Abstract
Native communities can resist the establishment and invasion of alien species through consumptive and/or competitive interactions. The extent of consumptive resistance from freshwater fish to the invasion of zebra mussels Dreissena polymorpha, a globally invasive Ponto-Caspian species, was assessed in two areas in Britain using stable isotope analysis, where mixing models predicted the contribution of putative prey resources (including zebra mussel) to fish diet. Across the sites and species, only roach Rutilus rutilus were predicted to have a diet where zebra mussels contributed highly (predicted contribution: 44%), with literature suggesting that their functional morphology would have facilitated their consumption of this prey item. Predicted contributions of zebra mussels to common bream Abramis brama diet was comparatively low (29%), despite them being present to much larger sizes than roach, and with pike Esox lucius, perch Perca fluviatilis and pikeperch Sander lucioperca also predicted to have low dietary contributions of zebra mussels (0.08%, 24% and 24%, respectively). These results suggest the consumptive resistance to its invasion in Britain has been low and, correspondingly, if there is a management desire to further limit the invasion of zebra mussels then relying on biological resistance to limit their invasion appears to be insufficient.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alien species are a pervasive agent of global change (Simberloff et al., 2013), with the consequences of invasions including both ecological and economic damage (Crystal-Ornelas & Lockwood, 2020; Diagne et al., 2021). Although the number of alien species introduced into new regions remains relatively high at global levels (Dawson et al., 2017; Turbelin et al., 2017), the proportion of these introductions that develop damaging invasions tends to be relatively low (Gozlan, 2008). Nevertheless, an area that has produced a relatively high number of globally invasive aquatic species is the Ponto-Caspian region (Ricciardi & MacIsaac, 2000), which comprises a large geographic extension around the Black, Azov and Caspian seas that is characterised by variable climatic and habitat conditions that provide its native species with relatively high adaptation capacity (Gallardo & Aldridge, 2013). Invasive species that originate from this region include amphipods (e.g. killer shrimp Dikerogammarus villosus (Sowinsky, 1894)) and molluscs (e.g. zebra mussel Dreissena polymorpha (Pallas, 1771)) (Gallardo & Aldridge, 2015).
In the risk management of introduced alien species, an important component is understanding the processes by which introduced species are able to colonise novel ecosystems (Dominguez Almela et al., 2020), where the establishment of the population and its integration into the native food web are fundamentally important (Britton & Gozlan, 2013; Britton et al., 2019). There are a number of hypotheses on factors that influence the establishment of an introduced species and thus whether it will go on to develop an invasive population (Catford et al., 2009). These include the biological resistance hypothesis, which posits that communities of native species can mediate the establishment and performance of invasive species through competitive and/or consumptive interactions (Britton, 2012; Alofs & Jackson, 2014). In freshwaters ecosystems, evidence suggests that the mechanisms of biological resistance are stronger from consumptive resistance than competitive resistance (Alofs & Jackson, 2014), with functionally diverse communities having relatively high resistance against multiple invaders (Byun et al., 2020).
Zebra mussels are a global invader whose continued range expansion has been facilitated by their transport on boats (Robertson et al., 2020). Zebra mussels have been implicated in substantial changes in invaded freshwaters, including increased water clarity, modified nutrient cycling, altered composition of the benthic community and changes in the fish community (Hansen et al., 2020; Robertson et al., 2020). Furthermore, the ability of zebra mussels to efficiently filter water (4 L/day; Silverman et al., 1996) has been shown to impact dissolved oxygen concentrations (Strayer, 2009; Wong et al., 2021) and phytoplankton/zooplankton densities (Wong et al., 2021), resulting in cascading effects (Higgins & Vander Zanden, 2010; Rennie et al., 2013). However, zebra mussels can also be important dietary component of the resident fish community, with studies in North America suggesting that at least six fish species are potential predators of these mussels, with this facilitated by their mouth size, teeth and chewing abilities (French, 1993). Magoulick & Lewis (2002) reported that zebra mussels were an important part of the diet of blue catfish Ictalurus furcatus (Lesueur, 1840), freshwater drum Aplodinotus grunniens (Rafinesque, 1819) and redear sunfish Lepomis microlophus (Günther, 1859). Laboratory experiments on round goby Neogobius melanostomus (Pallas, 1814) suggested they preferred consuming zebra mussels over species of clams and snails (Ghedotti et al., 1995). Studies on roach Rutilus rutilus (Linnaeus, 1758) indicated that larger fish would consume more mussels than smaller individuals, with fish below 160 mm in length considered as unable to consume them (Prejs et al., 1990). In a Canadian study, Wong et al. (2021) argued that although some fish may feed on zebra mussels, they would still use other prey resources as their primary dietary items and so would not act as a biological control of these invaders. The bias of studies on zebra mussels to North America indicates there are considerable knowledge gaps on their invasion elsewhere in their range, especially in relation to their integration into invaded food webs and how they contribute to consumer diets, and thus what the extent of the biological resistance is to their invasion.
Zebra mussels were introduced into Great Britain in 1824 and have since spread to a number of discrete lakes, rivers and canals, with Aldridge et al. (2004) suggesting relatively recent increases in their abundance and distribution. However, there remains limited knowledge on their invasion in Britain, including to what extent they are dietary components of fish communities, where species present include roach (as per Prejs et al., 1990). Also present is the common bream Abramis brama (Linnaeus, 1758), which Nagelkerke & Sibbing (2011) considered to not be an efficient consumer of zebra mussels due to limitations set by the crushing power of their pharyngeal teeth. The objective of the current study was to assess the dietary contributions of zebra mussels to the fish community of two invaded and spatially discrete sites of Britain, where dietary contributions were predicted from the ecological application of stable isotope analysis. These dietary contributions were predicted and compared between omnivorous and piscivorous native and alien fishes, given these will differ in their putative prey resources, with the effect of body lengths of individual fish also tested on the dietary predictions. We posit that zebra mussels are an important component of the diet of at least some fish species in invaded waters in Britain and these fish species would have thus provided some consumptive resistance to their invasion.
Materials and methods
Site details
Site 1 was the Gloucester–Sharpness Canal in western England that is connected to the River Severn estuary at its downstream end (51.7249 N, 2.4733 W). This canal was constructed to allow for navigation of goods from the sea to Gloucester docks; 25 m wide and 5 m deep (Nolan & Britton, 2018). Sampling was focused in the areas downstream of Gloucester, where the fish assemblage is dominated by the cyprinid species roach Rutilus rutilus and common bream Abramis brama (‘bream’ hereafter), with perch Perca fluviatilis (Linnaeus, 1758) and invasive pikeperch Sander lucioperca (Linnaeus, 1758) also present (J.R. Britton, personal observations). Site 2 was the River Bure in eastern England. This river is 87 km in length, flowing south-east towards Breydon Water estuary at Great Yarmouth. Along with its tributaries, the Rivers Ant and Thurne, it forms the northern area of the Broads National Park, a wetland of significant ecological importance (Natural England, 2020; Winter et al., 2021c). The River Ant is mentioned as being colonised by zebra mussels in Aldridge et al. (2004). The area is characterised by multiple small shallow lakes termed ‘Broads’ (mediaeval peat diggings) less than 40 ha in size. As the landscape is generally flat, the catchment is tidal for approximately 45 km inland. The focal area under study here was upstream of the upper limit of saline incursion at Horning (Winter et al., 2021c), where the river was approximately 25 m wide with depths to 1.5 m. Depths of the connected lakes were typically ~ 1.5 m and did not exceed 3 m. Across the two reaches, the fish assemblage has limited diversity, being dominated by roach and bream, with perch and pike Esox lucius (Linnaeus, 1758) also present (Table 1). Both sites had zebra mussels present in all sampled areas, being abundant on areas such as submerged boat moorings.
Sampling
The main sampling period for site 1 was April to June 2017 where, due to the habitat of the canal (depth, width, boat traffic), sampling by traditional capture methods (e.g. electric fishing, seine netting) was not feasible. Rod-and-line angling was used as an alternative, where scales were collected from captured fish and used subsequently in stable isotope analysis (SIA). This scale collection was facilitated by the Environment Agency, the inland fishery regulatory body of England, who established a network of anglers within the area, where the participating anglers recorded their catches and were trained in the collection of scale samples that were used initially in age and growth analyses for fisheries management purposes (Nolan & Britton, 2019). Following their capture, the fish were identified to species, measured (fork length, nearest mm), and between 3 and 5 scales removed from the area below the dorsal fin and above the lateral line, and stored in a small paper envelope. In the same period, putative prey resources were sampled using long-handled sweep-nets, where macro-invertebrate samples were dominated by Gammaridae and Chironomidae. Also present in these samples were zebra mussels and macrophytes.
The main sampling period for site 2 was September 2018. Roach and perch were also sampled from the main river by angling, as sampling by typical fish capture methods (e.g. electric fishing, seine netting) was not feasible for the same reasons as per site 1. The captured fish were identified to species, measured for fork length (nearest mm) and a pelvic fin biopsy taken and frozen. Concomitantly, samples of amphipods were collected using a sweep net, with subsequent identification in the laboratory indicating the presence of species of Gammaridae and the invasive killer shrimp. This method was also used to collect samples of zebra mussels from colonised areas (e.g. boat moorings). There were few areas of macrophyte in the main river and so samples were not collected for subsequent analysis as per site 1. For bream, samples were also collected in the vicinity of the areas where the putative prey resources were collected using rod-and-line angling as per the roach and perch, with the same biometric data recorded and fin biopsy taken, but with the samples collected in April 2018. Only bream that subsequently foraged in the areas around the tagging site were used in analyses, as other individuals present were identified as spawning migrants from reaches further downstream and the River Thurne (Winter et al., 2021a, b, c).
Stable isotope analysis
For the fish at site 1, scales were the tissue used for SIA, with scale decalcification not performed prior to their analysis. This was because although comparisons of acidified versus non-acidified scales have revealed some differences in their isotopic data, the actual changes tend to be minor and not considered biologically relevant. For example, Ventura & Jeppesen (2010) demonstrated that the method resulted in mean changes in δ13C (± SD) of 0.18 ± 0.12 and in δ15N of −0.21 ± 0.24. The preparation of the scales thus comprised of cleaning with distilled water and then the removal of the outer portion of the scale for analysis. The latter process was completed to ensure the analysed tissue represented only the most recent growth of the fish (generally, the last full year of growth; Hutchinson & Trueman, 2006; Bašić & Britton, 2015). For the fish analysed, only one scale was used per individual, as this provided enough material for analysis. For fish at site 2, the fin biopsies were used for SIA. For the analyses of macro-invertebrates at both sites, three replicate samples were used per family, where a sample comprised of between one and three individuals (dependent on their body sizes). Zebra mussels were used individually, with preparation involving removal of their shell and a sample of soft tissue taken for SIA. All samples were dried to constant mass at 60 °C prior to their analysis at the Cornell Isotope Laboratory, New York, U.S.A. Stable isotope analytical details were as per Busst & Britton (2017), with lipid correction not necessary as C:N ratios indicated very low lipid content (< 4.0; Post et al., 2007).
Data analyses and mixing models
Differences in the stable isotope data between the omnivorous fish species per site were initially tested in ANCOVA, with fish fork length included as a covariate, with differences in length within the piscivorous fish then were tested in one-way ANOVA per site (α = 0.05 in all cases). For visual representation of the stable isotopes data per site, the isotopic niches of each fish species, expressed as standard ellipse areas (SEAc), were calculated in the R package ‘SIBER’ (Jackson et al., 2011), where the ellipses enclose 40% of the stable isotopes data and so represent the core niche of the species (Jackson et al., 2012).
The stable isotopes data for both sites were then applied to Bayesian mixing models which allowed for diet predictions based on posterior probability distributions of the contribution of relative proportions of the putative prey resources per fish species. For both sites, two models were run in the package MixSIAR (Stock et al., 2018) to predict the dietary contributions of the putative prey resources: (1) omnivorous fishes; and (2) piscivorous fishes. Separate models were run as these species required the use of different putative prey resources, with Carassou et al. (2008) suggesting that mixing models for piscivorous fish should use specific fractionation factors in models rather than use those used for dietary predictions of fishes that tend to feed primarily on invertebrates (Table 2). For the omnivores, bream and perch were the consumer species used in the models for site 1, and bream, perch and roach for site 2. For the piscivorous fishes, pikeperch were the consumer species used for site 1 and pike for site 2. Each model was then run at two levels: (1) at the population/community level (‘population’ hereafter; providing mean predicted dietary contributions per species); and (2) at the individual level (predicted diet composition per individual fish). All models were run using normal run length (chain length: 100,000 iterations with burn-in of 50,000, with posterior thinning (thin: 50) and 3 chains). Model diagnostics were based on Gelman-Rubin and Geweke (Stock & Semmens, 2016) to track algorithm convergence.
Mixing model prey resources and fractionation factors
For site 1, the putative prey resources used in the mixing models for the omnivorous fish species were macrophytes (‘plants’), macro-invertebrates (‘invertebrates’) and zebra mussel. For the ‘invertebrates’ prey resource, the stable isotopes data of Gammaridae and Chironomidae were tested for the significance of their differences using t-tests. Differences in δ13C were not significant (t-test: t12.8 = -1.58, p = 0.14), but were significant for δ15N (t12.4 = 2.41, p = 0.03) (Fig. 1a). However, the mean difference in δ15N between the groups of 0.73 ‰ was not considered biologically significant (Fig. 1a). Correspondingly, for their use as putative prey resources in the mixing model, their data were combined into a single resource. In these models, the isotopic fractionation values between the prey resources and consumers used were initially based on values generated from laboratory feeding trials of Busst & Britton (2017) (Table 2). However, it was decided these fractionation factors should use higher standard deviation values around the mean values in order to account for uncertainties relating to the different consumer species, and scales being used as the analysed tissue rather than fin tissue (Table 2). Moreover, different fractionation factors were used for the different prey resources, where the factors used were highest for plants and reduced for invertebrates and then zebra mussels/fish (Table 2). This was to account for patterns in the fractionation factors detected between prey resources of that differ in their protein content, where prey items of lower protein content tend to have higher fractionation factors (Busst & Britton, 2017; Table 2). For site 2, the putative prey resources differed to site 1, comprising Gammaridae, killer shrimp and zebra mussel, where the stable isotope values per resources were sufficiently different to enable them to be entered separately into models (Table 2). While the basis of the fractionation factors used at site 2 was as per site 1, the values used were different to site 1 as fin tissue was now the analysed tissue, which tends to have more depleted δ13C and enriched δ15N values than scales (Busst et al., 2015). For the piscivorous fish models, the same fractionation factors were used as per the omnivore models (Table 2). In site 1, the putative prey resources for pikeperch were freshwater prey fishes (common bream, perch and roach of lengths ≤ 170 mm), ‘invertebrates’ and zebra mussel. In site 2, the putative prey resources for pike were also freshwater prey fishes (common bream, perch and roach with length ≤ 170 mm), and then Gammaridae, killer shrimp and zebra mussel.
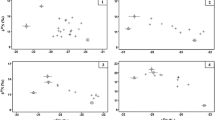
Stable isotope data and isotopic niches (as standard ellipse areas; SEAc) for the consumer species used in the stable isotope mixing models, where a site 1 (circle point, dashed ellipse: bream; triangle point, continuous ellipse: perch; square point, dotted ellipse: pikeperch); and b site 2 (circle point, continuous ellipse: bream; triangle point, dashed ellipse: perch; cross point, dotted ellipse: roach; square point, dashed-dotted ellipse: pike). Mean values are displayed for the putative prey resources (± SD)
All statistical analysis and graphical outputs were performed using R version 4.0.5 (R Core Team, 2021).
Results
Stable isotope data
For the analysed omnivorous fish species in site 1, perch were between 121 and 217 mm (mean ± SD: 172 ± 28 mm), whereas common bream were 127 to 476 mm (mean ± SD: 297 ± 126 mm). Differences in their stable isotope data were not significant between species (δ13C: F1,22 = 1.56, p = 0.22; δ15N: F1,22 = 0.46, p = 0.50; Fig. 1a), where the effect of fish length as a covariate was also not significant (δ13C: p = 0.33; δ15N: p = 0.87). For the piscivorous pikeperch in site 1, their lengths were 345 to 780 mm (mean 573 ± 107 mm), which had a significant effect on δ13C (F1,12 = 16.87, p < 0.01) but not δ15N (F1,12 = 0.08, p = 0.77).
For site 2, common bream lengths were 286 to 491 mm (mean: 414 ± 50 mm), perch were 121 to 224 mm (mean: 151 ± 32) and roach were 98 to 217 mm (143 ± 35). Differences between the species in their stable isotope data were not significant between species (δ13C: F1, 44 = 2.22, p = 0.12; δ15N: F1, 44 = 2.23, p = 0.12) (Fig. 1b), with the effect of fish length as a covariate also not significant (δ13C: p = 0.62; δ15N: p = 0.44). For the piscivorous pike, their lengths were 471 to 1014 mm (mean: 754 ± 147 mm), and length had a significant effect on δ13C (F1,13 = 5.39, p = 0.04), but not δ15N (F1,13 = 2.71, p = 0.12).
Stable isotope mixing models—omnivorous fishes
In site 1, the mixing models for bream and perch at both population and individual levels predicted that the ‘invertebrates’ prey resource had the highest contribution to the diets of both fish species, whereas zebra mussels were predicted to have a low dietary contribution irrespective of fish size (< 16%; Table 3, Fig. 2). In site 2, population models predicted killer shrimp was prey resource with the highest contribution to the diets of for bream and perch, followed by Gammaridae, and with zebra mussels again predicted to have a low dietary contribution. The individual models provided similar predictions, with Gammaridae predicted as the main prey resource for these two fish species, followed by killer shrimp (Table 4). Predictions for roach at the population level suggested a greater contribution of zebra mussels to diet than the other fish species (44%), but with the individual models suggesting Gammaridae was their main dietary source (Table 4). For roach, predictions suggested zebra mussels had relatively high dietary contributions for all fish sizes (Fig. 3).
Stable isotope mixing models—piscivorous fishes
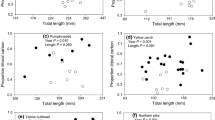
For pikeperch in site 1, the mixing models predicted that prey fish resources were had the highest dietary contributions at both the population level and individual level (Table 5), with this evident for all fish lengths present (Fig. 4a). Predictions for zebra mussel were 24% at the population level and 17% at the individual level (Table 5, Fig. 4a). For site 2, the predictions for pike also indicated that prey fish resources also contributed highly to their diet, irrespective of fish size, followed by killer shrimp (Table 5, Fig. 4b). Both zebra mussels and Gammaridae were predicted to contribute less than 18% of dietary resources (Table 5, Fig. 4b).
Estimated dietary prey contributions to the diet of individual piscivorous fishes by body length (0–1): a pikeperch with fish (dark grey), invertebrates (light grey) and zebra mussel (black), and b pike with fish (dark grey), Gammaridae (white), killer shrimp (light grey) and zebra mussels (black). Prey source shown as overall mean per individual
Discussion
The increased distribution of zebra mussels in British freshwaters (Aldridge et al., 2004) has presented a novel prey resource for the native fish communities. Across the five fish species analysed here, only roach, a fish ubiquitous to British lowland freshwaters (Britton, 2007), had predictions that zebra mussels were contributing strongly to their diet. The population level model prediction of 44% is consistent with Prejs et al. (1990), who suggested a predator–prey interaction between the two species, and Millane et al. (2012) who detected roach consumed them in Lough Sheelin, Ireland, in all seasons. For the other fish species, zebra mussels were never predicted to comprise of more than 29% of their diets at the population level.
Common bream of over 400 mm were present in samples taken from both sites, with their diets predicted to primarily consist of ‘invertebrate’ prey resources (covering Gammaridae, Chironomidae and killer shrimp, depending on the site). These results are consistent with other dietary studies on common bream that have highlighted the importance of macro-invertebrates as prey resources for the species, with Chironomidae often being a dominant prey source (e.g. Giles et al., 1990; Kakareko, 2001). Although not sampled here, zooplankton can also be important in their diet, although usually only in fish of below 200 mm and in situations where there is a lack of macro-invertebrates available (Kakareko, 2001). Zebra mussels were predicted to be only a minor component of their diet in both sites here, with this potentially related to their extensive branchial sieve which could make difficult to for them to ingest molluscs and limitations on their ability to crush the mussels imposed by the architecture of their pharyngeal teeth (Nagelkerke & Sibbing, 2011). Similarly, the analysed perch primarily consumed the ‘invertebrates’ prey resource at the population level, with this also consistent with other studies suggesting the importance of similar prey resources in their diet (Giles et al., 1990; Dörner et al., 2003). While there were no studies suggesting perch regularly prey upon zebra mussel, Mavrin & Strel’nikova (2011) suggested that when the species are in sympatry, with zebra mussels feeding extensively on zooplankton, juvenile perch might have to switch to macro-invertebrate prey from their early stages. This was also noted by Shcherbina & Bezmaternykh (2019), who also suggested that zebra mussel colonies also provide refuge for macro-invertebrates, increasing the difficulty of prey detection for perch during foraging. Roach were suggested in the literature as an important consumer of zebra mussels (Prejs et al., 1990; Millane et al., 2012) and, while they were only analysed in one site, the diet predictions here suggested all of the analysed fish had consumed some zebra mussels in recent months. However, this included individuals of below 160 mm length, whereas empirical studies have suggested a clear size threshold where these fish would not generally prey upon them (Prejs et al., 1990). Nagelkerke & Sibbing (2011) suggested that, functionally, roach are ‘superior’ in static crushing hard and brittle materials (like the shells of zebra mussels), compared to species such as common bream, emphasising those functional attributes such as gape size, strong pharyngeal teeth and the ability of chew prey items could be important for efficient feeding on these bivalves (French, 1993).
For piscivorous fish species, only fish above 170 mm were analysed, a size when the species tends to be an obligate piscivore (e.g. Nolan & Britton, 2018). Pikeperch primarily fed upon freshwater fish (70% at both population and individual levels), with a low contribution of the ‘invertebrates’ resource (< 10% both levels). Pike also had freshwater fish as their most important dietary resource, but with the alien killer shrimp also important in their diet. Recent studies on pike diet based on stable isotope data have indicated that macro-invertebrates can comprise a key component of their diet, irrespective of their body length, indicating some functional plasticity (Chapman & Mackay, 1990; Pedreschi et al., 2015; Nolan et al., 2019). Moreover, invasive killer shrimp have been detected as displacing native amphipod species in invaded waters, with fish species (including perch) responding by replacing native amphipods in their diet with the invasive ones (Eckmann et al., 2008). In site 2, invasive killer shrimp were in sympatry with native amphipods and with both making substantial contributions to fish diet generally. Correspondingly, the influence of this Ponto-Caspian amphipod on the diet of freshwater fish in Britain might prove to be higher than zebra mussel.
These results suggest that roach was the only analysed fish species that used zebra mussels within their diet in more than a minor way, suggesting that the extent of consumptive resistance on zebra mussel colonisation and spread was likely to be limited in the analysed sites. While roach are ubiquitous in most regions of lowland Britain, populations tend to be dominated by fish below 200 mm (Britton, 2007), which are unlikely to be capable of consuming large quantities of bivalves. Similarly, while French & Bur (1992) revealed that although freshwater drum Aplodinotus grunniens predated heavily on these molluscs, they suggested this predation pressure would have little effect on the invasion of zebra mussels due to their rapid reproduction and growth. This suggests that where there is a management requirement to prevent the establishment of zebra mussel populations, or reduce their abundance, then reliance on biological resistance and/or biological control methods is not recommended. Thus, the application of chemical control methods would be required instead, but while these can substantially reduce their populations, these methods tend to also cause a high degree of collateral damage in non-target species (Aldridge et al., 2004; Lydeard et al., 2004). Awareness on the use of biosecurity (i.e. cleaning and drying techniques) to prevent introductions, rather than manage new and on-going invasions, tends to be used more, but clearly does not reduce their impacts in already invaded waters (Melly & Hanrahan, 2020; Smith et al., 2020).
Conclusion
The results from two invaded sites in England, one in the east and one in the west, suggest that in lowland areas at least, the consumptive resistance to the invasion of zebra mussels in Britain appears to have been limited. While it is acknowledged that the spatial and temporal extent of our sampling was limited, the results were largely consistent with studies elsewhere in the invaded range of zebra mussels, suggesting that while more in-depth studies might be interesting, they are unlikely to provide any novel insights. However, our analyses were based only on stable isotope analysis, with other methods of dietary analyses not being used, such as stomach contents analyses, which can provide both consistent (e.g. Nolan & Britton, 2018) and contrary results (e.g. Locke et al., 2013; Hamidan et al., 2016). Nevertheless, these results do suggest that if the invasion of Britain by zebra mussels is going to be contained, a more active management approach must be taken to prevent spread of existing populations and the establishment of new ones.
Data availability
The datasets generated and analysed here are available from the corresponding author at reasonable request.
References
Aldridge, D. C., P. Elliott & G. D. Moggridge, 2004. The recent and rapid spread of the zebra mussel (Dreissena polymorpha) in Great Britain. Biological Conservation 119: 253–261.
Alofs, K. M. & D. A. Jackson, 2014. Meta-analysis suggests biotic resistance in freshwater environments is driven by consumption rather than competition. Ecology 95: 3259–3270.
Bašić, T. & J. R. Britton, 2015. Utility of fish scales from stock assessment surveys in stable isotope analysis for initial assessments of trophic relationships in riverine fish communities. Journal of Applied Ichthyology 31: 296–300.
Britton, J. R., 2007. Reference data for evaluating the growth of common riverine fishes in the UK. Journal of Applied Ichthyology 23: 555–560.
Britton, J. R., 2012. Testing strength of biotic resistance against an introduced fish: inter-specific competition or predation through facultative piscivory? PLoS ONE 7: e31707.
Britton, J. R. & R. E. Gozlan, 2013. How many founders for a biological invasion? Predicting introduction outcomes from propagule pressure. Ecology 94: 2558–2566.
Britton, J. R., C. Gutmann Roberts, F. Amat Trigo, E. T. Nolan & V. De Santis, 2019. Predicting the ecological impacts of an alien invader: experimental approaches reveal the trophic consequences of competition. Journal of Animal Ecology 88: 1066–1078.
Busst, G. M. & J. R. Britton, 2015. Quantifying the growth consequences for crucian carp Carassius carassius of competition from non-native fishes. Ecology of Freshwater Fish 24: 489–492.
Busst, G. M. & J. R. Britton, 2017. Comparative trophic impacts of two globally invasive cyprinid fishes reveal species-specific invasion consequences for a threatened native fish. Freshwater Biology 62: 1587–1595.
Byun, C., S. de Blois & J. Brisson, 2020. Restoring functionally diverse communities enhances invasion resistance in a freshwater wetland. Journal of Ecology 108: 2485–2498.
Carassou, L., M. Kulbicki, T. J. R. Nicola & N. V. C. Polunin, 2008. Assessment of fish trophic status and relationships by stable isotope data in the coral reef lagoon of New Caledonia, southwest Pacific. Aquatic Living Resources 21: 1–12.
Catford, J. A., R. Jansson & C. Nilsson, 2009. Reducing redundancy in invasion ecology by integrating hypotheses into a single theoretical framework. Diversity and Distributions 15: 22–40.
Chapman, L. J. & W. C. Mackay, 1990. Ecological Correlates of Feeding Flexibility in Northern Pike (Esox lucius). Journal of Freshwater Ecology 5: 313–322.
Crystal-Ornelas, R. & J. L. Lockwood, 2020. The ‘known unknowns’ of invasive species impact measurement. Biological Invasions 22: 1513–1525.
Dawson, W., D. Moser, M. Van Kleunen, H. Kreft, J. Pergl, P. Pyšek, P. Weigelt, M. Winter, B. Lenzner, T. M. Blackburn & E. E. Dyer, 2017. Global hotspots and correlates of alien species richness across taxonomic groups. Nature Ecology & Evolution 1: 1–7.
Diagne, C., B. Leroy, A. C. Vaissière, R. E. Gozlan, D. Roiz, I. Jarić, J. M. Salles, C. J. Bradshaw & F. Courchamp, 2021. High and rising economic costs of biological invasions worldwide. Nature 592: 571–576.
Dominguez Almela, V., S. C. Palmer, P. K. Gillingham, J. M. Travis & J. R. Britton, 2020. Integrating an individual-based model with approximate Bayesian computation to predict the invasion of a freshwater fish provides insights into dispersal and range expansion dynamics. Biological Invasions 22: 1461–1480.
Dörner, H., S. Berg, L. Jacobsen, S. Hülsmann, M. Brojerg & A. Wagner, 2003. The feeding behaviour of large perch Perca fluviatilis (L.) in relation to food availability: a comparative study. Hydrobiologia 506: 427–434.
Eckmann, R., M. Mörtl, D. Baumgärtner, C. Berron, P. Fischer, D. Schleuter & A. Weber, 2008. Consumption of amphipods by littoral fish after the replacement of native Gammarus roeseli by invasive Dikerogammarus villosus in Lake Constance. Aquatic Invasions 3: 187–191.
French III, J.R.P, & M.T. Bur, 1992. Predation of the zebra mussel (Dreissena polymorpha) by freshwater drum in western Lake Erie. In Nalepa, T.F. & D.W. Schloesser (eds), Publication of an organization other than the U.S. geological survey. Lewis Publishers: 453–463
French III, J. R. P., 1993. How Well Can Fishes Prey on Zebra Mussels in Eastern North America? Fisheries 18: 13–19.
Gallardo, B. & D. C. Aldridge, 2013. Priority setting for invasive species management: risk assessment of Ponto-Caspian invasive species into Great Britain. Ecological Applications 23: 352–364.
Gallardo, B. & D. C. Aldridge, 2015. Is Great Britain heading for a Ponto-Caspian invasional meltdown? Journal of Applied Ecology 52: 41–49.
Ghedotti, M. J., J. C. Smihula & G. R. Smith, 1995. Zebra mussel predation by round gobies in the laboratory. Journal of Great Lakes Research 21: 665–669.
Giles, N., M. Street & R. M. Wright, 1990. Diet composition and prey preference of tench, Tinca tinca (L.), common bream, Abramis brama (L.), perch, Perca fluviatilis L. and roach, Rutilus rutilus (L.), in two contrasting gravel pit lakes: potential trophic overlap with wildfowl. Journal of Fish Biology 37: 945–957.
Gozlan, R. E., 2008. Introduction of non-native freshwater fish: is it all bad? Fish and Fisheries 9: 106–115.
Hansen, G. J., T. D. Ahrenstorff, B. J. Bethke, J. D. Dumke, J. Hirsch, K. E. Kovalenko, J. F. LeDuc, R. P. Maki, H. M. Rantala & T. Wagner, 2020. Walleye growth declines following zebra mussel and Bythotrephes invasion. Biological Invasions 22: 1481–1495.
Hamidan, N., M. C. Jackson & J. R. Britton, 2016. Diet and trophic niche of the endangered fish Garra ghorensis in three Jordanian populations. Ecology of Freshwater Fish 25: 455–464.
Higgins, S. N. & M. J. Vander Zanden, 2010. What a difference a species makes: a meta–analysis of dreissenid mussel impacts on freshwater ecosystems. Ecological Monographs 80: 179–196.
Hutchinson, J. J. & C. N. Trueman, 2006. Stable isotope analyses of collagen in fish scales: Limitations set by scale architecture. Journal of Fish Biology 69: 1874–1880.
Jackson, A. L., A. C. Parnell, R. Inger & S. Bearhop, 2011. Comparing isotopic niche widths among and within communities: SIBER – Stable Isotope Bayesian Ellipses in R. Journal of Animal Ecology 80: 595–602.
Jackson, M. C., I. Donohue, A. L. Jackson, J. R. Britton, D. M. Harper & J. Grey, 2012. Population-level metrics of trophic structure based on stable isotopes and their application to invasion ecology. PLoS ONE 7: e31757.
Kakareko, T., 2001. The diet, growth and condition of common bream, Abramis brama (L.) in wŁocŁawek reservoir. Acta Ichthyologica Et Piscatoria 31: 37–53.
Locke, S. A., G. Bulte, M. R. Forbes & D. J. Marcogliese, 2013. Estimating diet in individual pumpkinseed sunfish Lepomis gibbosus using stomach contents, stable isotopes and parasites. Journal of Fish Biology 82: 522–537.
Lydeard, C., R. H. Cowie, W. F. Ponder, A. Bogan, P. Bouchet, S. Clark, K. Cummings, T. Frest, O. Gargominy & D. Herbert, 2004. The global decline of nonmarine mollusks. BioScience 54: 321–330.
Magoulick, D. D. & L. C. Lewis, 2002. Predation on exotic zebra mussels by native fishes: effects on predator and prey. Freshwater Biology 47: 1908–1918.
Mavrin, A. S. & A.P Strel’nikova, 2011. Feeding, development, and growth of juvenile perch perca fluviatilis in mesocosms in the presence of filter-feeding zebra mussel Dreissena polymorpha pallas. Inland Water Biology 4: 232–241.
Melly, D. & J. Hanrahan, 2020. Tourist biosecurity awareness and risk mitigation for outdoor recreation: Management implications for Ireland. Journal of Outdoor Recreation and Tourism 31: 100313.
Millane, M., M. O’Grady, K. Delanty & M. Kelly-Quinn, 2012. An assessment of fish predation on the zebra mussel, Dreissena Polymorpha (Pallas 1771) after recent colonisation of two managed brown trout lake fisheries in Ireland. Biology and Environment: Proceedings of the Royal Irish Academy 112B: 1–9.
Nagelkerke, L. & F. A. Sibbing, 2011. Efficiency of feeding on zebra mussel (Dreissena polymorpha) by common bream (Abramis brama), white bream (Blicca bjoerkna), and roach (Rutilus rutilus): The effects of morphology and behavior. Canadian Journal of Fisheries and Aquatic Sciences 53: 2847–2861.
Natural England, 2020. Designated Sites View. https://designatedsites.naturalengland.org.uk/ [Accessed on 23/07/20]
Nolan, E. T. & J. R. Britton, 2018. Diet of invasive pikeperch Sander lucioperca: developing non-destructive tissue sampling for stable isotope analysis with comparisons to stomach contents analysis. Knowledge & Management of Aquatic Ecosystems 419: 49.
Nolan, E. T. & J. R. Britton, 2019. Spatial variability in the somatic growth of pikeperch Sander lucioperca, an invasive piscivorous fish. Ecology of Freshwater Fish 28: 330–340.
Nolan, E. T., C. Gutmann Roberts & J. R. Britton, 2019. Predicting the contributions of novel marine prey resources from angling and anadromy to the diet of a freshwater apex predator. Freshwater Biology 64: 1542–1554.
Pedreschi, D., S. Mariani, J. Coughlan, C. C. Voigt, M. O’Grady, J. Caffrey & M. Kelly-Quinn, 2015. Trophic flexibility and opportunism in pike Esox lucius. Journal of Fish Biology 87: 876–894.
Post, D. M., C. A. Layman, D. A. Arrington, G. Takimoto, J. Quattrochi & C. G. Montana, 2007. Getting to the fat of the matter: models, methods and assumptions for dealing with lipids in stable isotope analyses. Oecologia 152(1): 179–189.
Prejs, A., K. Lewandowski & A. Stańczykowska-Piotrowska, 1990. Size-selective predation by roach (Rutilus rutilus) on zebra mussel (Dreissena polymorpha): field stuides. Oecologia 83: 378–384.
R Core Team, 2021. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/.
Rennie, M. D., D. O. Evans & J. D. Young, 2013. Increased dependence on nearshore benthic resources in the Lake Simcoe ecosystem after dreissenid invasion. Inland Waters 3: 297–310.
Ricciardi, A. & H. J. MacIsaac, 2000. Recent mass invasion of the North American Great Lakes by Ponto-Caspian species. Trends in Ecology & Evolution 15: 62–65.
Robertson, J. J., T. M. Swannack, M. McGarrity & A. N. Schwalb, 2020. Zebra mussel invasion of Texas lakes: estimating dispersal potential via boats. Biological Invasions 22: 3425–3455.
Shcherbina, G. K. & V. Bezmaternykh, 2019. Effect of zebra mussel Dreissena polymorpha (Pallas, 1771) (Mollusca, Dreissenidae) and perch Perca fluviatilis (L.) (Pisces, Perciidae) of Different Age Groups on the Structure and Main Characteristics of Macrozoobenthos in Experimental Mesocosms. Inland Water Biology 12: 190–198.
Silverman, H., J. W. Lynn, E. C. Achberger & T. H. Dietz, 1996. Gill Structure in Zebra Mussels: Bacterial-Sized Particle Filtration. American Zoologist 36: 373–384.
Simberloff, D., J. L. Martin, P. Genovesi, V. Maris, D. A. Wardle, J. Aronson, F. Courchamp, B. Galil, E. García-Berthou, M. Pascal & P. Pyšek, 2013. Impacts of biological invasions: what’s what and the way forward. Trends in Ecology & Evolution 28: 58–66.
Smith, E. R. C., H. Bennion, C. D. Sayer, D. C. Aldridge & M. Owen, 2020. Recreational angling as a pathway for invasive non-native species spread: awareness of biosecurity and the risk of long distance movement into Great Britain. Biological Invasions 22: 1135–1159.
Stock, B. C. & B. X. Semmens, 2016. Unifying error structures in commonly used biotracer mixing models. Ecology 97(10): 2562–2569.
Stock, B. C., A. L. Jackson, E. J. Ward, A. C. Parnell, D. L. Phillips & B. X. Semmens, 2018. Analyzing mixing systems using a new generation of Bayesian tracer mixing models. PeerJ 6: 5096.
Strayer, D. L., 2009. Twenty years of zebra mussels: lessons from the mollusk that made headlines. Frontiers in Ecology and the Environment 7: 135–141.
Turbelin, A. J., B. D. Malamud & R. A. Francis, 2017. Mapping the global state of invasive alien species: patterns of invasion and policy responses. Global Ecology and Biogeography 26: 78–92.
Ventura, M. & E. Jeppesen, 2010. Evaluating the need for acid treatment prior to δ13C and δ15N analysis of freshwater fish scales: Effects of varying scale mineral content, lake productivity and CO2 concentration. Hydrobiologia 644(1): 245–259.
Winter, E. R., A. M. Hindes, S. Lane & J. R. Britton, 2021a. Movements of common bream Abramis brama in a highly connected, lowland wetland reveal sub-populations with diverse migration strategies. Freshwater Biology 66: 1410–1422.
Winter, E. R., A. M. Hindes, S. Lane & J. R. Britton, 2021b. Acoustic telemetry reveals strong spatial preferences and mixing during successive spawning periods in a partially migratory common bream population. Aquatic Sciences 83: 1–14.
Winter, E. R., A. M. Hindes, S. Lane & J. R. Britton, 2021c. Dual-isotope isoscapes for predicting the scale of fish movements in lowland rivers. Ecosphere 12: e03456.
Wong, C. H. S., E. C. Enders & C. T. Hasler, 2021. Limited evidence of zebra mussel (Dreissena polymorpha) consumption by freshwater drum (Aplodinotus grunniens) in Lake Winnipeg. Journal of Great Lakes Research 47: 592–602.
Acknowledgements
We acknowledge the assistance of Andrew Hindes, Steve Lane and Brecht Morris for their help in coordinating the fish sampling. We thank all of the anglers involved in the fish sampling.
Funding
VDA was supported by an iCASE studentship from the Natural Environment Research Council (Grant Number NE/R008817/1) and the Environment Agency; EW was supported by the EU LIFE + Nature and Biodiversity Programme: LIFE14NAT/UK/000054, as well as funding and resource support from the Environment Agency; and EN was supported by the Environment Agency and Severn Rivers Trust.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study design. EN, EW and JRB collected the data. VDA led analyses and writing, assisted by EN, EW and JRB. All authors contributed to revising the manuscript and approved its submission.
Corresponding author
Ethics declarations
Conflict of interest
None of the co-authors have a conflict/competing interest.
Ethical approval
All regulated procedures were completed under UK Home Office project licence 70/8063 and after ethical review.
Informed consent
All authors reviewed the manuscript and agreed with its contents.
Additional information
Handling editor: Manuel Lopes Lima
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dominguez Almela, V., Nolan, E.T., Winter, E.R. et al. Dietary contributions of the alien zebra mussel Dreissena polymorpha in British freshwater fish suggest low biological resistance to their invasion. Hydrobiologia 849, 2253–2265 (2022). https://doi.org/10.1007/s10750-022-04861-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-022-04861-5