Abstract
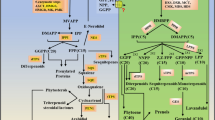
Short cutting is the main method of tea propagation. The callus formation at the base of cuttings has a certain absorption function. The early callus emergence is conducive to the resistance of cuttings to water and nutrient stress, thus promoting rooting and survival. In order to investigate the effect of light quality on the callus formation at the base of cuttings, this study used white (WL), red (660 nm, RL), and blue (430 nm, BL) light emitting diodes (LEDs) to treat tea cuttings. The results showed that compared with WL, BL promoted callus formation and RL inhibited callus formation. In order to explore the internal molecular mechanism of light quality affecting the difference of cutting callus formation, phytohormone determination and full-length transcriptome sequencing were conducted on mature leaves under different light quality treatments. The analysis of phytohormone showed that the contents of abscisic acid (ABA), indole-3-carboxylic acid (ICA), trans-zeatin (tZ), gibberellin 9 (GA9) and 5-deoxystrigol (5DS) were consistent with the callus formation trend, which showed BL > WL > RL. Full-length transcriptome sequencing was performed on mature leaves of cuttings under darkness (DL, CK), WL, RL and BL. Kyoto encyclopedia of genes and genomes (KEGG) analysis of genes, many genes participated in the “plant hormone signal transduction (ko04075)”, such as auxin early response protein AUX/IAA, auxin response factor ARF, two-component response regulator B-ARR, and DELLA genes had highest expression under BL. Meanwhile, auxin synthesis and transport related genes were analyzed,, including L-tryptophan–pyruvate aminotransferase 1-like TAA1, flavin monooxygenase gene YUCCA, PIN1,3,4 and PIN-LIKES6 and 7, which also had the highest expression under BL. In conclusion, this study revealed the molecular mechanism of light quality regulating the callus formation of tea cutting, and provided new insights for the application of light quality in tea breeding.







Similar content being viewed by others
Data availability
The raw data for RNA-seq have been uploaded to the NCBI SRA with accession number PRJNA928604.
References
Achard P, Cheng H, De Grauwe L, Decat J, Schoutteten H, Moritz T, Van Der Straeten D, Peng J, Harberd NP (2006) Integration of plant responses to environmentally activated phytohormonal signals. Science 311:91–94. https://doi.org/10.1126/science.1118642
Achard P, Gong F, Cheminant S, Alioua M, Hedden P, Genschik P (2008a) The cold-inducible CBF1 factor-dependent signaling pathway modulates the accumulation of the growth-repressing DELLA proteins via its effect on gibberellin metabolism. Plant Cell 20:2117–2129. https://doi.org/10.1105/tpc.108.058941
Achard P, Renou JP, Berthomé R, Harberd NP, Genschik P (2008b) Plant DELLAs restrain growth and promote survival of adversity by reducing the levels of reactive oxygen species. Curr Biol 18:656–660. https://doi.org/10.1016/j.cub.2008.04.034
Adamowski M, Friml J (2015) PIN-dependent auxin transport: action, regulation, and evolution. Plant Cell 27:20–32. https://doi.org/10.1105/tpc.114.134874
Arend M, Schnitzler JP, Ehlting B, Hänsch R, Lange T, Rennenberg H, Himmelbach A, Grill E, Fromm J (2009) Expression of the Arabidopsis mutant ABI1 gene alters abscisic acid sensitivity, stomatal development, and growth morphology in gray poplars. Plant Physiol 151:2110–2119. https://doi.org/10.1104/pp.109.144956
Barbez E, Kubeš M, Rolčík J, Béziat C, Pěnčík A, Wang B, Rosquete MR, Zhu J, Dobrev PI, Lee Y, Zažímalovà E, Petrášek J, Geisler M, Friml J, Kleine-Vehn J (2012) A novel putative auxin carrier family regulates intracellular auxin homeostasis in plants. Nature 485:119–122. https://doi.org/10.1038/nature11001
Bernasconi P (2010) Effect of synthetic and natural protein tyrosine kinase inhibitors on auxin efflux in zucchini (Cucurbita pepo) hypocotyls. Physiol Plant 96:205–210
Béziat C, Barbez E, Feraru MI, Lucyshyn D, Kleine-Vehn J (2017) Light triggers PILS-dependent reduction in nuclear auxin signalling for growth transition. Nat Plants 3:17105. https://doi.org/10.1038/nplants.2017.105
Blilou I, Xu J, Wildwater M, Willemsen V, Paponov I, Friml J, Heidstra R, Aida M, Palme K, Scheres B (2005) The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433:39–44. https://doi.org/10.1038/nature03184
Bustillo-Avendaño E, Ibáñez S, Sanz O, Sousa Barros JA, Gude I, Perianez-Rodriguez J, Micol JL, Del Pozo JC, Moreno-Risueno MA, Pérez-Pérez JM (2018) Regulation of Hormonal Control, Cell Reprogramming, and patterning during De Novo Root Organogenesis. Plant Physiol 176:1709–1727. https://doi.org/10.1104/pp.17.00980
Casanova-Sáez R, Voß U (2019) Auxin Metabolism Controls Developmental decisions in land plants. Trends Plant Sci 24:741–754. https://doi.org/10.1016/j.tplants.2019.05.006
Cheminant S, Wild M, Bouvier F, Pelletier S, Renou JP, Erhardt M, Hayes S, Terry MJ, Genschik P, Achard P (2011) DELLAs regulate chlorophyll and carotenoid biosynthesis to prevent photooxidative damage during seedling deetiolation in Arabidopsis. Plant Cell 23:1849–1860. https://doi.org/10.1105/tpc.111.085233
de Lucas M, Davière JM, Rodríguez-Falcón M, Pontin M, Iglesias-Pedraz JM, Lorrain S, Fankhauser C, Blázquez MA, Titarenko E, Prat S (2008) A molecular framework for light and gibberellin control of cell elongation. Nature 451:480–484. https://doi.org/10.1038/nature06520
Fan M, Xu C, Xu K, Hu Y (2012) LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res 22:1169–1180. https://doi.org/10.1038/cr.2012.63
Feraru E, Feraru MI, Barbez E, Waidmann S, Sun L, Gaidora A, Kleine-Vehn J (2019) PILS6 is a temperature-sensitive regulator of nuclear auxin input and organ growth in Arabidopsis thaliana. Proc Natl Acad Sci U S A 116:3893–3898. https://doi.org/10.1073/pnas.1814015116
Friml J, Vieten A, Sauer M, Weijers D, Schwarz H, Hamann T, Offringa R, Jürgens G (2003) Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 426:147–153. https://doi.org/10.1038/nature02085
Fukaki H, Tameda S, Masuda H, Tasaka M (2002) Lateral root formation is blocked by a gain-of-function mutation in the SOLITARY-ROOT/IAA14 gene of Arabidopsis. Plant J 29:153–168. https://doi.org/10.1046/j.0960-7412.2001.01201.x
Fukaki H, Nakao Y, Okushima Y, Theologis A, Tasaka M (2005) Tissue-specific expression of stabilized SOLITARY-ROOT/IAA14 alters lateral root development in Arabidopsis. Plant J 44:382–395. https://doi.org/10.1111/j.1365-313X.2005.02537.x
Goh T, Kasahara H, Mimura T, Kamiya Y, Fukaki H (2012) Multiple AUX/IAA-ARF modules regulate lateral root formation: the role of Arabidopsis SHY2/IAA3-mediated auxin signalling. Philos Trans R Soc Lond B Biol Sci 367:1461–1468. https://doi.org/10.1098/rstb.2011.0232
Goren R, Altman A, Giladi I (1979) Role of Ethylene in Abscisic Acid-induced callus formation in Citrus Bud cultures. Plant Physiol 63:280–282. https://doi.org/10.1104/pp.63.2.280
Hu Y, Bao F, Li J (2000) Promotive effect of brassinosteroids on cell division involves a distinct CycD3-induction pathway in Arabidopsis. Plant J 24:693–701. https://doi.org/10.1046/j.1365-313x.2000.00915.x
Iwase A, Mitsuda N, Koyama T, Hiratsu K, Kojima M, Arai T, Inoue Y, Seki M, Sakakibara H, Sugimoto K, Ohme-Takagi M (2011) The AP2/ERF transcription factor WIND1 controls cell dedifferentiation in Arabidopsis. Curr Biol 21:508–514. https://doi.org/10.1016/j.cub.2011.02.020
Krecek P, Skupa P, Libus J, Naramoto S, Tejos R, Friml J, Zazímalová E (2009) The PIN-FORMED (PIN) protein family of auxin transporters. Genome Biol 10:249. https://doi.org/10.1186/gb-2009-10-12-249
Laskowski M, Grieneisen VA, Hofhuis H, Hove CA, Hogeweg P, Marée AF, Scheres B (2008) Root system architecture from coupling cell shape to auxin transport. PLoS Biol 6:e307. https://doi.org/10.1371/journal.pbio.0060307
Li J, Wen J, Lease KA, Doke JT, Tax FE, Walker JC (2002) BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 110:213–222. https://doi.org/10.1016/s0092-8674(02)00812-7
Liu YY, Wang RL, Zhang P, Sun LL, Xu J (2016) Involvement of reactive oxygen species in lanthanum-induced inhibition of primary root growth. J Exp Bot 67:6149–6159. https://doi.org/10.1093/jxb/erw379
Lu Y, Sasaki Y, Li X, Mori IC, Matsuura T, Hirayama T, Sato T, Yamaguchi J (2015) ABI1 regulates carbon/nitrogen-nutrient signal transduction independent of ABA biosynthesis and canonical ABA signalling pathways in Arabidopsis. J Exp Bot 66:2763–2771. https://doi.org/10.1093/jxb/erv086
Maierhofer T, Diekmann M, Offenborn JN, Lind C, Bauer H, Hashimoto K, KA S A-R, Luan S, Kudla J, Geiger D, Hedrich R (2014) Site- and kinase-specific phosphorylation-mediated activation of SLAC1, a guard cell anion channel stimulated by abscisic acid. Sci Signal 7:ra86. https://doi.org/10.1126/scisignal.2005703
Marchant A, Bhalerao R, Casimiro I, Eklöf J, Casero PJ, Bennett M, Sandberg G (2002) AUX1 promotes lateral root formation by facilitating indole-3-acetic acid distribution between sink and source tissues in the Arabidopsis seedling. Plant Cell 14:589–597. https://doi.org/10.1105/tpc.010354
Mason MG, Mathews DE, Argyros DA, Maxwell BB, Kieber JJ, Alonso JM, Ecker JR, Schaller GE (2005) Multiple type-B response regulators mediate cytokinin signal transduction in Arabidopsis. Plant Cell 17:3007–3018. https://doi.org/10.1105/tpc.105.035451
Mravec J, Skůpa P, Bailly A, Hoyerová K, Krecek P, Bielach A, Petrásek J, Zhang J, Gaykova V, Stierhof YD, Dobrev PI, Schwarzerová K, Rolcík J, Seifertová D, Luschnig C, Benková E, Zazímalová E, Geisler M, Friml J (2009) Subcellular homeostasis of phytohormone auxin is mediated by the ER-localized PIN5 transporter. Nature 459:1136–1140. https://doi.org/10.1038/nature08066
Nam KH, Li J (2002) BRI1/BAK1, a receptor kinase pair mediating brassinosteroid signaling. Cell 110:203–212. https://doi.org/10.1016/s0092-8674(02)00814-0
Nolan TM, Vukašinović N, Liu D, Russinova E, Yin Y (2020) Brassinosteroids: Multidimensional regulators of Plant Growth, Development, and stress responses. Plant Cell 32:295–318. https://doi.org/10.1105/tpc.19.00335
Olatunji D, Geelen D, Verstraeten I (2017) Control of endogenous auxin levels in Plant Root Development. Int J Mol Sci 18DOI. https://doi.org/10.3390/ijms18122587
OuYang F, Mao JF, Wang J, Zhang S, Li Y (2015) Transcriptome analysis reveals that Red and Blue Light regulate growth and phytohormone metabolism in Norway Spruce [Picea abies (L.) Karst]. PLoS ONE 10:e0127896DOI. https://doi.org/10.1371/journal.pone.0127896
Péret B, Swarup K, Ferguson A, Seth M, Yang Y, Dhondt S, James N, Casimiro I, Perry P, Syed A, Yang H, Reemmer J, Venison E, Howells C, Perez-Amador MA, Yun J, Alonso J, Beemster GT, Laplaze L, Murphy A, Bennett MJ, Nielsen E, Swarup R (2012) AUX/LAX genes encode a family of auxin influx transporters that perform distinct functions during Arabidopsis development. Plant Cell 24:2874–2885. https://doi.org/10.1105/tpc.112.097766
Quan J, Ni R, Wang Y, Sun J, Ma M, Bi H (2022) Effects of different growth regulators on the Rooting of Catalpa bignonioides Softwood Cuttings. Life (Basel) 12. https://doi.org/10.3390/life12081231
Raghavendra AS, Gonugunta VK, Christmann A, Grill E (2010) ABA perception and signalling. Trends Plant Sci 15:395–401. https://doi.org/10.1016/j.tplants.2010.04.006
Sauer M, Kleine-Vehn J (2019) PIN-FORMED and PIN-LIKES auxin transport facilitators. Dev 146 DOI. https://doi.org/10.1242/dev.168088
Shang B, Xu C, Zhang X, Cao H, Xin W, Hu Y (2016) Very-long-chain fatty acids restrict regeneration capacity by confining pericycle competence for callus formation in Arabidopsis. Proc Natl Acad Sci U S A 113:5101–5106. https://doi.org/10.1073/pnas.1522466113
Shen Y, Fan K, Wang Y, Wang H, Ding S, Song D, Shen J, Li H, Song Y, Han X, Qian W, Ma Q, Ding Z (2022) Red and Blue Light affect the formation of adventitious roots of tea cuttings (Camellia sinensis) by regulating hormone synthesis and signal transduction pathways of mature Leaves. Front Plant Sci 13:943662. https://doi.org/10.3389/fpls.2022.943662
Shin SY, Choi Y, Kim SG, Park SJ, Park JS, Moon KB, Kim HS, Jeon JH, Cho HS, Lee HJ (2022) Submergence promotes auxin-induced callus formation through ethylene-mediated post-transcriptional control of auxin receptors. Mol Plant 15:1947–1961. https://doi.org/10.1016/j.molp.2022.11.001
Statements & Declarations
Stepanova AN, Robertson-Hoyt J, Yun J, Benavente LM, Xie DY, Dolezal K, Schlereth A, Jürgens G, Alonso JM (2008) TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 133:177–191. https://doi.org/10.1016/j.cell.2008.01.047
Sun L, Feraru E, Feraru MI, Waidmann S, Wang W, Passaia G, Wang ZY, Wabnik K, Kleine-Vehn J (2020) PIN-LIKES coordinate Brassinosteroid Signaling with Nuclear Auxin Input in Arabidopsis thaliana. Curr Biol 30:1579–1588e1576. https://doi.org/10.1016/j.cub.2020.02.002
Suo J, Zhou C, Zeng Z, Li X, Bian H, Wang J, Zhu M, Han N (2021) Identification of regulatory factors promoting embryogenic callus formation in barley through transcriptome analysis. BMC Plant Biol 21:145. https://doi.org/10.1186/s12870-021-02922-w
Swarup R, Péret B (2012) AUX/LAX family of auxin influx carriers-an overview. Front Plant Sci 3:225. https://doi.org/10.3389/fpls.2012.00225
Swarup R, Friml J, Marchant A, Ljung K, Sandberg G, Palme K, Bennett M (2001) Localization of the auxin permease AUX1 suggests two functionally distinct hormone transport pathways operate in the Arabidopsis root apex. Genes Dev 15:2648–2653. https://doi.org/10.1101/gad.210501
Swarup R, Kargul J, Marchant A, Zadik D, Rahman A, Mills R, Yemm A, May S, Williams L, Millner P, Tsurumi S, Moore I, Napier R, Kerr ID, Bennett MJ (2004) Structure-function analysis of the presumptive Arabidopsis auxin permease AUX1. Plant Cell 16:3069–3083. https://doi.org/10.1105/tpc.104.024737
Swarup K, Benková E, Swarup R, Casimiro I, Péret B, Yang Y, Parry G, Nielsen E, De Smet I, Vanneste S, Levesque MP, Carrier D, James N, Calvo V, Ljung K, Kramer E, Roberts R, Graham N, Marillonnet S, Patel K, Jones JD, Taylor CG, Schachtman DP, May S, Sandberg G, Benfey P, Friml J, Kerr I, Beeckman T, Laplaze L, Bennett MJ (2008) The auxin influx carrier LAX3 promotes lateral root emergence. Nat Cell Biol 10:946–954. https://doi.org/10.1038/ncb1754
Tanaka H, Hashimoto N, Kawai S, Yumoto E, Shibata K, Tameshige T, Yamamoto Y, Sugimoto K, Asahina M, Ikeuchi M (2022) Auxin-induced WUSCHEL-RELATED HOMEOBOX13 mediates asymmetric activity of callus formation upon cutting. Plant Cell Physiol. https://doi.org/10.1093/pcp/pcac146
Tatematsu K, Kumagai S, Muto H, Sato A, Watahiki MK, Harper RM, Liscum E, Yamamoto KT (2004) MASSUGU2 encodes Aux/IAA19, an auxin-regulated protein that functions together with the transcriptional activator NPH4/ARF7 to regulate differential growth responses of hypocotyl and formation of lateral roots in Arabidopsis thaliana. Plant Cell 16:379–393. https://doi.org/10.1105/tpc.018630
Tian Q, Reed JW (1999) Control of auxin-regulated root development by the Arabidopsis thaliana SHY2/IAA3 gene. Development 126:711–721. https://doi.org/10.1242/dev.126.4.711
Uehara T, Okushima Y, Mimura T, Tasaka M, Fukaki H (2008) Domain II mutations in CRANE/IAA18 suppress lateral root formation and affect shoot development in Arabidopsis thaliana. Plant Cell Physiol 49:1025–1038. https://doi.org/10.1093/pcp/pcn079
Vanneste S, Friml J (2009) Auxin: a trigger for change in plant development. Cell 136:1005–1016. https://doi.org/10.1016/j.cell.2009.03.001
Wang X, Kota U, He K, Blackburn K, Li J, Goshe MB, Huber SC, Clouse SD (2008) Sequential transphosphorylation of the BRI1/BAK1 receptor kinase complex impacts early events in brassinosteroid signaling. Dev Cell 15:220–235. https://doi.org/10.1016/j.devcel.2008.06.011
Wang P, Cheng T, Wu S, Zhao F, Wang G, Yang L, Lu M, Chen J, Shi J (2014) Phylogeny and molecular evolution analysis of PIN-FORMED 1 in angiosperm. PLoS ONE 9:e89289. https://doi.org/10.1371/journal.pone.0089289
Wang Y, Pang D, Ruan L, Liang J, Zhang Q, Qian Y, Zhang Y, Bai P, Wu L, Cheng H, Cui Q, Wang L, Wei K (2022) Integrated transcriptome and hormonal analysis of naphthalene acetic acid-induced adventitious root formation of tea cuttings (Camellia sinensis). BMC Plant Biol 22:319. https://doi.org/10.1186/s12870-022-03701-x
Yang Y, Hammes UZ, Taylor CG, Schachtman DP, Nielsen E (2006) High-affinity auxin transport by the AUX1 influx carrier protein. Curr Biol 16:1123–1127. https://doi.org/10.1016/j.cub.2006.04.029
Zhai S, Cai W, Xiang ZX, Chen CY, Lu YT, Yuan TT (2021) PIN3-mediated auxin transport contributes to blue light-induced adventitious root formation in Arabidopsis. Plant Sci 312:111044. https://doi.org/10.1016/j.plantsci.2021.111044
Zhao Y (2010) Auxin biosynthesis and its role in plant development. Annu Rev Plant Biol 61:49–64. https://doi.org/10.1146/annurev-arplant-042809-112308
Zhao Y (2018) Essential roles of local Auxin Biosynthesis in Plant Development and in adaptation to environmental changes. Annu Rev Plant Biol 69:417–435. https://doi.org/10.1146/annurev-arplant-042817-040226
Zhao Y, Christensen SK, Fankhauser C, Cashman JR, Cohen JD, Weigel D, Chory J (2001) A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 291:306–309. https://doi.org/10.1126/science.291.5502.306
Zheng C, Ma JQ, Ma CL, Shen SY, Liu YF, Chen L (2019) Regulation of growth and flavonoid formation of tea plants (Camellia sinensis) by Blue and Green Light. J Agric Food Chem 67:2408–2419. https://doi.org/10.1021/acs.jafc.8b07050
Funding
This research was funded by the Technology System of Modern Agricultural Industry in Shandong Province (SDAIT19-01) and the Special Foundation for Distinguished Taishan Scholar of Shandong Province (ts201712057), the Project of Agricultural Science and Technology Fund in Shandong Province (2019LY002, 2019YQ010, and 2019TSLH0802), Shandong Agricultural Seed Improvement Project (2020LZGC010), the Project of Rizhao Natural Science Foundation Youth Fund (RZ2021ZR48). The Livelihood Project of Qingdao City (22-3-7-xdny-5-nsh). The Agricultural Science and Technology Innovation Project of Shandong Academy of Agricultural Sciences (CXGC2023F18, CXGC2023A11). Natural Science Foundation of Shandong Province (ZR2023QC086).
Author information
Authors and Affiliations
Contributions
Yaozong Shen conducted the experiment, analyzed the data, and wrote a manuscript. Xiao Han participated in the manuscript writing and picture production. Zhaotang Ding, Kai Fan and Yu Wang participated in the experimental design and revised the manuscript. Hui Wang conducted the experiment, collected samples and statistical data. Jiazhi Shen revised the manuscript. He Li, Shibo Ding and Dapeng Song collected samples. All authors contributed to the article and approved the submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Communicated by Ben Zhang.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shen, Y., Han, X., Wang, H. et al. Full-length transcriptome and metabolism revealed the difference of callus formation of tea cutting under white, red and blue light. Plant Growth Regul 101, 715–726 (2023). https://doi.org/10.1007/s10725-023-01052-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-023-01052-7