Abstract
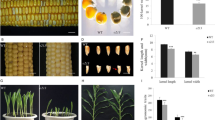
Endosperm development plays an important role in the determination of grain weight in maize. In this context, a spontaneous recessive shrunken kernel mutant, shrunken1-m (sh1-m), identified from improved maize inbred line Zheng58 (Gai-Z58) was studied. Physiological and microscopic analysis revealed that the sh1-m mutant significantly increased the content of amylose, and its starch granules showed distinctive morphology. Utilizing 1877 recessive sh1-m individuals from a BC1 segregating population of B73/sh1-m and 19 molecular markers, the sh1-m gene was limited to a 2.1 kb interval within the GRMZM2G089713 gene. A 5-bp substitution in sh1-m covering the 3′ end of the thirteenth exon and the 5′ splicing site of the thirteenth intron led to a missense mutation and a mis-splicing site that resulted in early translational termination. sh1 reference mutant sh1-912A (shrunken-912A) had a single transversion at the 3′ splice junction of the second intron resulting in a mis-splicing site causing the 13-bp spliced out in the third exon and the initiation codon to shift to 118-bp downstream. Further genetic allelism tests between them confirmed that sh1-m is a novel allele of the Sh1 locus. Sh1 was constitutively expressed in all tested tissues, and its expression pattern/level was not significantly changed in developing seeds of the sh1-m mutant. Transcript mRNA patterns/levels of some other genes involved in starch biosynthesis was significantly altered in the sh1-m mutant. These results provide direct evidence that sh1-m plays a key role in starch biosynthesis and provide valuable information for grain quality research and breeding in maize.




Similar content being viewed by others
Abbreviations
- AGPase:
-
ADP-glucose pyrophosphorylase
- SS:
-
Starch synthases
- GBSS:
-
Granule-bound starch synthase
- SBE:
-
Starch branching enzymes
- DBE:
-
Debranching enzyme
- CTAB:
-
Cetyl trimethyl ammonium bromide
- SSR:
-
Simple sequence repeat
- HTGS:
-
High throughput genomic sequences
- PCR:
-
Polymerase chain reaction
- InDel:
-
Insertion/deletion
- SNP:
-
Single nucleotide polymorphism
- ORF:
-
Open reading frame
- US:
-
United States
- NCBI:
-
National Center for Biotechnology Information
- R:
-
Root
- S:
-
Stem
- L:
-
Leaf
- E:
-
Ear
- T:
-
Tassel
- DAP:
-
Days after pollination
- EMS:
-
Ethyl methane sulfonate
References
Anderson JM, Robertson DS, Morris DW (1991) Molecular characterization of four shrunken mutations induced in Mutator lines in Zea mays L. Plant Sci 77(1):93–101
Bae JM, Giroux MJ, Hannah LC (1990) Cloning and characterization of the Brittle-2 gene of maize. Maydica 35(4):317–322
Bhave MR, Lawrence S, Barton C, Hannah LC (1990) ldentification and molecular characterization of Shrunken-2 cDNA clones of maize. Plant Cell 2(6):581–588
Boyer CD, Liu KC (1985) The interaction of endosperm genotype and genetic background. 1. Differences in chromatographic profiles of starches from nonmutant and mutant endosperms. Starke 37:73
Boyer CD, Preiss J (1981) Evidence for independent genetic control of the multiple forms of maize endosperm branching enzymes and starch synthases. Plant Physiol 67(6):1141–1145
Burr B, Burr FA (1981) Detection of changes in maize DNA at the shrunken locus due to the intervention of Ds elements. Cold Spring Harb Symp Quant Biol 45:463–465
Cao HP, Imparl-Redosevich J, Guan HP, Keeling PL, James MG, Myers AM (1999) Identification of the soluble starch synthase activities of maize endosperm. Plant Physiol 120(1):205–215
Carlson SJ, Chourey PS (1996) Evidence for plasma membrane-associated forms of sucrose synthase in maize. Mol Gen Genet 252(3):303–310
Carlson SJ, Chourey PS, Helentjaris T, Datta R (2002) Gene expression studies on developing kernels of maize sucrose synthase (SuSy) mutants show evidence for a third SuSy gene. Plant Mol Biol 49(1):15–29
Chai XJ, Wang PW, Guan SY, Xu YW (2005) Reducing the maize amylopectin content through RNA interference manipulation. J Plant Physiol Mol Biol 31(6):625–630 (in Chinese)
Chourey PS (1981) Genetic control of sucrose synthase in maize endosperm. Mol Gen Genet 184:372–376
Chourey PS, Nelson OE (1976) The enzymatic deficiency conditioned by the shrunken-1 mutations in maize. Biochem Genet 14(11–12):1041–1055
Chourey PS, Schwartz D (1971) Ethyl methanesulfonate induced mutations of the Sh protein in maize. Mutat Res 12:151–157
Courage-Tebbe U, Doring HP, Fedoroff N, Starlinger P (1983) The controlling element Ds at the Shrunken locus in Zea mays: structure of the unstable sh-m5933 allele and several revertants. Cell 34(2):383–393
Dinges JR, Colleoni C, James MG, Myers AM (2003) Mutational analysis of the pullulanase-type debranching enzyme of maize indicates multiple functions in starch metabolism. Plant Cell 15(3):666–680
Dooner HK (1981) Regulation of the enzyme UFGT by the controlling element Ds in bz-m4, an unstable mutant in maize. Cold Spring Harbor Symp Quant Biol 45:457–462
Dooner HK, Nelson OE (1977) Controlling element-induced alterations in UDPglucose: flavonoid glucosyltransferase, the enzyme specified by the bronze locus in maize. Proc Natl Acad Sci USA 74(12):5623–5627
Doring HP, Pahl I, Durany M (1990) Chromosomal rearrangements caused by the aberrant transposition of double Ds elements are formed by Ds and adjacent non- Ds sequences. Mol Gen Genet 224(1):40–48
Duncan KA, Hardin SC, Huber SC (2006) The three maize sucrose synthase isoforms differ in distribution, localization, and phosphorylation. Plant Cell Physiol 47(7):959–971
Dunn GM, Kramer HH, Whistler RL (1953) Gene dosage effects on corn endosperm carbohydrates. Agron J 45:101–104
Fisher DK, Boyer CD, Hannah LC (1993) Starch branching enzyme II from maize endosperm. Plant Physiol 102(3):1045–1046
Fisher DK, Gao M, Kim KN, Boyer CD, Guiltinan MJ (1996) Allelic analysis of the maize amylose-extender locus suggests that independent genes encode starch-branching enzymes IIa and IIb. Plant Physiol 110(2):611–619
Gao M, Wanat J, Stinard PS, James MG, Myers AM (1998) Characterization of dull1, a maize gene coding for a novel starch synthase. Plant Cell 10(3):399–412
Guan SY, Wang PW, Liu HJ, Liu GN, Ma YY, Zhao LN (2011) Production of high-amylose maize lines using RNA interference in sbe2a. Afr J Biotechnol 10(68):15229–15237
Guo ZH, Zhang JW, Wang D, Chen ZH (2008) Using RNAi technology to produce high-amylose potato plants. Sci Agric Sin 41:494–501 ( in Chinese)
Gupta HS, Raman B, Agrawal PK, Mahajan V, Hossain F, Thirunavukkarasu N (2013) Accelerated development of quality protein maize hybrid through marker-assisted introgression of opaque-2 allele. Plant Breed 132(1):77–82
Hannah LC (2005) Starch synthesis in the maize endosperm. Maydica 50:497–506
Hannah LC, James M (2008) The complexities of starch biosynthesis in cereal endosperm. Curr Opin Biotechnol 19(2):160–165
Hennen-Bierwagen TA, Liu FS, Marsh RS, Kim S, Gan QL, Tetlow IJ et al (2008) Starch biosynthetic enzymes from developing maize endosperm associate in multisubunit complexes. Plant Physiol 146:1892–1908
Holder DG, Glover DV, Shannon JC (1974) Interaction of shrunken-2 with five other carbohydrate genes in corn endosperm. Crop Sci 14:643–646
Hutchison CB (1921) Heritable characters of maize vii. shrunken endosperm. J Hered 12(2):76–83
James MG, Robertson DS, Myers AM (1995) Characterization of the maize gene sugary1, a determinant of starch composition in kernels. Plant Cell 7(4):417–429
Jeon JS, Ryoo N, Hahn TR, Walia H, Nakamura Y (2010) Starch biosynthesis in cereal endosperm. Plant Physiol Biochem 48:383–392
Jiang HX, Campbell M, Blanco M, Jane JL (2010) Characterization of maize amylose-extender (ae) mutant starches: Part II. Structures and properties of starch residues remaining after enzymatic hydrolysis at boiling-water temperature. Carbohydr Polym 80(1):1–12
Jiang LL, Yu XM, Qi X, Yu Q, Deng S, Bai B et al (2013) Multigene engineering of starch biosynthesis in maize endosperm increases the total starch content and the proportion of amylose. Transgenic Res 22(6):1133–1142
Kim KN, Fisher DK, Gao M, Guiltinan MJ (1998) Molecular cloning and characterization of the amylose-extender gene encoding starch branching enzyme IIB in maize. Plant Mol Biol 38(6):945–956
Kirchberger S, Leroch M, Huynen MA, Wahl M, Neuhaus HE, Tjaden J (2007) Molecular and biochemical analysis of the plastidic ADP-glucose transporter (ZmBT1) from Zea mays. J Biol Chem 282(31):22481–22491
Kramer HH, Pfahler PL, Whistler RJ (1958) Gene interaction in maize affecting endosperm properties. Agronomy J 50(4):207–210
Li Q, Wan JM (2005) SSRHunter: development of a local searching software for SSR sites. Hereditas 27(5):808–810
Liang JS, Zhang JH, Cao XZ (2001) Grain sink strength may be related to the poor grain filling of indica-japonica rice (Oryza sativa) hybrids. Physiol Plant 112(4):470–477
Lin LS, Guo DW, Zhao LX, Zhang XD, Wang J, Zhang FM et al (2016) Comparative structure of starches from high-amylose maize inbred lines and their hybrids. Food Hydrocoll 52:19–28
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-DDCT method. Methods 25(4):402–408
Martin C, Smith AM (1995) Starch Biosynthesis. Plant Cell 7(7):971–985
Myers AM, Morell MK, James MG, Ball SG (2000) Recent progress toward understanding biosynthesis of the amylopectin crystal. Plant Physiol 122(4):989–997
Nelson O, Pan D (1995) Starch synthesis in maize endosperm. Annu Rev Plant Physiol Plant Mol Biol 46:475–496
Nelson OE, Rines HW (1962) The enzymatic deficiency in the waxy mutant of maize. Biochem Biophys Res Commun 4(9):297–300
Ortiz DF, Rowland LJ, Gregerson RG, Strommer JN (1988) Insertion of Mu into the Shrunken 1 gene of maize affects transcriptional and post-transcriptional regulation of Sh1 RNA. Mol Gen Genet 214(1):135–141
Prasanna BM, Pixley K, Warburton ML, Xie CX (2010) Molecular marker-assisted breeding for maize improvement in Asia. Mol Breed 26(2):339–356
Preiss J, Sivak MN (1998) Biochemistry, molecular biology and regulation of starch synthesis. Genet Eng 20:177–223
Prioul JL, Mechin V, Lessard P, Thevenot C, Grimmer M, Chateau-Joubert S et al (2008) A joint transcriptomic, proteomic and metabolic analysis of maize endosperm development and starch filling. Plant Biotechnol J 6:855–869
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81(24):8014–8018
Schwartz D (1960) Electrophoretic and immunochemical studies with endosperm proteins of maize mutants. Genetics 45(10):1419–1427
Slattery CJ, Kavakli IH, Okita TW (2000) Engineering starch for increased quantity and quality. Trends Plant Sci 5(7):291–298
Takeda Y, Preiss J (1993) Structures of B90 (sugary) and W64A (normal) maize starches. Carbohydr Res 240:265–275
Teng B, Zhang C, Zhang Y, Wu JD, Li ZF, Luo ZX et al (2015) Comparison of amylopectin structure and activities of key starch synthesis enzymes in the grains of rice single-segment substitution lines with different Wx alleles. Plant Growth Regul 77:117–124
Tian ZX, Qian Q, Liu QQ, Yan MX, Liu XF, Yan CJ et al (2009) Allelic diversities in rice starch biosynthesis lead to a diverse array of rice eating and cooking qualities. Proc Natl Acad Sci USA 106:21760–21765
Tracy WF (1997) History, genetics, and breeding of supersweet [shrunken2] sweet corn. Plant Breed Rev 14:189–236
Wang YJ, White P, Pollak L, Jane J (1993) Characterization of starch structures of 17 maize endosperm mutant genotypes with Oh43 inbred line background. Cereal Chem 70(2):171–179
Werr W, Frommer WB, Maas C, Starlinger P (1985) Sturcture and sucrose synthase gene on 9 of Zea mays L. EMBO J 4(6):1373–1380
Whitt SR, Wilson LM, Tenaillon MI, Gaut BS, Buckler ES (2002) Genetic diversity and selection in the maize starch pathway. Proc Natl Acad Sci USA 99(20):12959–12962
Wilson LM, Whitt SR, Ibanez AM, Rocheford TR, Goodman MM, Buckler ES (2004) Dissection of maize kernel composition and starch production by candidate gene association. Plant Cell 16(10):2719–2733
Xiong N, Liu J, Yuan J, Zhou XQ, Liu Y, Yu DN et al (2007) Rice. Determination of amylase content. ISO 6647-1 (in Chinese)
Xu J, Liu L, Xu YB, Chen CR, Rong TZ, Ali FH et al (2013) Development and characterization of simple sequence repeat markers providing genome-wide coverage and high resolution in maize. DNA Res 20(5):497–509
Yang D, Guohua W, Ying X, Chang D, Zhao X (2008) Determination of starch in foods. GB/T 5009.9–2008 (in Chinese)
Yao Y, Thompson DB, Guiltinan MJ (2004) Maize starch-branching enzyme isoforms and amylopectin structure. In the absence of starch-branching enzyme IIb, the further absence of starch-branching enzyme Ia leads to increased branching. Plant Physiol 136(3):3515–3523
Zhang XL, Colleoni C, Ratushna V, Sirghie-Colleoni M, James MG, Myers AM (2004) Molecular characterization demonstrates that the Zea mays gene sugary2 code for starch synthase isoform SSIIa. Plant Mol Biol 54(6):865–879
Zhao YJ, Li N, Li B, Li ZX, Xie GN, Zhang JR (2015) Reduced expression of starch branching enzyme IIa and IIb in maize endosperm by RNAi constructs greatly increases the amylose content in kernel with nearly normal morphology. Planta 241:449–461
Acknowledgements
We thank Marty Sachs in the Maize Genetics COOP Stock Center for furnishing seeds of sh1-912A (Stock 912A), and Guoqi Yao (colleague) for furnishing SSR primers. This work was supported by Natural Science Foundation of Shandong Province (ZR2016CB52), National Natural Science Foundation of China (31701443), Major Science and Technology Projects of Shandong Province (2015ZDJS03001), Agricultural Science and Technology Innovation Project of Shandong Academy of Agricultural Sciences (CXGC2017B01), National Key Research and Development Plan (2017YFD0101204) and Research on the foundation and advanced technology of the science and Technology Department of Henan (162300410179).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
10725_2017_309_MOESM1_ESM.tif
Supplementary material 1 Sequence analysis of Sh1 ORFs. The start (ATG) or stop (TAG/TAA) codons are enclosed with yellow and red boxes, respectively. The blue box indicates three nucleotide (CAG to ATA) substitution at 1,993-1,995 bp in the sh1-m coding region. The black box indicates a single base transversion from C to G at 2,183 bp in sh1-912A. (TIF 9163 KB)
10725_2017_309_MOESM2_ESM.tif
Supplementary material 2 Schematic diagram of Sh1 genomic structure. a B73; b Gai-Z58, SNPs and InDels indicate the allelic variation compared with B73, the red asterisk indicates the SNP causing a missense mutation Arg (615) Lys; c sh1-m, SNPs and InDels indicate the allelic variation compared with wild type lines B73 and Gai-Z58, the gray box indicates the thirteenth intron that did not splice out, nucleotides with green and blue font indicate the difference in the exon and intron, respectively, CC with red star indicates the mis-splicing site at the 5′ splice junction of the thirteenth intron and the TAA with red font indicates stop codon; d sh1-912A, SNPs and InDels indicate the allelic variation compared with wild type lines (B73 and Gai-Z58) and sh1-m, nucleotides with blue font and red star indicate the mis-splicing site at the 3′ splice junction of the second intron and the 13-bp nucleotides with green font indicate the sequence at the start of the third exon that spliced out resulted from a new splice site (AG with green font and green star) in this exon. The red asterisk indicates the SNP causing a missense mutation Ala (728) Gly. Black boxes and numbers above and below them indicate exons, their order and their length, respectively. Black short lines indicate introns and numbers above them indicate their length. Green and blue vertical lines indicate SNPs in exon and intron, respectively, red vertical lines indicate InDels. The different length of these vertical lines indicates different SNPs or InDels nearby (TIF 2086 KB)
10725_2017_309_MOESM3_ESM.tif
Supplementary material 3 Multiple amino acid sequence alignment of sh1-m, sh1-912A, B73 and Gai-Z58. The orange line indicates the GT1_Sucrose_synthase region. The red box indicates an Ala to Gly missense mutation in the sh1-912A encoded protein. Dark blue and pink colors indicate 100% and 75% homology levels, respectively (TIF 7600 KB)
10725_2017_309_MOESM4_ESM.tif
Supplementary material 4 Segregation of InDel marker P20 in an F2 population of B73/sh1-m. M DL-2000 marker, P1 B73, F1 B73×sh1-m, P2 sh1-m, 1-4 Homozygous Sh1Sh1 plants in F2, 5-8 Heterozygous Sh1sh1 plants in F2, 9-12 Homozygous sh1sh1 plants in F2 (TIF 1184 KB)
Rights and permissions
About this article
Cite this article
Guan, H., Dong, Y., Liu, C. et al. A splice site mutation in shrunken1-m causes the shrunken 1 mutant phenotype in maize. Plant Growth Regul 83, 429–439 (2017). https://doi.org/10.1007/s10725-017-0309-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0309-9