C., J. Steenbeek, K. Kaschner, F. Ben Rais Lasram, J. Aguzzi, E. Ballesteros, C. N. Bianchi, J. Corbera, T. Dailianis, R. Danovaro, M. Estrada, C. Froglia, B. S. Galil, J. M. Gasol, R. Gertwagen, J. Gil, F. Guilhaumon, K. Kesner-Reyes, M.-S. Kitsos, A. Koukouras, N. Lampadariou, E. Laxamana, C. M. López-Fé De La Cuadra, H. K. Lotze, D. Martin, D. Mouillot, D. Oro, S. Raicevich, J. Rius-Barile, J. I. Saiz-Salinas, C. San Vicente, S. Somot, J. Templado, X. Turon, D. Vafidis, R. Villanueva & E. Voultsiadou, 2010. The biodiversity of the Mediterranean sea: estimates, patterns, and threats. PLoS ONE 5: e11842. https://doi.org/10.1371/journal.pone.0011842.
Purcell, J. E., T. A. Shiganova, M. B. Decker & E. D. Houde, 2001. The ctenophore Mnemiopsis in native and exotic habitats: U.S. estuaries versus the Black sea basin. Hydrobiologia 451: 145–176. https://doi.org/10.1023/A:1011826618539.
Reusch, T. B. H., S. Bolte, M. Sparwel, A. G. Moss & J. Javidpour, 2010. Microsatellites reveal origin and genetic diversity of Eurasian invasions by one of the world’s most notorious marine invader, Mnemiopsis leidyi (Ctenophora). Molecular Ecology 19: 2690–2699. https://doi.org/10.1111/j.1365-294X.2010.04701.x.
Rozas, J. & R. Rozas, 1999. DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15: 174–175. https://doi.org/10.1093/bioinformatics/15.2.174.
Sasson, D. A. & J. F. Ryan, 2016. The sex lives of ctenophores: the influence of light, body size, and self-fertilization on the reproductive output of the sea walnut. Mnemiopsis Leidyi. Peerj 4: e1846. https://doi.org/10.7717/peerj.1846.
Schliep, K. P., 2011. Phangorn: phylogenetic analysis in R. Bioinformatics 27: 592–593. https://doi.org/10.1093/bioinformatics/btq706.
Schliep, K. P., A. A. Potts, D. A. Morrison & G. W. Grimm, 2016. Intertwining phylogenetic trees and networks. PeerJ 4: e2054v1. https://doi.org/10.7287/peerj.preprints.2054v1.
Schroeder, A., E. Camatti, M. Pansera & A. Pallavicini, 2023. Feeding pressure on meroplankton by the invasive ctenophore Mnemiopsis leidyi. Biological Invasions 25: 2007–2021. https://doi.org/10.1007/s10530-023-03023-5.
Shiganova, T. A., 1997. Mnemiopsis leidyi abundance in the Black sea and its impact on the pelagic community. In Özsoy, E. & A. Mikaelyan (eds), Sensivity to change: black sea, Baltic sea and north sea. NATO ASI Series, Kluwer Academic Publish, Alphen aan den Rijn.
Shiganova, T. A., 2020. Adaptive strategies of Mnemiopsis leidyi A. Agassiz 1865 in different environments of the Eurasian seas. Marine Pollution Bulletin 161: 111737. https://doi.org/10.1016/j.marpolbul.2020.111737.
Shiganova, T. & A. Malej, 2009. Native and non-native Ctenophores in the gulf of trieste, northern Adriatic sea. Journal of Plankton Research 31: 61–71. https://doi.org/10.1093/plankt/fbn102.
Shiganova, T., E.D. Christou, J.V. Bulgakova, I. Siokou-Frangou, S. Zervoudaki & A. Siapatis, 2004. Distribution and biology of the invader Mnemiopsis leidyi in the northern Aegean Sea and comparison with indigenous species Bolinopsis vitrea. In: Dumont, H., Shiganova, T.A., Niermann, U. (Eds.), Aquatic invasions in the black, caspian, and Mediterranean seas. Nato science series: IV, Earth and Environmental Sciences, Kluwer Academic Publishers Vol 35. pp. 113–135. https://doi.org/10.1007/1-4020-2152-6_4
Shiganova, T. A., U. Sommer, J. Javidpour, J. C. Molinero, A. Malej, A. S. Kazmin, M. Isinibilir, E. Christou, I. Siokou- Frangou, M. Marambio, V. Fuentes, Z. A. Mirsoyan, N. Gülsahin, F. Lombard, M. K. S. Lilley, D. L. Angel, B. S. Galil, D. Bonnet & F. Delpy, 2019. Patterns of invasive ctenophore Mnemiopsis leidyi distribution and variability in different recipient environments of the Eurasian seas: a review. Marine Environmental Research 152: 104791. https://doi.org/10.1016/j.marenvres.2019.104791.
Stephens, M., N. J. Smith & P. Donnelly, 2001. A new statistical method for haplotype reconstruction from population data. The American Journal of Human Genetics 68: 978–989. https://doi.org/10.1086/319501.
Studenikina, E. I., S. P. Volovik, I. A. Miryozan & G. I. Luts, 1991. The ctenophore Mnemiopsis leidyi in the sea of azov. Oceanology 3: 722–725.
Tamura, K., G. Stecher & S. Kumar, 2021. MEGA11: molecular evolutionary genetics analysis version 11. Molecular Biology and Evolution 38: 3022–3027. https://doi.org/10.1093/molbev/msab120.
Templado, J., 2014. Future trends of mediterranean biodiversity. In Goffredo, S. & Z. Dubinsky (eds), The Mediterranean sea. Springer, Netherlands.
Tendal, O. S., K. R. Jensen & H. U. Riisgård, 2007. Invasive ctenophore Mnemiopsis leidyi widely distributed in Danish waters. Aquatic Invasions 2: 455–460. https://doi.org/10.3391/ai.2007.2.4.19.
Tirelli, V., A. Goruppi, R. Riccamboni & M. Tempesta, 2021. Citizens’ eyes on Mnemiopsis: how to multiply sightings with a click! Diversity 13: 224. https://doi.org/10.3390/d13060224.
Tiselius, P. & L. F. Møller, 2017. Community cascades in a marine pelagic food web controlled by the non-visual apex predator Mnemiopsis leidyi. Journal of Plankton Research 39: 271–279. https://doi.org/10.1093/plankt/fbw096.
Vinogradov, M. E., E. A. Shushkina, I. A. Musaeva & P. Y. Sorokin, 1989. Ctenophore Mnemiopsis leidyi (A. Agassiz) (Ctenophora: lobata)—new settlers in the Black sea. Oceanology 29: 293–298.
White, T. J., T. Bruns, S. Lee & J. Taylor, 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In Innis, M. A., D. H. Gelfand, J. J. Shinsky & T. J. White (eds), PCR protocols: a guide to methods and applications Academic Press, San Diego: 315–322.
Zenetos, A., P. G. Albano, E. López Garcia, N. Stern, K. Tsiamis & M. Galanidi, 2022. Established non-indigenous species increased by 40% in 11 years in the Mediterranean sea. Mediterranean Marine Science. https://doi.org/10.12681/mms.29106.
Zupančič, G., L. Gosar & M. David, 2015. Report summarizing and analysing shipping patterns in the Adriatic sea—upgrade (final report), BALMAS project, work package 4. Activity 1: 37.
Acknowledgements
We gratefully acknowledge R. Auriemma, N. Bettoso, D. Borme, M. Celio, S. Ciriaco, A. de Olazabal, L. Faresi and A. Goruppi for their assistance with field work and technical support.
Funding
Open access funding provided by Alma Mater Studiorum - Università di Bologna within the CRUI-CARE Agreement. No specific funding was provided to carry out this study, but part of the samples analysed were collected in the framework of the NOCE DI MARE projects funded by the Regione Autonoma Friuli Venezia Giulia (legge regionale 30 marzo 2018, n°14, art 2, commi 51–55 e legge regionale 30 dicembre 2020, n°26, art. 4, commi 33–34, CUP F96C18000240002 and CUP: D29J21000900002). VT gratefully acknowledges the support of the National Biodiversity Future Center—NBFC, funded under the National Recovery and Resilience Plan (NRRP), Mission 4 Component 2 Investment 1.4—Call for tender No. 3138 of 16 December 2021, rectified by Decree no. 3175 of 18 December 2021 of Italian Ministry of University and Research funded by the European Union—NextGenerationEU; Project code CN_00000033, Concession Decree No. 1034 of 17 June 2022 adopted by the Italian Ministry of University and Research,CUP F83B22000050001. FC was partially supported by the RETURN Extended Partnership and received funding from the European Union Next-GenerationEU (National Recovery and Resilience Plan – NRRP, Mission 4, Component 2, Investment 1.3 – D.D. 1243 2/8/2022, PE0000005).
Author information
Authors and Affiliations
Contributions
EP, FC and VT conceptualized the study. VT carried out the field samplings. EP and FC carried out the analysis. EP, FC and VT contributed to analysing and visualising the data. The first draft of the manuscript was written by EP with input from all co-authors. All authors commented on previous versions of the manuscript, drafted the work and revised it critically. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have not conflicts of interest to declare that are relevant to the content of this article. The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Guest editors: Sidinei M. Thomaz, Cécile Fauvelot, Lee B. Kats, Jonne Kotta & Fernando M. Pelicice / Aquatic Invasive Species IV
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Putelli, E., Costantini, F. & Tirelli, V. A first molecular insight into the invasive ctenophore Mnemiopsis leidyi in the northern Adriatic sea. Hydrobiologia (2024). https://doi.org/10.1007/s10750-024-05597-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10750-024-05597-0