Abstract
Recent research hypothesised that the Siculo-Tunisian Strait might fit, at least for some species, the picture of a genetic transitional zone instead of a sharp genetic break between the Western and Eastern Mediterranean basins. The present study aimed at using the common Octopus, Octopus vulgaris as an empirical test-case to evaluate this hypothesis. To accomplish this goal, 458 new sequences of the mitochondrial gene cytochrome c oxidase subunit I were used. Combining the new sequences with those available on public databases, we assembled a dataset containing 920 sequences to investigate the spatial genetic patterns across 34 Mediterranean populations of O. vulgaris. The genetic structure of this species was assessed combining analysis of molecular variance and Median-Joining networks. Results supported the hypothesis of a complex spatial genetic pattern across the Sicilian channel. Contemporary factors, such as marine currents, likely affect the species’ genetic structuring across this area. Overall, our results highlighted that focusing the attention on the whole transitional area rather than on a unique genetic break might help to detect similar patterns across different species. Finally, acknowledging the occurrence of complex spatial genetic patterns across transitional zones may improve stock identification and management practices for commercially valuable species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Improving the knowledge on species’ spatial genetic patterns is crucial for future conservation and management plans, with an emphasis on those commercial species for which management strategies have not been sufficiently enforced (Lleonart & Maynou, 2003). This task is particularly important in the Mediterranean Sea, where anthropogenic pressures and ongoing climate change are likely to have large and increasing impacts on marine biodiversity (Lejeusne et al., 2010; Coll et al., 2012). The Mediterranean Sea represents a natural focus of research interest for marine biodiversity related to other World’s regions (Bianchi, 2007; Lejeusne et al., 2010; Coll et al., 2012). In fact, the Mediterranean harbors, on average, 7% of total World marine biodiversity, with high proportions of both species’ richness and endemics, despite it represents only 0.8% and 0.3% of the World ocean area and volume, respectively (Coll et al., 2012 and references therein). Moreover, the semi-enclosed and highly compartmentalized nature of the Mediterranean, make it a potential model to study marine ecosystems (Lejeusne et al., 2010; Coll et al., 2012). Because of the complex topography, the variable oceanographic and ecological features, the present-day variety of climatic conditions, and the complex geological history, the Mediterranean is subdivided into distinct sub-basins (Bas, 2009), marine ecoregions (Spalding et al., 2007) and biogeographic sectors (Bianchi, 2007). Thus, the Mediterranean offers the opportunity of investigating how the interplay between abiotic and biological factors shape interspecific as well as intraspecific diversity across transitional zones (Patarnello et al., 2007).
From a biogeographic perspective, increasing attention has been long paid to the boundary between the Western and the Eastern basins of the Mediterranean Sea, which lies between the Siculo-Tunisian strait (STS) and the eastern limit of the Ionian Sea (Villamor et al., 2018). Based on evidence from literature, Bianchi (2007) outlined four main hypotheses for the placement of the boundary between the Western and the Eastern Mediterranean basins. In the first hypothesis (i), the boundary was placed in the mid Ionian Sea, thus including Sicily, Calabria and the Gulf of Taranto in the Western Mediterranean basin. Alternatively (hypothesis ii), the boundary was placed to the middle of STS, with the Pantelleria Island included in the Western Mediterranean, whilst the Pelagie and Malta Islands in the Eastern Mediterranean. Another scenario (hypothesis iii) placed the boundary so that Western Mediterranean included yet the whole Sicily but not the Ionian coast of Calabria and the Gulf of Taranto. Finally, the location of the boundary was proposed to be at the Peloponnese and the Aegean Sea, rather than at the STS (hypothesis iv). From the genetic viewpoint, the picture is much more complex: most of investigations highlighted divergence between Western and Eastern Mediterranean populations, although other subdivisions of the Mediterranean Sea might have been overlooked (e.g., Borrero-Pérez et al., 2011; Mokhtar-Jamaï et al., 2011; Cossu et al., 2015). For instance, two genetic breaks (the Almeria-Oran front and the Gulf of Bejaia) delimit a wide mosaic hybrid zone along Algerian coasts, in which deeply divergent lineages of Mytilus galloprovincialis Lamarck, 1819 mix (El Ayari et al., 2019). Moreover, apart from the recognition of biogeographic subdivisions, tracing the precise boundary between the two major Mediterranean basins is still a debated issue. In fact, the location of genetic breaks varies among species, ranging from the STS (Borsa et al., 1997; Tarnowska et al., 2010; Souissi et al., 2018) to east off the Sicilian coasts (Sá-Pinto et al., 2010, 2012; Sanna et al., 2013) and even further east in the mid or eastern Ionian Sea (Boissin et al., 2011; Ragionieri & Schubart, 2013; Villamor et al., 2014; Deli et al., 2018). There are also species that do not show genetic structuring across this boundary area, including fishes (Fruciano et al., 2011; Schunter et al., 2011a, b; Barbieri et al., 2014) and marine invertebrates (Maltagliati et al., 2010; Fratini et al., 2016; Pannacciulli et al., 2017). Likely, this variety of outcomes reflects the interplay between the species’ evolutionary histories (Patarnello et al., 2007), the life-history traits and environmental factors such as oceanographic features or inadequate sampling (Pascual et al., 2017). In light of the above considerations, the whole area encompassing the south Tyrrhenian Sea, the Sicilian channel and the Ionian Sea could be viewed as a “transitional zone” between the Western and Eastern Mediterranean basins, which is characterised by heterogeneous levels of admixture in the area, as proposed by Villamor et al. (2018). According to these authors, this picture would fit better than a sharp genetic break, whereby there is an abrupt shift of allele frequencies at a geographic location, to explain the complex genetic patterns reported in this area so far.
In the present study, we used the common octopus, Octopus vulgaris Cuvier 1797 (Mollusca: Cephalopoda), as an empirical test-case of the hypothesis advanced by Villamor et al. (2018). The common octopus is a benthic cephalopod of commercial interest distributed in the whole Mediterranean Sea, Atlantic Ocean and North-West Pacific (Warnke et al., 2004). Although O. vulgaris is likely a complex of cryptic species rather than a widespread cosmopolitan species (Sales et al., 2013; De Luca et al., 2014), Mediterranean populations of O. vulgaris belong unambiguously to a single species (De Luca et al., 2016; Melis et al., 2018). The common octopus lives in a variety of habitats, such as rocky, sandy and muddy bottoms as well as seagrass beds, from the coastline to the outer edge of continental shelf at depths of approximately 200 m (Belcari et al., 2002). The species has a short-life cycle (9-15 months) including a paralarval stage (Smale & Buchan, 1981; Hernández-López et al., 2001). Paralarvae, which are morphologically very similar to adults, hatch from the eggs and play an important role in dispersal as they conduct an active pelagic life for 35–60 days depending on water temperature (Villanueva et al., 1997; Katsanevakis & Verriopoulos, 2006; O’Connor et al., 2007). Therefore, gene flow in O. vulgaris is likely to be affected by oceanographic features and biogeographic barriers (Maltagliati et al., 2002). In contrast to paralarvae, adult migrations are of very restricted range, including daily migrations for forage (Mather, 1994) and seasonal migrations of females in relation to reproductive behavior (Smale & Buchan, 1981). Reasonably, the contribution of adult migrations to gene flow can be considered negligible if compared to paralarval dispersal.
As O. vulgaris is a commercially valuable species in Mediterranean fisheries, species’ patterns of genetic diversity have been widely investigated from regional-to basin-scale level in the Mediterranean region (Casu et al., 2002; Maltagliati et al., 2002; Cabranes et al., 2008; Fadhlaoui-Zid et al., 2012; De Luca et al., 2016; Melis et al., 2018). Overall, the hypothesis of a single panmictic unit within the Mediterranean Sea was rejected. For instance, Melis et al. (2018) found shallow but significant genetic structuring at regional scale along Sardinian coasts by using microsatellites. De Luca et al. (2016) highlighted the strong genetic divergence of a population from the Strait of Messina to the other Mediterranean populations, which might clue into the isolation of the marine biota within this strait (Bianchi, 2007). On the other hand, an isolation-by-distance driven pattern of genetic variation was inferred along Spanish coasts (Cabranes et al., 2008). At larger spatial scales, the presence of a genetic break between the Western and Eastern Mediterranean basins was evidenced using allozymes (Maltagliati et al., 2002), and cytochrome c oxidase subunit III (Fadhlaoui-Zid et al., 2012) but it was not clearly supported by microsatellites (De Luca et al., 2016).
The objective of the present study was to investigate the spatial genetic structure of O. vulgaris across the Mediterranean Sea, focusing our attention on the Siculo-Tunisian Strait and the surrounding areas. We aimed at ascertaining whether spatial genetic variation fits a scenario characterized by a sharp phylogeographic break, taking into account two potential locations of it (namely Bianchi’s hypotheses ii and iii) or, alternatively, a transitional area with smoothed genetic change. To achieve this goal, we used the mitochondrial gene cytochrome c oxidase subunit I (COI), which is effective to infer both the historical and contemporary processes shaping the spatial genetic patterns in various marine invertebrates (Sanna et al., 2013, 2014; Cordero et al., 2014; Cossu et al., 2015; Pannacciulli et al., 2017; Villamor et al., 2018).
Materials and methods
COI amplification and sequencing
COI fragments of 641 bp were amplified from genomic DNA stored at – 20 °C using Folmer et al. (1994) primers LCO1490 (5′- GGTCAACAAATCATAAAGATATTGG- 3′) and HCO2198 (5′- TAAACTTCAGGGTGACCAAAAAATCA-3′). Each 25 µl PCR mixture contained approximately 100 ng of total genomic DNA, 0.40 µM of each primer, 1.75 U of Taq DNA Polymerase (Sigma-Aldrich®), 1 × reaction buffer and 200 µM of each dNTP. MgCl2 concentration was set at 2 mM, and 12.5 mg of BSA were added to the reaction mixture. PCRs were performed according to the following steps: 1 cycle of 3 min at 94°C, 35 cycles of 40 s at 94°C, 45 s at 51°C and 1 min and 40 s at 72°C, with a final extension performed at 72°C for 5 min. After that, PCR samples were cooled to 4°C. Negative controls were included for each primer to check for contamination. After electrophoresis on 2% agarose gels, the PCR products were purified using ExoSAP-IT (USB Corporation, Santa Clara, California, USA) and sequenced using an external sequencing core service (Macrogen Europe, Amsterdam, Netherlands). Dual peaks of similar height, which could be interpreted as evidence of mitochondrial pseudogenes in the nucleus (Numts) or heteroplasmy, were not observed in any of the electropherograms. The PCR products did not show any occurrence of aspecificity, excluding the possibility of multiple nuclear mtDNA-like sequences.
Dataset assembling
A dataset containing 920 partial sequences of the mitochondrial gene cytochrome c oxidase subunit I (COI) was assembled. Of these, 458 COI fragments (Table 1) were newly sequenced from genomic DNA obtained from previous studies (Casu et al., 2002; Fadhlaoui-Zid et al., 2012), whereas the remaining sequences were retrieved from Genbank (Fadhlaoui-Zid et al., 2013; Amor et al., 2014; Melis et al., 2018; Jassoud et al., unpubl. data), Barcode Of Life Data System (BOLD, Roura et al., 2019) and from supplementary material (De Luca et al., 2016). Only localities with a sample size of at least five sequences were retained in the present study. Overall, the dataset encompassed 34 Mediterranean localities (Online Resource 1: Table S1, Fig. 1).
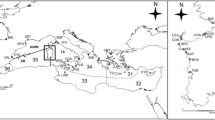
Sampling locations of Octopus vulgaris. Open circles mark localities at which Cytochrome c oxidase subunit I (COI) was newly sequenced, whereas closed circles mark localities at which COI partial sequences were retrieved from Genbank (see Online Resource 1: Table S1 for further details). Gray dotted lines outline three hypotheses advanced for the location of the biogeographic boundary between Western and Eastern Mediterranean (modified from Bianchi 2007). The inset shows the localities sampled in Sardinia. SPN Valencia, BAR Barcelona, PER Perpignan, SET Sete, ALG Alghero 2010-2012, AHO Alghero 1999, PTO Porto Torres, CSA Castelsardo, SUP Su Pallosu, ORI Oristano, TEU Teulada, GOL Gulf of Cagliari, OLB Olbia, SIN Siniscola, CFE Capo Ferrato, GAI Galite island, LEG Livorno, PSS Porto Santo Stefano, NA1 Napoli 2015 station 1, NA2 Napoli 2015 station 2, NAP Napoli 1999, STM Strait of Messina, SSI Channel of Sicily, MAL Malta, PPA Porto Palo, PCS Porto Cesareo, CKI Kerkennah island coastal waters, DWK Kerkennah island deep waters, ZAR Zarzis, TAJ Tajoura, ANC Ancona, CRZ Split, CRE Heraklion, SMS Samos
Data analysis
Sequences were visualised and checked with Chromas Pro 1.7.6 (http://technelysium.com.au) and aligned using the CLUSTAL W algorithm (Thompson et al., 1994) implemented in the software BioEdit 7.2.3 (Hall, 1999). Neither insertions nor deletions were found in the final alignment. Only one new haplotype was found in the Mediterranean, which was deposited in GenBank with the following accession number: MT812944.
The software POWSIM 4.1 (Ryman & Palm, 2006) adjusted for mtDNA data (Larsson et al., 2009) was used for testing the power of detecting population differentiation. For each independent simulation, 1000 pseudo-datasets simulating the same number of populations (N), haplotypes and population sampling size (S) as the real dataset were created. Each simulation tested the power of detecting population differentiation at a different level of genetic differentiation (FST = 0 to 0.2), whose range was chosen based upon observed pairwise FST values (Fig. 2). Two further simulation sets investigated the power of detecting population differentiation at small and uniform sampling sizes (S = 5 and 10). All simulations were run assuming a constant effective population size of 2000 individuals (Ne), no migration across demes, and varying the number of generations of genetic drift (T) in order to obtain the expected level of genetic differentiation at equilibrium. Finally, the effect of N and S on the expected level of statistical type II error (the proportion of simulations in which population differentiation is detected when FST = 0, which is expected to be around 5%) was assessed. Three independent sets of simulations (N = 10, 20, 30) were carried out varying the sampling size (S = 5 to 50 by steps of five individuals) and keeping generations of genetic drift constant (T = 0).
Analysis of molecular variance (AMOVA), as implemented in Arlequin 3.5.2.3 software (Excoffier & Lischer, 2010), was used to partition the genetic variance in the within- and among-sample components. Then, different grouping criteria were adopted depending on the Western-Eastern Mediterranean boundary considered, namely the boundary located at the STS, and off the eastern Sicilian coast (hypotheses ii and iii in Bianchi, 2007). Estimates of genetic divergence among all groups were calculated using Φ-statistics. Three hierarchical levels of subdivision were obtained: ΦCT (the degree of differentiation among basins), ΦSC (the degree of differentiation among samples within basins), and ΦST (the degree of differentiation among samples). Φ-statistics were computed using the genetic distances estimated with the Tamura (1992) model of sequence evolution, selected with the Bayesian information criterion (BIC) in MEGA 7.0.14 (Kumar et al., 2016). In addition to the Φ-statistics, which includes information on both haplotype frequency and genealogy, the Weir & Cockerham (1984) F-statistics, which takes into account only haplotype frequency, was also computed. The latter can be more powerful than Φ-statistics at detecting significant differentiation, when haplotype divergence does not contain any relevant information (Penant et al., 2013). Furthermore, since F-statistics does not depend on the degree of genetic distances among haplotypes, in terms of accumulation of mutations, it can detect population differentiation even after few generations of genetic drift. A non-parametric permutation method with 10000 replicates was used to test whether the derived indices were significantly different from zero.
A spatial analysis of molecular variance (SAMOVA) implemented in SAMOVA 2.0 (Dupanloup et al., 2002) was used to retrieve the likely number of hierarchical groups explaining most of the genetic structure. The method aims at finding the grouping scheme that maximizes variance among groups, without a prior population structure definition. The software was run with 100 random initial conditions for 10000 iterations, with number of tested groups, K = 2 to 7, and the most likely genetic structure was inferred either with or without spatial information on the sampling localities of Octopus vulgaris. The best partition obtained by SAMOVA was then used in the AMOVA framework as outlined above.
The relationships among haplotypes were analysed by a Median-Joining Network with default values (Bandelt et al., 1999) using the PopART v.1.7 software (Leigh & Bryant, 2015). Finally, the genetic structuring resulting from SAMOVA and the spatial distribution of haplotypes were superimposed on a map depicting the mean surface marine currents of the Mediterranean Sea over the years from 1998 to 2015. The map was Generated using the European Union (EU) Copernicus Marine Service Information (http://marine.copernicus.eu/): monthly mean currents over the period considered here were downloaded by Mediterranean Sea physical reanalysis (MEDREA 1987-2015) dataset (Simoncelli et al., 2014), whose model horizontal grid resolution is 1/16˚ (about 6–7 km). The resulting netCDF file was uploaded within the R 3.5.2 statistical environment (R Core Team, 2015) using the library ncdf4 1.17 (Pierce, 2019). The netcdf file was cropped in order to contain only months from July to December, which correspond to the reproductive and recruitment period of O. vulgaris in the Mediterranean (Garofalo et al., 2010). The new file was used to compute the resulting direction and mean current velocity over the period 1998–2015, which were plotted on a map using the R packages oceanmap 0.1.1 (Bauer, 2018), OceanView 1.0.5 (Soetaert, 2016) and RNetCDF 2.3.1 (Michna & Woods, 2017). Pie charts showing the genetic makeup of each population were superimposed on the map using the R package marmap 1.0.4 (Pante & Simon-Bouhet, 2013).
Results
Two data sets were obtained as the result of the different length of COI sequences that were available on public databases. Since the data set including the shortest sequences (500 bp) resulted in a reduction of both the number of haplotypes and the power of detecting genetic differentiation, only the alignment of 860 sequences that were 621 bp long was used for further analyses. Overall, 64 haplotypes were recorded, including sequences from eastern Atlantic (Online Resource 1: Table S2).
Based on the simulations, our data had sufficient power (> 95%) to detect weak population differentiation (FST ≥ 0.01). However, there was an excess of false positives, as population differentiation was wrongly detected in the 23% of pseudo-datasets (Online Resource 1: Fig. S1). Conversely, the test was too conservative when sample size was small (S = 5 and 10 individuals) as the rate of false positives was ten-fold less than its expectation (α-error = 0.004 and 0.007, respectively). Small sample size also decreased the power of detecting population differentiation when genetic divergence was weak. At S = 10 individuals, the power of detecting population differentiation reached 95% at FST ≥ 0.015. Halving of sample size (S = 5) increased up to five times the degree of genetic divergence (FST = 0.05) needed to get a power of detecting population differentiation > 95%. The detection rate of false positives depended on the number of populations and sample size (Online Resource 1: Fig. S1). In all cases, such rate did not exceed its expectation (α-error ≤ 5%) and tended to increase with the number of populations, whereas the effect of sampling size was less predictable. Therefore, only localities with S > 10 were retained for analyses that used summary statistics to estimate genetic divergence: 24 out of 34 populations were thus used within the AMOVA and SAMOVA frameworks. Such removal was enough to barely reduce the α-error rate to the expected level (α-error = 0.051).
Overall, Octopus vulgaris showed significant genetic structuring across the study area either based on Φ-statistics (overall ΦST = 0.065; p < 0.001) or F-statistics (overall FST = 0.073; p < 0.001). Regardless the metric used, AMOVA analyses highlighted significant genetic structuring between the Eastern and Western Mediterranean, either when the boundary was placed at the STS or shifted east off Sicily (Table 2 and Online Resource 1: Table S4). However, both metrics (ΦCT and FCT) were slightly higher when populations from Sicilian channel were grouped according to Bianchi’s (2007) biogeographical hypothesis “iii” (Fig. 1). The increase of the amount of variance explained by shifting eastward the boundary was greater when metric based only on haplotype frequency was used. Overall, such metric always showed a slightly greater genetic variance than metric accounting for mutations as well (Table 2 and Online Resource 1: Table S4).
Results based on SAMOVA, which searches for the grouping scheme that maximizes variance partitioning, highlighted an even more complex pattern (Table 2; Online resource 1: Table S2, Fig. S2). Populations along the Sicilian channel, DWK (Kerkennah Island Deep Waters, Tunisia) and TAJ (Tajoura, Lybian coast) were included in the Western Mediterranean group. Porto Santo Stefano (PSS, Tyrrhenian Sea) and all the remaining Eastern Mediterranean populations were ranked in the second and the third group, respectively. This grouping scheme showed an among-group variance component accounting for about 12% (Φ-statistics, ΦCT = 0.120; p < 0.001) and 12.5% (F-statistics, FCT = 0.125; p < 0.001) of total genetic variance.
The median-joining network highlighted a shallow level of divergence among haplotypes as most of them were separated by one mutational step and did not evidence any genetic break between the Western and eastern Mediterranean basins (Fig. 3). The four most common haplotypes (hereafter H1, H2, H10 and H17; Online resource 1: Table S3) were widespread across basins and localities, albeit their spatial distribution was uneven (Fig. 3). The frequency of H1 and H2, which were by far the most common haplotypes in the eastern Atlantic, decreased from west to east; however, the latter spread into the eastern Mediterranean reaching non-negligible frequencies, whereas the former was mainly restricted to the western basin (Fig. 4). Conversely, H10 and H17 showed more complex spatial patterns: H10 mainly spread across the Central Mediterranean, reaching the highest frequencies in a region encompassing the Sardinian and Sicilian channels, as well as the Tunisian coast north of the Gulf of Gabes (Fig. 4), H17 was the most geographically widespread haplotype in the Mediterranean, with its frequency decreasing from the Aegean Sea (eastern Mediterranean) to the Balearic Sea (western Mediterranean). The remaining haplotypes either occurred in a unique population (private haplotypes) or showed a restricted geographic distribution: for instance, the spatial distribution of H25, H27 and H31 (Fig. 3) encompassed an area delimited by the mainland coast of Italy in the East and the northern coast of Sardinia in the west (Fig. 4).
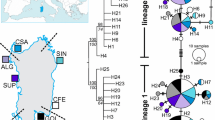
Median-Joining Network (MJN) showing the relationships among haplotypes of Octopus vulgaris. Circles represent haplotypes, whose size is proportional to their frequency. To ease the graphic visualisation, the length of edges connecting haplotypes is not proportional to the number of mutations, which are drawn as hatchmarks. Circles are coloured based on the frequency of haplotypes in each population. Shades of green are used for Atlantic populations; shades of white, yellow, orange and red for western Mediterranean populations; shades of magenta, purple and blue for eastern Mediterranean populations. Black dots represent median vectors, which may be interpreted as intermediate unsampled haplotypes
Haplotype frequencies superimposed on a map of monthly of marine currents. Pie charts show the genetic makeup of each population: slices correspond to the proportion of main haplotypes plus the cumulated frequencies of private and remaining haplotypes in each population. The map of marine currents was assembled from a dataset of monthly mean currents ranging from 1998 to 2015, generated using E.U. Copernicus Marine Service Information (http://marine.copernicus.eu/). Mean current speed and resulting direction have been computed based on the reproductive and recruitment period of Octopus vulgaris. The software R 4.0.2 (R Core Team, 2015) was used to subset the NETCDF file, compute mean current speed and direction and to plot the map. Monthly mean currents from July to December were used based on the period of reproduction and recruitment reported by Garofalo et al. (2010)
Discussion
The present study supports the hypothesis suggested by Villamor et al. (2018): the Sicilian channel and the neighbouring areas should be viewed as a transitional area, characterised by complex connectivity patterns between the Western and the Eastern Mediterranean. Consistent with this picture, genetic patterns observed in O. vulgaris across the STS roughly match those reported for Patella caerulea Linnaeus, 1758 in the same area (Villamor et al., 2018), as well as those of other species, regardless their different life-history traits (e.g., Maltagliati et al., 2010; Barbieri et al., 2014; Deli et al., 2018, 2019). Conversely, other species show a sharp genetic break across the Sicilian channel or east off it such as, for instance, the limpets P. rustica Linnaeus, 1758 and P. ulyssiponensis Gmelin, 1791 (Sá-Pinto et al., 2010, 2012). Such a stark contrast with P. caerulea (Villamor et al., 2014, 2018) is not counterintuitive as biogeographic boundaries can affect in different ways even closely related species with comparable biological traits (Galarza et al., 2009). Likely, dispersal and connectivity cannot be generalized for a given geographic area, as they depend on each species’ life history and biological traits (Fratini et al., 2016; Pascual et al., 2017), as well as the evolutionary history and the possible presence of endogenous genetic barriers among formerly isolated lineages as envisioned by the coupling hypothesis (Bierne et al., 2011). Therefore, focusing the attention on the whole transitional area instead of a unique genetic break may increase the ease of detecting common patterns across species as the need for geographically coincident steep changes of allele frequencies is released in the former case.
Our results do not support an abrupt shift of haplotype frequencies at a precise location as it could be expected for a sharp genetic break. Indeed, either the biogeographic boundary located at the STS or east off Sicily (hypotheses ii and iii in Bianchi, 2007) are supported by AMOVA analyses, although different metrics yielded slightly different results (Table 2; Online Resource 1: Table S4). The amount of genetic variation explained by partitioning populations according to the two barriers hypothesized by Bianchi (2007) that were considered in the present study is comparable when accounting for both haplotype frequency and genealogy (Online Resource 1: Table S4). Otherwise, genetic differentiation among groups resulting from the eastward shift of biogeographical boundary (hypothesis iii) is greater than that assessed based on the hypothesis ii (11.80 and 9.40%, respectively), when only haplotype frequency is considered (Table 2). Reasonably, the spatial genetic pattern observed across this area for populations of Octopus vulgaris is more complex than a simple shift of haplotype frequencies in east–west direction. In addition to a north-west south-east gradient of genetic variation, the present study highlights the partial isolation of the Gulf of Gabes from the rest of the Central Mediterranean. According to the best grouping scheme retrieved by SAMOVA, only CKI (Kerkennah Coastal Waters) and ZAR (Zarzis) on the Tunisian coast rank in the same group with ANC (Ancona) and CRE (Heraklion, Crete) in the Adriatic and Aegean Sea, respectively. This result also fits with the low levels of genetic diversity reported on CKI and ZAR by Fadhlaoui-Zid et al. (2012). Conversely, all the other Eastern Mediterranean populations clustered with the Western Mediterranean populations. Such apparently chaotic genetic pattern may reflect the influence of the complex hydrographical regime that characterizes the Sicilian channel and neighboring areas. The pattern of marine currents in the Sicilian channel is seasonally variable and influenced by the presence of eddies and gyres (Poulain & Zambianchi, 2007; Ciappa, 2009 and references therein). Atlantic water, advected toward STS by the Algerian current, split into two branches, one of which flows south-eastward along Sicilian coast (the Atlantic Ionian Stream, AIS) and the other southward over the Tunisian shelf at 13°E, about 160 km east off the Tunisian coast (the Atlantic Tunisian Current, ATC) (Poulain & Zambianchi, 2007; Ciappa, 2009). Whereas the former is stronger in summer and weaker in winter, the opposite trend has been observed for the ATC, which is almost undetectable in summer. The winter pattern of this current is still debated: whereas drifter trajectories indicate intrusion of AW in the Gulf of Gabes along the mainstream of ATC (Poulain & Zambianchi, 2007), patterns of chlorophyll concentration suggest that ATC is deviated eastward at about 35°N by an anticyclonic gyre (the Sydra gyre) and reach Lybian coast flowing along its border (Ciappa, 2009). Atlantic water can still reach Lybian coast in summer by a current that flows southward from the southeastern tip of Sicily and then merge with the eastern arm of the Sydra gyre (Ciappa, 2009).
The spatial distribution of haplotype frequencies fits quite well with the pattern of marine currents, with the most widespread haplotypes showing a gradual rather than a steep change of their frequencies in a northwest-southeast direction (Fig. 4). Not surprisingly, DWK and TAJ (Kerkennah deep waters and Tajoura, respectively) show a genetic makeup more similar to populations on the northern side of the Sicilian channel than to the other Tunisian populations (Kerkennah coastal waters and Zarzis). DWK is located close to the mainstream of the ATC, whereas TAJ is close to a trait of coast that can be reached either by the ATC (Poulain & Zambianchi, 2007) or by the southward current flowing by the southeastern tip of Sicily (Ciappa, 2009). Thus, spatial genetic patterns of O. vulgaris observed in the northern sector of the Sicilian Channel, at Kerkennah deep waters and Tajoura might reflect a stronger advection of paralarvae from the Western Mediterranean related to the southern Tunisian coast and the Gulf of Gabes. The isolation of the latter area as well as that of Adriatic and Aegean seas related to the Mediterranean Sea, might explain why scattered populations as Kerkennah coastal waters, Zarzis, Ancona and Heraklion show similar genetic makeups. If the frequency and geographic distribution of the main haplotypes reflects the species’ demographic history in the Mediterranean, we may speculate that this populations still retain the genetic signature of older historical events.
The comparison of measures of genetic differentiation based on Φ- and F-statistics also supports the role of marine currents in shaping the spatial genetic patterns of O. vulgaris. The slightly higher value of the latter measure indicates that accumulation of mutations, which is accounted for the estimation of Φ-statistics parameters and reflects the relative contribution of past evolutionary history, has a little effect on genetic structuring (Fadhlaoui-Zid et al., 2012). Conversely, F-statistics is more powerful at detecting population differentiation after few generations of genetic drift, as it relies only on haplotype frequencies to estimate genetic divergence (Penant et al., 2013).
A last interesting remark deals with the case of PSS population, which does not group with any other Mediterranean population according to SAMOVA (Online resource 1: Fig. S2). This population displayed a peculiar genetic makeup (Fig. 3): in contrast to other Western Mediterranean populations, the most common haplotypes show a relatively low frequency. In fact, PSS is characterized by the occurrence of high-frequency haplotypes that are rare elsewhere, apart from LEG, NAP and PTO (haplotypes 25, 27 and 31, Fig. 3). Based on the life history traits of O. vulgaris, we cannot rule out that this pattern reflect temporal rather than spatial genetic variation, even though neither the present study (Online resource 1: Table S5) nor Melis et al. (2018) evidenced the occurrence of temporal genetic subdivision. Even though it is not geographically widespread in the present study, it may still occur at local scales as in other marine invertebrates in which recruitment and growth are affected by environmental factors. For instance, spatio-temporal genetic variation was found by Calderón et al. (2012) during a five-years genetic monitoring of the sea urchin, Paracentrotus lividus (Lamarck, 1816). Notwithstanding an overall genetic homogeneity across the timeframe of the study, the genetic composition of some studied populations changed markedly in a particular year, with the appearance of many private haplotypes. Such a change in the genetic makeup was related to the unusual circulation pattern in that year, which likely favored an abnormal migration of recruits from genetically heterogeneous source populations (Calderón et al., 2012). The same scenario cannot be discarded for O. vulgaris, as marine circulation in the Tyrrhenian Sea is characterized by a high degree of seasonal and inter-annual variability, with the formation of mesoscale eddies and gyres (Iacono et al., 2013). Such oceanographical features coupled with the sea surface temperature variation may influence recruitment and growth in O. vulgaris. For instance, the number and extent of spawning and nursery grounds show inter-annual variability associated with environmental conditions in the Central Mediterranean (Garofalo et al., 2010). Genetic monitoring over different years is needed to disentangle these two competing, but not mutually exclusive scenarios in O.vulgaris.
Conclusions
The present study does not support the presence of a sharp genetic break across the Siculo-Tunisian strait in Mediterranean populations of O. vulgaris. The complex genetic patterns observed in this species are another clue in on the hypothesis that the Central Mediterranean might be a transition zone. Nevertheless, as many species show a sharp genetic break across the STS or east off this channel, more studies realising a dense geographic sampling around this area are needed to assess if and how widespread this genetic pattern is in marine organisms with different life-histories.
Although investigating the drivers that shape spatial genetic structure is beyond the scope of the present study, our results indicate that spatial genetic structure in O. vulgaris is influenced by contemporary factors as oceanographic features. However, marine currents might not be the only or the main factor contributing to shape the genetic structure observed in the present study. In addition to exogenous (due to oceanographic and/or environmental factors), endogenous genetic barriers (due to historical isolation followed by secondary contact) are important factors underpinning complex genetic patterns, as outlined by the coupling hypothesis (Bierne et al., 2011). The use of nuclear markers and joint biophysical-genetic models, which combine larval dispersal and spatially explicit genetic models, may shed light into the spatial distribution of genetic variation expected over ecologically relevant timescales. These models may thus help better define the relative contributions of historical and contemporary factors on species’ genetic structuring.
Moreover, the limited connectivity between the Gulf of Gabes and the other regions may have important implications for common octopus fisheries, suggesting that populations located in this area might be regarded as an independent stock, relying mostly on self-recruitment. The results outlined above further stress the need of designing an exhaustive geographical sampling to unravel phylogeographical patterns in areas that are characterized by a complex oceanography as the Central Mediterranean. As evidenced by Fratini et al. (2016), limited sampling of both populations and individuals per population might hamper the detection of genetic structuring in marine invertebrates with larval dispersal stages.
Data availability
The datasets generated and/or analysed during the current study are available in the FIGSHARE repository, https://doi.org/10.6084/m9.figshare.12957380.
References
Amor, M. D., M. D. Norman, H. E. Cameron & J. M. Strugnell, 2014. Allopatric speciation within a cryptic species complex of Australasian octopuses. PLoS ONE 9:
Bandelt, H. J., P. Forster & A. Röhl, 1999. Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution 16: 37–48.
Barbieri, M., F. Maltagliati, M. I. Roldán & A. Castelli, 2014. Molecular contribution to stock identification in the small-spotted catshark, Scyliorhinus canicula (Chondrichthyes, Scyliorhinidae). Fisheries Research 154: 11–16.
Bas, C., 2009. The Mediterranean: a synoptic overview. Contributions in Marine Science 5: 25–39.
Bauer, R.K. 2018. Oceanmap: a plotting toolbox for 2D oceanographic data. https://CRAN.R-project.org/package=oceanmap. Accessed 9 Jul 2019
Belcari, P., D. Cuccu, M. Gonzalez, A. Srairi & P. Vidoris, 2002. Distribution and abundance of Octopus vulgaris Cuvier, 1797 (Cephalopoda: Octopoda) in the Mediterranean Sea. Scientia Marina 62: 157–166.
Bianchi, C. N., 2007. Biodiversity issues for the forthcoming tropical Mediterranean Sea. Hydrobiologia 580: 7–21.
Bierne, N., J. Welch, E. Loire, F. Bonhomme & P. David, 2011. The coupling hypothesis: why genome scans may fail to map local adaptation genes. Molecular Ecology 20: 2044–2072.
Boissin, E., S. Stöhr & A. Chenuil, 2011. Did vicariance and adaptation drive cryptic speciation and evolution of brooding in Ophioderma longicauda (Echinodermata: ophiuroidea), a common Atlanto-Mediterranean ophiuroid? Molecular Ecology 20: 4737–4755.
Borrero-Pérez, G. H., M. Gonzàlez-Wangüemert, C. Marcos & A. Pérez-Ruzafa, 2011. Phylogeography of the Atlanto-Mediterranean sea cucumber Holothuria (Holothuria) mammata: the combined effects of historical processes and current oceanographical pattern. Molecular Ecology 20: 1964–1975.
Borsa, P., M. Naciri, L. Bahri, L. Chikhi & F. J. Garcia de Leon, 1997. Zoogéographie infra-spécifique de la mer Méditerranée: analyse des données génétiques populationnelles sur seize espèces atlanto-méditerranéennes (poissons et invertébrés). Vie et milieu 47: 295–305.
Cabranes, C., P. Fernandez-Rueda & J. L. Martinez, 2008. Genetic structure of Octopus vulgaris around the Iberian Peninsula and Canary Islands as indicated by microsatellite DNA variation. ICES Journal of Marine Science 65: 12–16.
Calderón, I., L. Pita, S. Brusciotti, C. Palacín & X. Turon, 2012. Time and space: genetic structure of the cohorts of the common sea urchin Paracentrotus lividus in Western Mediterranean. Marine Biology 159: 187–197.
Casu, M., F. Maltagliati, M. Meloni, D. Casu, P. Cossu, G. Binelli, M. Curini-Galletti & A. Castelli, 2002. Genetic structure of Octopus vulgaris (Mollusca, Cephalopoda) from the Mediterranean Sea as revealed by a microsatellite locus. Italian Journal of Zoology 71: 473–486.
Ciappa, A. C., 2009. Surface circulation patterns in the Sicily Channel and Ionian Sea as revealed by MODIS chlorophyll images from 2003 to 2007. Continental Shelf Research 29: 2099–2109.
Coll, M., C. Piroddi, C. Albouy, F. B. R. Lasram, W. W. L. Cheung, V. Christensen, V. S. Karpouzi, F. Guilhaumon, D. Mouillot, M. Paleczny, M. L. Palomares, J. Steenbeck, P. Trujillo, R. Watsin & D. Pauly, 2012. The Mediterranean Sea under siege: spatial overlap between marine biodiversity, cumulative threats and marine reserves. Global Ecology and Biogeography 21: 465–480.
Cordero, D., J. B. Peña & C. Saavedra, 2014. Phylogeographic analysis of introns and mitochondrial DNA in the clam Ruditapes decussatus uncovers the effects of Pleistocene glaciations and endogenous barriers to gene flow. Molecular Phylogenetics and Evolution 71: 274–287.
Cossu, P., G. L. Dedola, F. Scarpa, D. Sanna, T. Lai, F. Maltagliati, M. Curini-Galletti & M. Casu, 2015. Patterns of spatial genetic variation in Patella ulyssiponensis: insights from the western Mediterranean marine ecoregion. Hydrobiologia 755: 39–55.
De Luca, D., G. Catanese, G. Procaccini & G. Fiorito, 2014. An integration of historical records and genetic data to the assessment of global distribution and population structure in Octopus vulgaris. Frontiers in Ecology and Evolution 2: 55.
De Luca, D., G. Catanese, G. Procaccini & G. Fiorito, 2016. Octopus vulgaris (Cuvier, 1797) in the Mediterranean Sea: genetic diversity and population structure. PLoS ONE 11:
Deli, T., E. Kalkan, S. Ü. Karhan, S. Uzunova, A. Keikhosravi, R. Bilgin & C. D. Schubart, 2018. Parapatric genetic divergence among deep evolutionary lineages in the Mediterranean green crab, Carcinus aestuarii (Brachyura, Portunoidea, Carcinidae), accounts for a sharp phylogeographic break in the Eastern Mediterranean. BMC Evolutionary Biology 18: 53.
Deli, T., C. Kiel & D. Schubart, 2019. Phylogeographic and evolutionary history analyses of the warty crab Eriphia verrucosa (Decapoda, Brachyura, Eriphiidae) unveil genetic imprints of a late Pleistocene vicariant event across the Gibraltar Strait, erased by postglacial expansion and admixture among refugial lineages. BMC Evolutionary Biology 19: 105.
Dupanloup, I., S. Schneider & L. Excoffier, 2002. A simulated approach to define the genetic structure of populations. Molecular Ecology 11: 2571–2581.
El Ayari, T., N. T. El Menif, B. Hamer, A. E. Cahill & N. Bierne, 2019. The hidden side of a major marine biogeographic boundary: a wide mosaic hybrid zone at the Atlantic-Mediterranean divide reveals the complex interaction between natural and genetic barriers in mussels. Heredity 122: 770–784.
Excoffier, L. & H. E. L. Lischer, 2010. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Research 10: 564–567.
Fadhlaoui-Zid, K., L. Knittweis, D. Aurelle, C. Nafkha, S. Ezzeddine, F. Fiorentino, H. Ghmati, L. Ceriola, O. Jarboui & F. Maltagliati, 2012. Genetic structure of Octopus vulgaris (Cephalopoda, Octopodidae) in the central Mediterranean Sea inferred from the mitochondrial COIII gene. Comptes Rendus Biologies 335: 625–636.
Fadhlaoui-Zid, K., D. Sanna, D. Aurelle, S. Ezzeddine, A. Castelli, M. Casu & F. Maltagliati, 2013. Phylogeographical analysis of Octopus vulgaris (Mollusca, Cephalopoda) in the Mediterranean: insights from the COI mitochondrial gene. Rapports et procès verbaux des réunions - Commission Internationale pour l’Exploration Scientifique de la Mer Méditerranée 40: 712.
Folmer, O., M. Black, W. Hoeh, R. Lutz & R. Vrijenhoek, 1994. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology 3: 294–299.
Fratini, S., L. Ragionieri, T. Deli, A. Harrer, I. A. M. Marino, S. Cannicci, L. Zane & C. D. Schubart, 2016. Unravelling population genetic structure with mitochondrial DNA in a notional panmictic coastal crab species: sample size makes the difference. BMC Evolutionary Biology 16: 150.
Fruciano, C., R. Hanel, P. V. Debes, C. Tigano & V. Ferrito, 2011. Atlantic-Mediterranean and within-Mediterranean molecular variation in Coris julis (L. 1758) (Teleostei, Labridae). Marine Biology 158: 1271–1286.
Galarza, J. A., G. F. Turner, E. Macpherson & C. Rico, 2009. Patterns of genetic differentiation between two co-occurring demersal species: the red mullet (Mullus barbatus) and the striped red mullet (Mullus surmuletus). Canadian Journal of Fisheries and Aquatic Sciences 66: 1478–1490.
Garofalo, G., L. Ceriola, M. Gristina, F. Fiorentino & R. Pace, 2010. Nurseries, spawning grounds and recruitment of Octopus vulgaris in the Strait of Sicily, central Mediterranean Sea. ICES Journal of Marine Science 67: 1363–1371.
Hall, T. A., 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41: 95–98.
Hernández-López, J. L., J. J. Castro-Hernández & V. Hernández-García, 2001. Age determined from the daily deposition of concentric rings on common octopus (Octopus vulgaris) beaks. Fishery Bulletin 99: 679–694.
Katsanevakis, S. & G. Verriopoulos, 2006. Modelling the effect of temperature on hatching and settlement patterns of meroplanktonic organisms: the case of octopus. Scientia Marina 70: 699–708.
Iacono, R., E. Napolitano, S. Marullo, V. Artale & A. Vetrano, 2013. Seasonal variability of the tyrrhenian sea surface geostrophic circulation as assessed by altimeter data. Journal of Physical Oceanography 43: 1710–1732.
Kumar, S., G. Stecher & K. Tamura, 2016. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution 33: 1870–1874.
Larsson, L. C., J. Charlier, L. Laikre & N. Ryman, 2009. Statistical power for detecting genetic divergence—organelle versus nuclear markers. Conservation Genetics 10: 1255–1264.
Leigh, J. W. & D. Bryant, 2015. PopART: full-feature software for haplotype network construction. Methods in Ecology and Evolution 6: 1110–1116.
Lejeusne, C., P. Chevaldonné, C. Pergent-Martini, C. F. Boudouresque & T. Pérez, 2010. Climate change effects on a miniature ocean: the highly diverse, highly impacted Mediterranean Sea. Trends in Ecology and Evolution 25: 250–260.
Lleonart, J. & F. Maynou, 2003. Fish stock assessments in the Mediterranean: state of the art. Scientia Marina 67: 37–49.
Maltagliati, F., P. Belcari, D. Casu, M. Casu, P. Sartor, G. Vargiu & A. Castelli, 2002. Allozyme genetic variability and gene flow in Octopus vulgaris (Cephalopoda, Octopodidae) from the Mediterranean Sea. Bulletin of Marine Science 71: 473–486.
Maltagliati, F., G. Di Giuseppe, M. Barbieri, A. Castelli & F. Dini, 2010. Phylogeography and genetic structure of the edible sea urchin Paracentrotus lividus (Echinodermata: echinoidea) inferred from the mitochondrial cytochrome b gene. Biological Journal of the Linnean Society 100: 910–923.
Mather, J. A., 1994. ‘Home’choice and modification by juvenile Octopus vulgaris (Mollusca: Cephalopoda): specialized intelligence and tool use? Journal of Zoology 233: 359–368.
Melis, R., L. Vacca, D. Cuccu, M. Mereu, A. Cau, M. C. Follesa & R. Cannas, 2018. Genetic population structure and phylogeny of the common octopus Octopus vulgaris Cuvier, 1797 in the western Mediterranean Sea through nuclear and mitochondrial markers. Hydrobiologia 807: 277–296.
Michna, P. & M. Woods 2017. RNetCDF: Interface to NetCDF datasets. R package version 1.9-1. https://CRAN.R-project.org/package=RNetCDF. Accessed 9 Jul 2019
Mokhtar-Jamaï, K., M. Pascual, J.-B. Ledoux, R. Coma, J.-P. Féral, J. Garrabou & D. Aurelle, 2011. From global to local genetic structuring in the red gorgonian Paramuricea clavata: the interplay between oceanographic conditions and limited larval dispersal. Molecular Ecology 20: 3291–3305.
O’Connor, M., J. F. Bruno, S. D. Gaines, B. S. Halpern, S. E. Lester, B. P. Kinlan & J. M. Weiss, 2007. Temperature control of larval dispersal and the implications for marine ecology, evolution, and conservation. Proceedings of the National Academy of Sciences of the United States of America 104: 1266–1271.
Pannacciulli, F. G., F. Maltagliati, C. De Guttry & Y. Achituv, 2017. Phylogeography on the rocks: the contribution of current and historical factors in shaping the genetic structure of Chthamalus montagui (Crustacea, Cirripedia). PLoS ONE 12:
Pante, E. & B. Simon-Bouhet, 2013. marmap: a package for importing, plotting and analyzing bathymetric and topographic data in R. PloS ONE 8:
Pascual, M., B. Rives, C. Schunter & E. Macpherson, 2017. Impact of life history traits on gene flow: a multispecies systematic review across oceanographic barriers in the Mediterranean Sea. PLoS ONE 12:
Patarnello, T., F. A. M. J. Volckaert & R. Castilho, 2007. Pillars of hercules: is the Atlantic-Mediterranean transition a phylogeographic break? Molecular Ecology 16: 4426–4444.
Penant, G., D. Aurelle, J.-P. Feral & A. Chenuil, 2013. Planktonic larvae do not ensure gene flow in the edible sea urchin Paracentrotus lividus. Marine Ecology Progress Series 480: 155–170.
Pierce, D. 2019. ncdf4: Interface to Unidata netCDF (Version 4 or Earlier) Format Data Files. R package version 1.16.1. https://CRAN.R-project.org/package=ncdf4. Accessed 9 Jul 2019
Poulain, P. M. & E. Zambianchi, 2007. Surface circulation in the central Mediterranean Sea as deduced from Lagrangian drifters in the 1990s. Continental Shelf Research 27: 981–1001.
Ragionieri, L. & C. D. Schubart, 2013. Population genetics, gene flow, and biogeographical boundaries of Carcinus aestuarii (Crustacea: Brachyura: Carcinidae) along the European Mediterranean coast. Biological Journal of the Linnean Society 109: 771–790.
R Core Team 2015. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/. Accessed 9 Jul 2019
Roura, Á., M. Amor, Á. F. González, Á. Guerra, E. D. Barton & J. M. Strugnell, 2019. Oceanographic processes shape genetic signatures of planktonic cephalopod paralarvae in two upwelling regions. Progress in Oceanography 170: 11–27.
Ryman, N. & S. Palm, 2006. POWSIM: a computer program for assessing statistical power when testing for genetic differentiation. Molecular Ecology Notes 6: 600–602.
Sales, J. B. D. L., P. S. D. Rego, A. W. S. Hilsdorf, A. A. Moreira, M. Haimovici, A. R. Tomás, B. B. Batista, R. A. Marinho, U. Markaida, H. Schneider & I. Sampaio, 2013. Phylogeographical features of Octopus vulgaris and Octopus insularis in the Southeastern Atlantic based on the analysis of mitochondrial markers. Journal of Shellfish Research 32: 325–339.
Sá-Pinto, A., S. J. E. Baird, C. Pinho, P. Alexandrino & M. Branco, 2010. A three-way contact zone between forms of Patella rustica (Mollusca: Patellidae) in the central Mediterranean Sea. Biological Journal of the Linnean Society 100: 154–169.
Sá-Pinto, A., M. Branco, P. Alexandrino, M. C. Fontaine & S. J. E. Baird, 2012. Barriers to gene flow in the marine environment: insights from two common intertidal limpet species of the Atlantic and Mediterranean. PLoS ONE 7:
Sanna, D., P. Cossu, G. L. Dedola, F. Scarpa, F. Maltagliati, A. Castelli, P. Franzoi, T. Lai, B. Cristo, M. Curini-Galletti, P. Francalacci & M. Casu, 2013. Mitochondrial DNA Reveals Genetic Structuring of Pinna nobilis across the Mediterranean Sea. PLoS ONE 8:
Sanna, D., G. L. Dedola, F. Scarpa, T. Lai, P. Cossu, H. Ben Halaya, M. Curini-Galletti, P. Francalacci & M. Casu, 2014. New mitochondrial and nuclear markers for the Mediterranean marine bivalve Pinna nobilis. Mediterranean Marine Science 15: 416–422.
Schunter, C., J. Carreras-Carbonell, E. Macpherson, J. Tintoré, E. Vidal-Vijande, A. Pascual, P. Guidetti & M. Pascual, 2011a. Matching genetics with oceanography: directional gene flow in a Mediterranean fish species. Molecular Ecology 20: 5167–5181.
Schunter, C., J. Carreras-Carbonell, S. Planes, E. Sala, E. Ballesteros, M. Zabala, J. G. Harmelin, M. Harmelin-Vivien, E. Macpherson & M. Pascual, 2011b. Genetic connectivity patterns in an endangered species: the dusky grouper (Epinephelus marginatus). Journal of Experimental Marine Biology and Ecology 401: 126–133.
Simoncelli, S., C. Fratianni, N. Pinardi, A. Grandi, M. Drudi, P. Oddo & S. Dobricic 2014. Mediterranean Sea physical reanalysis (MEDREA 1987-2015) [Data set]. Copernicus Monitoring Environment Marine Service (CMEMS). https://doi.org/10.25423/MEDSEA_REANALYSIS_PHYS_006_004
Smale, M. J. & P. R. Buchan, 1981. Biology of Octopus vulgaris off the East coast of South Africa. Marine Biology 65: 1–12.
Soetaert, K. 2016. OceanView: visualisation of oceanographic data and model output. R package version 1.0.4. https://CRAN.R-project.org/package=OceanView. Accessed 9 Jul 2019
Souissi, A., F. Bonhomme, M. Manchado, L. Bahri-Sfar & P. A. Gagnaire, 2018. Genomic and geographic footprints of differential introgression between two divergent fish species (Solea spp.). Heredity 121: 579–593.
Spalding, M. D., H. E. Fox, G. R. Allen, N. Davidson, Z. A. Ferdaña, M. Finlayson, B. S. Halpern, M. A. Jorge, A. Lombana, S. A. Lourie, K. D. Martin, E. McManus, J. Molnar, C. A. Recchia & J. Robertson, 2007. Marine ecoregions of the World: a bioregionalization of coastal and shelf areas. Bioscience 57: 573–583.
Tamura, K., 1992. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G + C content biases. Molecular Biology and Evolution 9: 678–687.
Tarnowska, K., A. Chenuil, R. Nikula, J.-P. Féral & M. Wolowicz, 2010. Complex genetic population structure of the bivalve Cerastoderma glaucum in a highly fragmented lagoon habitat. Marine Ecology Progress Series 406: 173–184.
Thompson, J. D., D. G. Higgins & T. J. Gibson, 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research 22: 4673–4680.
Villamor, A., F. Costantini & M. Abbiati, 2014. Genetic structuring across marine biogeographic boundaries in rocky shore invertebrates. PLoS ONE 9:
Villamor, A., F. Costantini & M. Abbiati, 2018. Multilocus phylogeography of Patella caerulea (Linnaeus, 1758) reveals contrasting connectivity patterns across the Eastern-Western Mediterranean transition. Journal of Biogeography 45: 1301–1312.
Villanueva, R., C. Nozais & S. V. Boletsky, 1997. Swimming behavior and food searching in planktonic Octopus vulgaris Cuvier from hatching to settlement. Journal of Experimental Marine Biology and Ecology 208: 169–184.
Warnke, K., R. Soller, D. Blohm & U. Saint-Paul, 2004. A new look at geographic and phylogenetic relationships within the species group surrounding Octopus vulgaris (Mollusca, Cephalopoda): indications of very wide distribution from mitochondrial DNA sequences. Journal of Zoological Systematics and Evolutionary Research 42: 306–312.
Weir, B. S. & C. C. Cockerham, 1984. estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Acknowledgements
This work was supported by the Tunisian Ministry of Higher Education and Scientific Research.
Funding
Open access funding provided by Università degli Studi di Sassari within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
All authors, who agreed to be listed, read and approved the submitted version of the manuscript. KFZ and FM designed the study and all authors contributed to conceive the work, DS, FS, TL and MC acquired the data, PC carried out data analysis and interpretation and drafted the work, which was critically revised by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest
Ethical approval
The study did not involve endangered or protected species. New DNA sequences were obtained from genomic DNA stored at – 20 °C that had been isolated from tissue samples collected in previous studies (See materials and methods for further details). Therefore, no specific permissions were required for locations and activities involved in this study.
Additional information
Handling editor: Cécile Fauvelot
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fadhlaoui-Zid, K., Cossu, P., Sanna, D. et al. Spatial genetic patterns of Octopus vulgaris Mediterranean populations support the hypothesis of a transitional zone across the Siculo-Tunisian Strait. Hydrobiologia 848, 4225–4240 (2021). https://doi.org/10.1007/s10750-021-04634-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-021-04634-6