Abstract
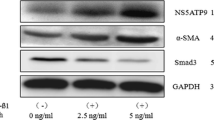
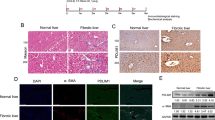
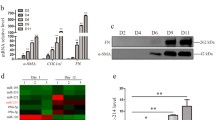
Liver fibrosis is a global public health problem, and the activation of hepatic stellate cells (HSCs) is the main driving force for liver fibrosis. However, the activation mechanism of HSCs is still not fully understood. In this study, we screened out 854 differentially expressed genes [Log2 fold change absolute: log2 FC(abs) ≥ 1] in activated LX-2 cells. Subsequently, we performed functional analyses of these differentially expressed genes. Gene Ontology enrichment analysis showed that the target genes were mainly enriched in processes such as positive regulation of cell migration involved in sprouting angiogenesis, negative regulation of keratinocyte proliferation, and nuclear inclusion bodies. Kyoto Encyclopedia of Gene and Genome signaling pathway enrichment analysis revealed that dysregulated genes were involved in signaling pathways such as pantothenate and coenzyme A biosynthesis and riboflavin metabolism. The microarray results were validated by reverse transcription-quantitative polymerase chain reaction, which indicated that the microarray results were reliable and that the tripartite motif containing 15 (TRIM15) had the highest absolute value of Log2FC. Additionally, the effect of TRIM15 on the proliferation, migration, and activation of LX-2 cells was assessed using overexpression plasmids and siRNA transfections. TRIM15 promoted the proliferation and migration of LX-2 cells and positively regulated the expression of α-smooth muscle actin and type I collagen. Collectively, the data revealed the gene expression profiles of quiescent and activated LX-2 cells and the involvement of TRIM15 in the activation of LX-2 cells. Hereby, TRIM15 could be a novel target of the HSC activation mechanism.






Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request for non‑commercial purposes, without breaching participant confidentiality.
References
Bai F, Huang Q, Nie J, Lu S, Lu C, Zhu X, Wang Y, Zhuo L, Lu Z, Lin X (2017) Trolline ameliorates liver fibrosis by inhibiting the nf-kappab pathway, promoting HSC apoptosis and suppressing autophagy. Cell Physiol Biochem 44:436–446. https://doi.org/10.1159/000485009
Bataller R, Brenner DA (2005) Liver fibrosis. J Clin Invest 115:209–218. https://doi.org/10.1172/JCI24282
Bataller R, North KE, Brenner DA (2003) Genetic polymorphisms and the progression of liver fibrosis: a critical appraisal. Hepatology 37:493–503. https://doi.org/10.1053/jhep.2003.50127
Chang Y, Li H (2020) Hepatic antifibrotic pharmacotherapy: are we approaching success? J Clin Transl Hepatol 8:222–229. https://doi.org/10.14218/JCTH.2020.00026
Chen W, Zhang Z, Yao Z, Wang L, Zhang F, Shao J, Chen A, Zheng S (2018) Activation of autophagy is required for Oroxylin A to alleviate carbon tetrachloride-induced liver fibrosis and hepatic stellate cell activation. Int Immunopharmacol 56:148–155. https://doi.org/10.1016/j.intimp.2018.01.029
Chen Y, Chen X, Ji YR, Zhu S, Bu FT, Du XS, Meng XM, Huang C, Li J (2020) PLK1 regulates hepatic stellate cell activation and liver fibrosis through Wnt/beta-catenin signalling pathway. J Cell Mol Med 24:7405–7416. https://doi.org/10.1111/jcmm.15356
Crawford LJ, Johnston CK, Irvine AE (2018) TRIM proteins in blood cancers. J Cell Commun Signal 12:21–29. https://doi.org/10.1007/s12079-017-0423-5
Dooley S, ten Dijke P (2012) TGF-beta in progression of liver disease. Cell Tissue Res 347:245–256. https://doi.org/10.1007/s00441-011-1246-y
Du G, Wang J, Zhang T, Ding Q, Jia X, Zhao X, Dong J, Yang X, Lu S, Zhang C, Liu Z, Zeng Z, Safadi R, Qi R, Zhao X, Hong Z, Lu Y (2020) Targeting Src family kinase member Fyn by Saracatinib attenuated liver fibrosis in vitro and in vivo. Cell Death Dis 11:118. https://doi.org/10.1038/s41419-020-2229-2
Ellis EL, Mann DA (2012) Clinical evidence for the regression of liver fibrosis. J Hepatol 56:1171–1180. https://doi.org/10.1016/j.jhep.2011.09.024
Fang J, Yao X, Hou M, Duan M, Xing L, Huang J, Wang Y, Zhu B, Chen Q, Wang H (2020) ApoL1 induces kidney inflammation through RIG-I/NF-kappaB activation. Biochem Biophys Res Commun 527:466–473. https://doi.org/10.1016/j.bbrc.2020.04.054
Hammond SM, Aartsma-Rus A, Alves S, Borgos SE, Buijsen RAM, Collin RWJ, Covello G, Denti MA, Desviat LR, Echevarria L, Foged C, Gaina G, Garanto A, Goyenvalle AT, Guzowska M, Holodnuka I, Jones DR, Krause S, Lehto T, Montolio M, Van Roon-Mom W, Arechavala-Gomeza V (2021) Delivery of oligonucleotide-based therapeutics: challenges and opportunities. EMBO Mol Med 13:e13243. https://doi.org/10.15252/emmm.202013243
Higashi T, Friedman SL, Hoshida Y (2017) Hepatic stellate cells as key target in liver fibrosis. Adv Drug Deliv Rev 121:27–42. https://doi.org/10.1016/j.addr.2017.05.007
Hong Y, Li S, Wang J, Li Y (2018) In vitro inhibition of hepatic stellate cell activation by the autophagy-related lipid droplet protein ATG2A. Sci Rep 8:9232. https://doi.org/10.1038/s41598-018-27686-6
Hu GX, Wan ZY, Shao JL, Zhang Y, Zhang LL, Gong ZJ (2012) Effects of hydroxycamptothecin on TGFb1, a-SMA and collagen I expression in rat hepatic satellite cells. Zhonghua Gan Zang Bing Za Zhi 20:453–457. https://doi.org/10.3760/cma.j.issn.1007-3418.2012.06.015
Jiang MD, Zheng SM, Xu H, Zeng WZ, Zhang Y, Sun HP, Wang YX, Qin JP, Wu XL (2008) An experimental study of extracellular signal-regulated kinase and its interventional treatments in hepatic fibrosis. Hepatobiliary Pancreat Dis Int 7:51–57
Jiang Y, Zhu F, Wu GS, Wang KA, Wang C, Yu Q, Zhu BH, Sun Y, Xia ZF (2020) Microarray and bioinformatics analysis of circular RNAs expression profile in traumatic lung injury. Exp Ther Med 20:227–234. https://doi.org/10.3892/etm.2020.8686
Kumar P, Smith T, Raeman R, Chopyk DM, Brink H, Liu Y, Sulchek T, Anania FA (2018) Periostin promotes liver fibrogenesis by activating lysyl oxidase in hepatic stellate cells. J Biol Chem 293:12781–12792. https://doi.org/10.1074/jbc.RA117.001601
Lee OH, Lee J, Lee KH, Woo YM, Kang JH, Yoon HG, Bae SK, Songyang Z, Oh SH, Choi Y (2015) Role of the focal adhesion protein TRIM15 in colon cancer development. Biochim Biophys Acta 1853:409–421. https://doi.org/10.1016/j.bbamcr.2014.11.007
Li X, Shang Y, Yao W, Li Y, Tang N, An J, Wei Y (2020) Comparison of transcriptomics changes induced by TCS and MTCS exposure in human hepatoma HepG2 cells. ACS Omega 5:10715–10724. https://doi.org/10.1021/acsomega.0c00075
Liczner C, Duke K, Juneau G, Egli M, Wilds CJ (2021) Beyond ribose and phosphate: Selected nucleic acid modifications for structure-function investigations and therapeutic applications. Beilstein J Org Chem 17:908–931. https://doi.org/10.3762/bjoc.17.76
Liu X, Rosenthal SB, Meshgin N, Baglieri J, Musallam SG, Diggle K, Lam K, Wu R, Pan SQ, Chen Y, Dorko K, Presnell S, Benner C, Hosseini M, Tsukamoto H, Brenner D, Kisseleva T (2020) Primary alcohol-activated human and mouse hepatic stellate cells share similarities in gene-expression profiles. Hepatol Commun 4:606–626. https://doi.org/10.1002/hep4.1483
Uchil PD, Pawliczek T, Reynolds TD, Ding S, Hinz A, Munro JB, Huang F, Floyd RW, Yang H, Hamilton WL, Bewersdorf J, Xiong Y, Calderwood DA, Mothes W (2014) TRIM15 is a focal adhesion protein that regulates focal adhesion disassembly. J Cell Sci 127:3928–3942. https://doi.org/10.1242/jcs.143537
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, Kinzler KW (2013) Cancer genome landscapes. Science 339:1546–1558. https://doi.org/10.1126/science.1235122
Wang Q, Dai X, Yang W, Wang H, Zhao H, Yang F, Yang Y, Li J, Lv X (2015) Caffeine protects against alcohol-induced liver fibrosis by dampening the cAMP/PKA/CREB pathway in rat hepatic stellate cells. Int Immunopharmacol 25:340–352. https://doi.org/10.1016/j.intimp.2015.02.012
Wang Q, Wei S, Zhou H, Li L, Zhou S, Shi C, Shi Y, Qiu J, Lu L (2020) MicroRNA-98 inhibits hepatic stellate cell activation and attenuates liver fibrosis by regulating HLF expression. Front Cell Dev Biol 8:513. https://doi.org/10.3389/fcell.2020.00513
Wu YN, He LH, Bai ZT, Li X (2020) NRP1 is a prognostic factor and promotes the growth and migration of cells in intrahepatic cholangiocarcinoma. Cancer Manag Res 12:7021–7032. https://doi.org/10.2147/CMAR.S260091
Xiong WJ, Hu LJ, Jian YC, Wang LJ, Jiang M, Li W, He Y (2012) Wnt5a participates in hepatic stellate cell activation observed by gene expression profile and functional assays. World J Gastroenterol 18:1745–1752. https://doi.org/10.3748/wjg.v18.i15.1745
Yang Y, Yang F, Wu X, Lv X, Li J (2016) EPAC activation inhibits acetaldehyde-induced activation and proliferation of hepatic stellate cell via Rap1. Can J Physiol Pharmacol 94:498–507. https://doi.org/10.1139/cjpp-2015-0437
Zhou W, Chen H, Ruan Y, Zeng X, Liu F (2020) High expression of TRIM15 is associated with tumor invasion and predicts poor prognosis in patients with gastric cancer. J Invest Surg. https://doi.org/10.1080/08941939.2019.1705443
Author information
Authors and Affiliations
Contributions
HL designed the study. JZ and YC performed the experiments and collected the data. YT and SC analyzed the data. All authors contributed to preparation of the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, J., Chen, Y., Tian, Y. et al. Knockdown of TRIM15 inhibits the activation of hepatic stellate cells. J Mol Histol 52, 839–848 (2021). https://doi.org/10.1007/s10735-021-09997-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10735-021-09997-7