Abstract
Dehydrins (DHNs) are late embryogenesis-abundant (LEA) proteins, which enhance abiotic stress tolerance in plants. However, little is known regarding the function of DHNs in Suaeda salsa L. (S. salsa), which can grow in saline soil. Here, we successfully cloned and functionally characterized a dehydrin gene from S. salsa, designated SsDHN. Sequence alignment analysis showed that SsDHN belongs to SKn-type DHNs and shares relatively high level of homology with Spinacia oleracea COR47-like (XP_021846321.1). Quantitative RT-PCR analysis indicated that SsDHN expression level increased significantly under salt stress. We also generated SsDHN-overexpressing transgenic tobacco lines and analyzed their salt stress response. Seeds of transgenic tobacco plants grown under a range of salt concentrations (100, 200, and 300 mM) showed significantly higher germination rates relative to wild-type seeds. Transgenic plants had longer root length, lower relative electrical conductivity (REC), lower malondialdehyde (MDA) content, higher proline (PRO) content, increased peroxidase (SOD) activity, and less damage to the chloroplast ultrastructure. Our results showed that the transgenic tobacco plant lines had improved salt resistance and osmotic adjustment, enhanced reactive oxygen species scavenging ability, maintenance of the K+/Na+ balance, and reduced chloroplast membrane damage. These results suggest that the SsDHN gene may be used for improving abiotic stress tolerance in economically important crops.







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this article.
References
Adem GD, Roy SJ, Zhou M, Bowman JP, Shabala S (2014) Evaluating contribution of ionic, osmotic and oxidative stress components towards salinity tolerance in barley. BMC Plant Biol 14:113. https://doi.org/10.1186/1471-2229-14-113
Aguayo P, Sanhueza J, Noriega F, Ochoa M, Lefeuvre R, Navarrete D, Fernández M, Valenzuela S (2016) Overexpression of an SKn-dehydrin gene from Eucalyptus globulus and Eucalyptus nitens enhances tolerance to freezing stress in Arabidopsis. Trees 30:1785–1797. https://doi.org/10.1007/s00468-016-1410-9
Ahanger MA, Akram NA, Ashraf M, Alyemeni MN, Wijaya L, Ahmad P (2017) Plant responses to environmental stresses—from gene to biotechnology. AoB Plants 9:plx02. https://doi.org/10.1093/aobpla/plx025
Anwaar SA, Ali S, Ali S, Ishaque W, Farid M, Farooq MA, Najeeb U, Abbas F, Sharif M (2015) Silicon (Si) alleviates cotton (Gossypium hirsutum L.) from zinc (Zn) toxicity stress by limiting Zn uptake and oxidative damage. Environ Sci Pollut Res Int 22:3441–3450. https://doi.org/10.1007/s11356-014-3938-9
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24:23–58. https://doi.org/10.1080/07352680590910410
Bartwal A, Mall R, Lohani P, Guru SK, Arora S (2013) Role of secondary metabolites and brassinosteroids in plant defense against environmental stresses. J Plant Growth Regul 32:216–232. https://doi.org/10.1007/s00344-012-9272-x
Battaglia M, Olvera-Carrillo Y, Garciarrubio A, Campos F, Covarrubias AA (2008) The enigmatic LEA proteins and other hydrophilins. Plant Physiol 148:6–24. https://doi.org/10.1104/pp.108.120725
Ben Rejeb K, Abdelly C, Savouré A (2014) How reactive oxygen species and proline face stress together. Plant Physiol Biochem 80:278–284. https://doi.org/10.1016/j.plaphy.2014.04.007
Bhagi P, Zhawar VK, Gupta AK (2013) Antioxidant response and Lea genes expression under salt stress and combined salt plus water stress in two wheat cultivars contrasting in drought tolerance. Ind J Exp Biol 51:746–757. https://doi.org/10.1109/JBHI.2013.2261819
Bojórquez-Quintal E, Ruiz-Lau N, Velarde-Buendía A, Echevarría-Machado I, Pottosin I, Martínez-Estévez M (2016) Natural variation in primary root growth and K+ retention in roots of habanero pepper (Capsicum chinense) under salt stress. Funct Plant Biol 43:1114–1125. https://doi.org/10.1071/FP15391
Bray EA (1993) Molecular responses to water deficit. Plant Physiol 103:1035–1040. https://doi.org/10.1104/pp.103.4.1035
Brini F, Hanin M, Lumbreras V, Amara I, Khoudi H, Hassairi A (2007) Overexpression of wheat dehydrin DHN-5 enhances tolerance to salt and osmotic stress in Arabidopsis thaliana. Plant Cell Rep 26:2017–2026. https://doi.org/10.1007/s00299-007-0412-x
Cao Y, Xiang X, Geng M, You Q, Huang X (2017) Effect of HbDHN1 and HbDHN2 genes on abiotic stress responses in Arabidopsis. Front Plant Sci 8:470. https://doi.org/10.3389/fpls.2017.00470
Close TJ (1996) Dehydrins: emergence of a biochemical role of a family of plant dehydration proteins. Physiol Plant 97:795–803. https://doi.org/10.1034/j.1399-3054.1996.970422.x
Close TJ (2010) Dehydrins: a commonalty in the response of plants to dehydration and low temperature. Physiol Plantarum 100:291–296. https://doi.org/10.1111/j.1399-3054.1997.tb04785.x
Graether SP, Boddington KF (2014) Disorder and function: a review of the dehydrin protein family. Front Plant Sci 5:576. https://doi.org/10.3389/fpls.2014.00576
Grelet J, Benamar A, Teyssier E, Avelange-Macherel MH, Grunwald D, Macherel D (2005) Identification in pea seed mitochondria of a late-embryogenesis abundant protein able to protect enzymes from drying. Plant Physiol 137:157–167. https://doi.org/10.1104/pp.104.052480
Guntzer F, Keller C, Meunie JD (2012) Benefits of plant silicon for crops: a review. Agron Sustain Dev 32:201–213. https://doi.org/10.1007/s13593-011-0039-8
Guo X, Zhang L, Zhu J, Liu H, Wang A (2017) Cloning and characterization of SiDHN, a novel dehydrin gene from Saussurea involucrata Kar. et Kir. that enhances cold and drought tolerance in tobacco. Plant Sci 256:160–169. https://doi.org/10.1016/j.plantsci.2016.12.007
Guo JR, Dong XX, Li Y, Wang BS (2020a) NaCl treatment markedly enhanced pollen viability and pollen preservation time of euhalophyte Suaeda salsa via up regulation of pollen development-related genes. J Plant Res 133:57–71. https://doi.org/10.1007/s10265-019-01148-0
Guo JR, Lu CX, Zhao FC, Gao S, Wang BS (2020b) Improved reproductive growth of euhalophyte Suaeda salsa under salinity is correlated with altered phytohormone biosynthesis and signal transduction. Funct Plant Biol 47:170–183. https://doi.org/10.1071/Fp19215
Hara M (2010) The multifunctionality of dehydrins: an overview. Plant Signal Behav 5:503–508. https://doi.org/10.4161/psb.11085
Hariadi Y, Marandon K, Tian Y, Jacobsen SE, Shabala S (2011) Ionic and osmotic relations in quinoa (Chenopodium quinoa Willd.) plants grown at various salinity levels. J Exp Bot 62:185–193. https://doi.org/10.1093/jxb/erq257
Hughes SL, Schart V, Malcolmson J, Hogarth KA, Martynowicz DM, Tralman-Baker E, Patel SN, Graether SP (2013) The importance of size and disorder in the cryoprotective effects of dehydrins. Plant Physiol 163:1376–1386. https://doi.org/10.1104/pp.113.226803
Hundertmark M, Hincha DK (2008) LEA (Late embryogenesis abundant) proteins and their encoding gene in Arabidopsis thaliana. BMC Genome 9:118. https://doi.org/10.1186/1471-2164-9-118
Jin X, Cao D, Wang Z, Ma L, Tian K, Liu Y, Gong Z, Zhu X, Jiang C, Li Y (2019) Genome-wide identification and expression analyses of the LEA protein gene family in tea plant reveal their involvement in seed development and abiotic stress responses. Sci Rep 9:14123. https://doi.org/10.1038/s41598-019-50645-8
Kirungu JN, Magwanga RO, Pu L, Cai X, Xu Y, Hou Y, Zhou Y, Cai Y, Hao F, Zhou Z, Wang K, Liu F (2020) Knockdown of Gh_A05G1554 (GhDHN_03) and Gh_D05G1729 (GhDHN_04) dehydrin genes, reveals their potential role in enhancing osmotic and salt tolerance in cotton. Genomics 112:1902–1915. https://doi.org/10.1016/j.ygeno.2019.11.003
Kumar M, Lee SC, Kim JY, Kim SJ, Aye SS, Kim SR (2014) Over-expression of dehydrin gene, OsDhn1, improves drought and salt stress tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L.). J Plant Biol 57:383–393. https://doi.org/10.1007/s12374-014-0487-1
Li Q, Zhang X, Lv Q, Zhu D, Qiu T, Xu Y, Bao F, He Y, Hu Y (2017) Physcomitrella patens dehydrins (PpDHNA and PpDHNC) confer salinity and drought tolerance to transgenic Arabidopsis plants. Front Plant Sci 8:1316. https://doi.org/10.3389/fpls.2017.01316
Liu H, Yu C, Li H, Ouyang B, Wang T, Zhang J, Wang X, Ye Z (2015) Overexpression of ShDHN, a dehydrin gene from Solanum habrochaites enhances tolerance to multiple abiotic stresses in tomato. Plant Sci 231:198–211. https://doi.org/10.1016/j.plantsci.2014.12.006
Liu Y, Wang L, Zhang T, Yang X, Li D (2017) Functional characterization of KS-type dehydrin ZmDHN13 and its related conserved domains under oxidative stress. Sci Rep 7:7361. https://doi.org/10.1038/s41598-017-07852-y
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)). Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lv A, Wen W, Fan N, Su L, Zhou P, An Y (2021) Dehydrin MsDHN1 improves aluminum tolerance of alfalfa (Medicago sativa L.) by affecting oxalate exudation from root tips. Plant J 108:441–458. https://doi.org/10.1111/tpj.15451
Malik AA, Veltri M, Boddington KF, Singh KK, Graether SP (2017) Genome analysis of conserved dehydrin motifs in vascular plants. Front Plant Sci 8:709. https://doi.org/10.3389/fpls.2017.00709
Meng YC, Zhang HF, Pan XX, Chen N, Hu HF, Ha S, Khan A, Chen RG (2021) CaDHN3, a pepper (capsicum annuum L.) dehydrin gene enhances the tolerance against salt and drought stresses by reducing ROS accumulation. Int J Mol Sci. 22:3205. https://doi.org/10.3390/ijms22063205
Moore K, Roberts LJ (1998) Measurement of lipid peroxidation. Free Radic Res 28:659–671. https://doi.org/10.1002/0471140856.tx0204s00
Qiu H, Zhang L, Liu C, He L, Wang A, Liu HL, Zhu JB (2014) Cloning and characterization of a novel dehydrin gene, SiDhn2, from Saussurea involucrata Kar. et Kir. Plant Mol Biol 84:707–718. https://doi.org/10.1007/s11103-013-0164-7
Rascio AM, Russo L, Mazzucco C, Plantani G, Nicastro G, Di Fonzo N (2001) Enhanced osmotolerance of a wheat mutant selected for potassium accumulation. Plant Sci 160:441–448. https://doi.org/10.1016/s0168-9452(00)00404-0
Saavedra L, Svensson J, Carballo V, Izmendi D, Wellin B, Vidal S (2006) A dehydrin gene in Physcomitrella patens is required for salt and osmotic stress tolerance. Plant J 45:237–249. https://doi.org/10.1111/j.1365-313X.2005.02603.x
Saibi W, Feki K, Ben Mahmoud R, Brini F (2015) Durum wheat dehydrin (DHN-5) confers salinity tolerance to transgenic Arabidopsis plants through the regulation of proline metabolism and ROS scavenging system. Planta 242:1187–1194. https://doi.org/10.1007/s00425-015-2351-z
Serrano R, Montesinos C (2003) Molecular bases of desiccation tolerance in plant cells and potential applications in food dehydration. Food Sci Technol Int 9:157–161. https://doi.org/10.1177/1082013203035518
Shen X, Wang Z, Song X, Xu J, Jiang C, Zhao Y, Ma C, Zhang H (2014) Transcriptomic profiling revealed an important role of cell wall remodeling and ethylene signaling pathway during salt acclimation in Arabidopsis. Plant Mol Biol 86:303–317. https://doi.org/10.1007/s11103-014-0230-9
Song J, Wang BS (2015) Using euhalophytes to understand salt tolerance and to develop saline agriculture: suaeda salsa as a promising model. Ann Bot-London 115:541–553. https://doi.org/10.1093/aob/mcu194
Song J, Shi WW, Liu RR, Xu YG, Sui N, Zhou JC, Feng G (2017) The role of the seed coat in adaptation of dimorphic seeds of the euhalophyte Suaeda salsa to salinity. Plant Spec Biol 32:107–114. https://doi.org/10.1111/1442-1984.12132
Sun Y, Liu L, Sun S, Han W, Irfan M, Zhang X, Zhang L, Chen L (2021) AnDHN, a dehydrin protein from Ammopiptanthus nanus, mitigates the negative effects of drought stress in plants. Front Plant Sci 12:788938. https://doi.org/10.3389/fpls.2021.788938
Sunilkumar G, Vijayachandra K, Veluthambi K (1999) Preincubation of cut tobacco leaf explants promotes Agrobacterium-mediated transformation by increasing vir gene induction. Plant Sci 141:51–58. https://doi.org/10.1016/S0168-9452(98)00228-3
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Umezawa T, Fujita M, Fujita Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Engineering drought tolerance in plants: discovering and tailoring genes unlock the future. Curr Opin Biotechnol 17:113–122. https://doi.org/10.1016/j.copbio.2006.02.002
Vani B, Saradhi PP, Mohanty P (2001) Alteration in chloroplast structure and thylakoid membrane composition due to in vivo heat treatment of rice seedlings: correlation with the functional changes. J of Plant Physiol 158:583–592. https://doi.org/10.1078/0176-1617-00260
Ventura Y, Eshel A, Pasternak D, Sagi M (2015) The development of halophyte-based agriculture: past and present. Ann Bot 115:529–540. https://doi.org/10.1093/aob/mcu173
Verma G, Dhar YV, Srivastava D, Kidwai M, Chauhan PS, Bag SK et al (2017) Genome-wide analysis of rice dehydrin gene family: its evolutionary conservedness and expression pattern in response to PEG induced dehydration stress. PLoS ONE 12:e0176399. https://doi.org/10.1371/journal.pone.0176399
Vornam B, Gailing O, Derory J, Plomion C, Kremer A, Finkeldey R (2011) Characterisation and natural variation of a dehydrin gene in Quercus petraea (Matt.) Liebl. Plant Biol 13:881–887. https://doi.org/10.1111/j.1438-8677.2011.00446.x
Wang Y, Xu H, Zhu H, Tao Y, Zhang G, Zhang L, Zhang C, Zhan Z, Ma Z (2014) Classification and expression diversification of wheat dehydrin genes. Plant Sci 214:113–120. https://doi.org/10.1016/j.plantsci.2013.10.005
Xu H, Yang Y, Xie L, Li X, Feng C, Chen J, Xu C (2014) Involvement of multiple types of dehydrins in the freezing response in loquat (Eriobotrya japonica). PLoS One 9:e87575. https://doi.org/10.1371/journal.pone.0087575
Yang Y, He M, Zhu Z, Li S, Xu Y, Zhang C, Singer SD, Wang Y (2012) Identification of the dehydrin gene family from grapevine species and analysis of their responsiveness to various forms of abiotic and biotic stress. BMC Plant Biol 12:140. https://doi.org/10.1186/1471-2229-12-140
Zepeda-Jazo I, Shabala S, Chen Z, Pottosin II (2008) Na-K transport in roots under salt stress. Plant Signal Behav 3:401–403. https://doi.org/10.4161/psb.3.6.5429
Zhang QF, Li YY, Pang CH, Lu CM, Wang BS (2005) NaCl enhances thylakoid-bound SOD activity in the leaves of C-3 halophyte Suaeda salsa L. Plant Sci 168:423–430. https://doi.org/10.1016/j.plantsci.2004.09.002
Zhu W, Zhang L, Lv H, Zhang H, Zhang D, Wang X, Chen J (2014) The dehydrin wzy2 promoter from wheat defines its contribution to stress tolerance. Funct Integr Genomics 14:111–125. https://doi.org/10.1007/s10142-013-0354-z
Funding
This work was supported by the National Natural Science Foundation of China (Grant No. 31070448) and the Liaoning Province ‘2021 Special Project of Central Government Guiding local scientific technology development’ (2021JH6/10500164). The funding bodies had no roles in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have not disclosed any competing interest.
Additional information
Editorial Responsibiility: Yoichi Sakata.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

10725_2022_908_MOESM1_ESM.jpg
Supplementary file1 (JPG 100 KB)Supplementary Fig. S1 SsDHN clone (847 bp) was PCR amplified from the cDNA of S. salsa leaves via RT-PCR. (A) Products of conserved regions were obtained by RT-PCR. (B) 3′-RACE products were obtained by RT-PCR. (C) The full-length cDNA was obtained by RT-PCR.

10725_2022_908_MOESM2_ESM.jpg
Supplementary file2 (JPG 180 KB)Supplementary Fig. S2 SsDHN sequence information. (A) Nucleotide and deduced amino acid sequences of SsDHN cDNA. Nucleotides are numbered at right. The start (ATG) and stop codons (TAG) in the frames are boxed; (B) Multiple sequence alignment of SsDHN with homologous sequences. Bold lines indicate conserved amino acid sequences of S- and K-segment motifs. SsDHN (AGC55011.1); CqDHN (XP_021756500.1, XP_021732246.1); AcDHN (AFC98463.1); AhDHN (AGZ86543.1); SoDHN (XP_021846321.1); SgDHN (NCBI AEA29617.1); CbDHN (ABV56004.1); AtDHN (CAA62449.1); McDHN (XP_022152554.1); MeDHN (XP_021614140.1); CcDHN (ABC68275.1).


10725_2022_908_MOESM4_ESM.jpg
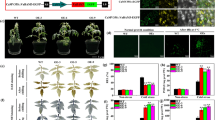
Supplementary file4 (JPG 86 KB)Supplementary Fig. S3 Molecular analysis of tobacco plant lines. (A) Electrophoregram of PCR products; (B) Southern blot analysis of SsDHN transgene in transgenic tobacco lines; (C) Western blot analysis of SsDHN expression in transgenic tobacco lines. M: DL2000 DNA marker, CK+: the Agrobacterium plasmid transformed into the PBI121-DHN recombinant vector was used as the positive control, CK-: WT tobacco, 1-6: transgenic tobacco.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, H., Zhang, L., Lin, J. et al. SsDHN, a dehydrin protein from Suaeda salsa, enhances salt stress tolerance in transgenic tobacco plants. Plant Growth Regul 99, 299–312 (2023). https://doi.org/10.1007/s10725-022-00908-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-022-00908-8