Abstract
Barley is a crucial cereal crop with a diverse genetic heritage that includes landraces-traditionally cultivated variants that have adapted to varying agro-climatic conditions over centuries. Preserving barley landraces in genebanks is of utmost importance for safeguarding genetic diversity and aiding future breeding endeavors.
This study aims an agro-morphological characterization of a selected collection of two row barley landraces conserved in the Osman Tosun Genebank. A total of 262 barley accessions were assessed in controlled field conditions to analyze various agro-morphological traits. The study unveiled notable disparities in the morphological traits of the genotypes, specifically in flag leaf anthocyanin, flag leaf sheath wax, spike shape, spike density, lemma color, grain color, lodging tendency, days to 50% flowering, plant height, spike length, spikelet quantity, and thousand grain weight, among other critical agronomical characteristics. To scrutinize the morphological disparities, the research utilized principal component analysis and ascertained that nearly 77.22% of the overall variance was accounted for by the first two principal components. The first four principal components accounted for 90.96% of the total variation, thus indicating significant genetic diversity among genotypes. The results of the cluster analysis revealed the presence of diverse genetic resources within the collection, with distinct groups of genotypes exhibiting similar traits being identified. The results of the agro-morphological characterization are a valuable resource for researchers and breeders, aiding them in making informed decisions regarding the management and application of barley landraces in breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Barley (Hordeum vulgare L.) is one of the oldest crop species and stands as a prominent cereal of significant global cultivation, spanning diverse environments that range from temperate to subtropical and from arid to semi-arid conditions. Domesticated barley (Hordeum vulgare ssp. vulgare) originated from its wild ancestor H. spontaneum in the Fertile Crescent of the Middle East approximately 10,000 years ago (Badr et al. 2000; Ullrich 2011; Kaur et al. 2022). Barley grain finds its primary applications in providing animal feed, producing malt products, and serving as a source of human consumption, respectively (Akar et al. 2004). Today, barley is still the world’s fourth largest crop in terms of cultivated area, after wheat, maize and rice in the world. European countries account for nearly half of the global barley cultivation area, where it stands as the second most cultivated crop after wheat. (FAO 2024).
The geographic area known as the Fertile Crescent and Türkiye are known as the most important center of diversity of many plant species including crops such as cereals and edible/forage legumes (Zohary et al. 2012a, b; FAO 2018). Türkiye is also located at the crossroads of numerous gene centers and along the historical migration pathways of significant gene pools (FAO 2018). The regions encompassing especially Southeastern Anatolia and Central Anatolia regions, ranks among the pivotal gene centers for barley (Karagöz 2017; Czembor and Czembor 2021). The different ecological conditions covering biotic and abiotic factors and plant domestication in this region have resulted in enormous plant genetic biodiversity for many species including wheat and barley (Teshome et al. 2001; Newton et al. 2010). Today, barley is the second widely grown cereal in Türkiye after wheat (FAO 2024). The crop has a wide number of different morphological forms such as winter, spring, two-row, six-row, awned, awnless, hooded, naked and hulled, white-grain, colored-grain, dwarf (Ullrich 2011). The commercial forms of barley, malting, feed and food types differ mainly in grain types and properties (Ullrich 2011). Barley displays resilience against challenging environmental factors including soil salinity, drought, cold, elevated altitudes and limited precipitation (Witzel et al. 2009; Ullrich 2011). Barley, classified as a diploid plant, possesses a large genome of 5.1 Gbp with > 80% of repetitive elements, constituting a size twofold that of the human genome, some mutant traits can be used as model traits for cereal crops (Sato 2020).
New cultivars with high yield, quality and adaptation to the changing environmental conditions especially tolerant to drought must continuously be developed to avoid food security problems and access a balanced diet for the growing population. On the other hand, using elite materials in plant breeding has caused cultivars to become genetically uniform, narrowed in variation and more vulnerable to biotic and abiotic stress factors such as drought, high temperature, salinity, diseases and pests (Allel et al. 2016; Kaur et al. 2022). The shrinking and homogeneous barley gene pool is a challenge for the development of new cultivars with superior tolerance to biotic and abiotic stresses (Newton et al. 2010; Marone et al. 2021). Agro-morphological characterization in plants is widely recognized as a crucial first step in evaluating genetic diversity. This process helps to differentiate between materials from different geographical regions, create core collections and identify priority accessions for optimal use in breeding programs. Several reports into barley assemblages across diverse global regions Bhutan (Konishi et al. 1993), Spain (Lasa et al. 2001), Slovakia (Žáková and Benková 2006), Jordan (Shakhatreh et al. 2010), Ethiopia (Abebe et al. 2010; Fantahun et al. 2023), North America (Mikel and Kolb 2008; Allel et al. 2016), Nordic countries (Bengtsson et al. 2017) and India (Manjunatha et al. 2007; Verma and Sarkar 2010; Kaur et al. 2018, 2022; Manju et al. 2019) have underscored the significance of phenotypic characterization. The increasing shrinkage of plant gene pools has directed scientists and breeders to new sources of resistance. The selection pressure have led to the contraction of the genetic resource of contemporary crops (Wulff and Dhugga 2018; Pietrusińska et al. 2018). The gene pool that can be used in plant breeding is divided into three groups according to crossbreeding and affinity (Harlan and de Wet 1971; Von Bothmer et al. 2003). The primary gene pool group consists of modern new (elite) cultivars, old cultivars, landraces, and closely related species. The use of elite as parents has caused the genetically uniform variation to shrink and become more vulnerable to stress factors caused by climate change (Munoz-Amatriain et al. 2014). Next-generation breeding strategies leverage a wider range of genetic resources, including underutilized landraces and wild relatives, to unlock novel traits for adaptation to changing climates and environments (Razzaq et al. 2021).
Identifying the traits of landraces cultivated in the past can contribute to understanding the reasons behind farmers’ selections in the regions where these landraces have adapted. Such investigations can shed light on the selection methods used under diverse environmental conditions, thereby contributing to a more profound comprehension and characterization of locally cultivated landraces (Degu et al. 2023). Moreover, in barley farming, as with all cultivated plants, selecting genotypes that thrive under specific ecological conditions and cultivation techniques is essential for maximizing yield. This is particularly important in Türkiye, where diverse precipitation patterns create a mosaic of ecological landscapes. Consequently, evaluating the adaptability of recently developed cultivars and traditional landraces across the country is a critical need.
Turkish barley landraces serve as a significant genetic resource for adapting to changing environmental conditions because barley has been cultivated in Türkiye since its domestication (i), generally grown in arid conditions with limited and poor resources (ii), and distributed in agroecological regions with very different environmental conditions in terms of climate, soil, hydrography, and topography (iii). Genetic stock investigations in Türkiye were initiated by Professor Dr Osman Tosun in 1938. Between 1938 and 1975, a total of 6,106 barley accessions were collected for the Osman Tosun Gene Bank (OTGB), consisting of 3590 native and 2516 exotic accessions. These accessions constitute a diverse and unique collection, finely attuned to the specific environmental challenges of barley cultivation. Many of these landraces, facing the risk of oblivion and having undergone genetic erosion, are critically preserved within the OTGB for future research and breeding endeavors.
This study focused on exploiting barley germplasm resources by efficiently describing the diversity among 262 two-row barley genotypes, predominantly of Turkish landraces, with morphological data and agronomic performance. This study concentrated on the detailed analysis of eight morphological characteristics, encompassing flowering time, plant height, spike length, and thousand-grain weight. These traits serve as crucial indicators of yield potential and adaptation to various environments. The characterization of these landraces employed multivariate analysis techniques, including principal component analysis, cluster analysis, and correlation analysis. These methodologies enabled the identification of patterns of variation and grouping of landraces based on shared characteristics. This information yields valuable insights into the genetic diversity and potential of the collection. The phenotypic characterization data obtained in this study were utilized to optimize the selection process for a core collection within the OTGB. The aim of this core collection is to capture the maximum genetic diversity and representativeness of Turkish barley landraces, ensuring their efficient conservation and utilization in future research and breeding programs. Additionally, this research establishes a robust foundation for further research on the genetic basis of adaptation in Turkish barley landraces.
Material and method
Experimental conditions
The experiment was conducted at the research field of the Field Crops Central Research Institute, Ministry of Agriculture and Forestry, in Ikizce, Golbasi, Ankara, Türkiye. The experimental site location coordinates are latitude 39°36′33.1″ North and longitude 32°40′40.6″ East, with an elevation of 1050 m above mean sea level. The trial area represents semi-arid agroecological conditions and characterized by an average yearly precipitation of 308 mm. The farm’s soil composition ranges from sandy loam to loamy sand, with a pH level around 7.7 and barley has been widely grown in this area for centuries.
Plant materials and growing techniques
A comprehensive collection of 262 two-row barley genotypes was used in this study. The collection included 209 domestic landraces (DL) collected from different agro-ecological zones in Türkiye and 50 entries from exotic collections (EC) obtained from international gene banks. In particular, the unique location of Türkiye at the crossroads of three different floristic regions has contributed significantly to the rich genetic diversity of the DLs. The EC genotypes, originating from different geographical regions and climates, further broadened the analyzed gene pool and allowed for a more comprehensive assessment. Maps showing the origin of the genotypes throughout world and Türkiye were constructed using MS Excel. To prevent heterogeneity in the landraces, one spike for each landrace was selected from the seed multiplication trials conducted the previous year. Three widely grown two-row barley cultivars, Aydanhanim, Zeynelaga, and Tarm-92, were used as control cultivars. The seed from trashed spike of the genotype were sown by hand in a single row of one mater in length, with a spacing of 40 cm among the rows.
The trial was designed following an Augmented Block Design incorporating a total of 3 blocks. Every block consisted of 117 genotypes, with the final block being of an incomplete nature. The plots were managed fallowing the conventional barley practice except irrigation because of lack of rainfall. The plots were irrigated three times with 100 mm total water, at the three different developmental stage (stem elongation, pre and post-heading stages). Individual accessions were manually harvested and subjected to thresh.
Agro-morphological traits phenotyping
Days to 50% flowering time (DFL), plant height (PH), spike length (SL), row (node) number per spike (RNS), number of grains per spike (NGS), grain weight per spike (SGW), weight of the main stem (SPW), and thousand grain weight (TGW) were quantitatively measured. Growth habit, flag leaf anthocyanin, flag leaf sheath wax, spike shape, spike density, awn arbs, lemma awn, lemma color, grain color, lodging tendency and hull situation were also quantitevely evaluated in accordance with established as indicated in barley (Mahajan et al. 2020; Kaur et al. 2022).
Statistical analysis
The data were analyzed for descriptive statistics (mean, range, SD, and CV) and correlation analysis following the methods of Steel et al. (1980). The principal components analysis (PCA) was carried out with the standardized average values for each of the eight agronomic traits. The dissimilarity between the genotypes was calculated using the Wards method of clustering. The statistical analyses were made with the JMP-13 package program (SAS 2017).
Results and discussion
Origin and distribution of two-row barley genotypes
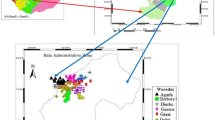
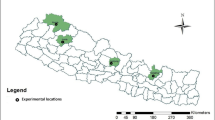
The two-row barley genotypes characterized in this study originated from 14 countries (from across four continents) (Table 1; Fig. 1a). The majority of the working collection consisted of Turkish landraces with 209 genotypes (Table 2). Within the foreign genotypes, 13 originated from the United States, 12 from Ethiopia, and 1 or 2 genotypes were sourced from various other countries. The local landraces were gathered from various locations with complex topography and environmental heterogeneity across Türkiye particularly in areas with widespread cultivation of two-row barley (Fig. 1b). Additionally, within the local landraces, numerous genotypes were from southeastern Türkiye, a region that is part of the Fertile Crescent, where barley was domesticated (Zohary et al. 2012a,b). Two-row barley shows better adaptation to low yielding environments than six-row barley and wheat, with higher productivity and lower yield variability (Newton et al. 2011; Dodig et al. 2020). Therefore, two-row barley is preferred in arid regions such as Central Anatolia with less favorable climates and soil conditions.
Agro-morphological variation of the genotypes
Understanding genetic diversity among different plant genetic materials is facilitated by agro-morphological traits. This study investigated a total of 19 agronomic traits, as previously mentioned. The agro-morphological traits exhibited a wide range of variation among the genotypes. Figure 2 presents the distribution of qualitative traits within two-row barley germplasm. It includes growth habit, flag leaf anthocyanin, flag leaf sheath wax, spike shape, spike density, awn barbs, lemma awn, lemma color, grain color, lodging tendency, and hull type. In Fig. 2, the number of variations within each trait category is shown, allowing for a comparison of the distribution of qualitative traits within OTGB barley germplasm. The landraces can be distinguished easily by their seed color, growth habit, spike length, whether they are naked or hulled, yield, and other significant agronomic traits (Degu et al. 2023). In their study on the characterization of barley landraces, Dondup et al. (2023) reported that accessions exhibited variations in seed colors, including black, blue, and white. They indicated that landraces originating from low altitudes predominantly had white seeds.
The genotypes showed a good fit to the normal distribution. The results of simple descriptive statistics, including the mean value, maximum value, minimum value, standard deviation (SD) and coefficient of variation (CV) of the remaining parameters were given in Table 2. The number of days to 50% flowering in barley genotypes varied from 201 to 232 days, with a standard deviation of 5.64 and a coefficient of variation of 2.64%. Control cultivars Tarm-92, Zeynelaga, and Aydanhanim reached 50% flowering at 206, 213, and 220 days, respectively. The earliest flowering genotype, OTGB955, was a domestic landrace collected from Ankara province, while the latest flowering genotypes, OTGB2803 and OTGB 2807, were from Ethiopia. Early and late maturity may indicate a desire for two-row barley in Central Anatolia. Plant height varied between 45.8 cm (OTGB629) and 113.6 cm (OTGB490) in barley genotypes and 73.0 cm, 89.4 cm, and 97.5 cm in control cultivars. Spike length, varied between 64.4 mm (OTGB1667) and 155.5 mm (OTGB316) in barley germplasm and 110.5 mm, 114.3 mm, and 122.6 mm in control cultivars. Row number of spikelet varied between 7.4 (OTGB1667) and 17.6 (OTGB332) in barley genotypes and 12.8, 14.9, and 17.3 in control cultivars. The number of grains per spike varied between 11.5 (OTGB773) and 34.2 (OTGB417) in barley genotypes and 21.0, 26.6 and 32.3 in control cultivars. Spike grain weight varied between 0.4 g (OTGB758 and OTGB2807) and 2.5 g (OTGB316 and OTGB907) in barley landraces and 1.2 g, 1.7 g, and 2.0 g in control cultivars. Single plant weight varied between 1.2 g (OTGB806) and 9.0 g (OTGB316) in barley germplasm and 2.7 g, 3.8 g, and 5.6 g in control cultivars. Thousand grain weight varied between 31.2 g (OTGB2807) and 76.9 g (OTGB1970) in barley landraces and 59.6 g, 65.9 g, and 61.8 g in control cultivars. OTGB316 had the highest value for the SL, SGW, and SPW parameters. OTGB1667 had the lowest value for SL and RNS and OTGB2807 for SGW and TGW (Table 2). In addition, all the minimum and maximum values of the characteristics examined were formed by landraces and the cultivars used as standards had a value in this range. The results were in line with some previous research. For example, in their study of 367 barley collections, Desta et al. (2024) reported a DFL range of 166–213 days, while Dziurdziak et al. (2021) identified a narrower DFL range (52.0–93.0 days) in a study involving 116 barley genotypes of diverse origins. In a separate investigation encompassing 18 naked barley genotypes, Massman et al. (2022) reported a flowering time ranging from 51 to 62 days. In a similar geographical region, Ergun et al. (2017) found that the heading date of 200 barley landraces ranged between 172 and 194 days. Verma et al. (2021) reported the DFL ranges of 65.0–125 days for 316 barley genotypes sown at different times. These findings illustrate the considerable variability in DFL values, attributed, in part, to disparities in growth conditions, cultivation periods, and genetic variations (Massman et al. 2022; Desta et al. 2024). Alemayehu and Parlevliet (1997) and Derbew et al (2013) found significant variability in plant height and time to maturity among barley genotypes. Desta et al. (2024) measured the spike length of barley landraces in the range of 3–14.4 cm with an average of 6.73 cm. Similarly, Verma et al. (2021) reported spike lengths in barley germplasm in the range of 4.5–13.1 cm. In this study, the spike length of barley germplasm was found to be in the range of 6.4–15.5 cm with an average of 11.1 cm, which is in agreement with previous research. In previous research, diverse ranges of thousand grain weight (TGW) values in barley genotypes have been reported, such as 17.77–67.23 g (Pasam et al. 2012), 30.5–58.8 g (Dziurdziak et al. 2021), 23.2–52.0 g (Mansour et al. 2014), and 3–68 g (Desta et al. 2024). In our study, the TGW values of barley germplasm were measured within the range of 31.2–76.9 g. Our study’s findings on the agro-morphological characteristics of barley germplasm largely corroborate the observations of previous reports. This extensive diversity can be primarily attributed to environmental and genetic factors, particularly the interplay between local stresses and genetic predisposition. For instance, variations in temperature and precipitation can influence plant height and spike length, while specific genes responsible for awn presence or grain color can be differentially expressed depending on breeding selection methods (Hu et al. 2018; Desta et al 2024).
The data were subjected to correlation analysis to determine the relationships between the quantitative agro-morphological traits studied. In the examination of the relationships among the studied characteristics, out of the 28 identified correlations, 4 were deemed insignificant, 2 were significant at the P < 0.05 level, and 22 were found to be highly significant at the P < 0.01 level. Furthermore, among these relationships, 3 were in the negative direction, while the remaining were in the positive direction. Correlation analysis showed that there were significant positive correlations between the traits, with the exception of DFL. DFL was the only quantitative attribute that displayed a negative correlation with PH, SGW, and TGW. Additionally, the relationships between DFL and PH, NGS and SPW were not statistically significant (Table 3). The highest correlations between the analyzed characters were observed between NGS and SGW (r = 0.819, P < 0.01) and between RNS and NGS (r = 0.794, P < 0.01). It was also found that increasing PH had a positive and large effect on SPW in barley germplasm (Table 3). Similar results with respect to maturity time and plant height in barley have also been reported in other research (Kisana et al. 1999; Khan et al. 2021). Similar to our results, positive and significant correlations of spike length with plant height, flowering time and thousand grain weight were reported by Akdeniz et al. (2004) and Desta et al. (2024). Also, Bhutta et al. (2005) reported a positive and significant relationship between plant height and thousand grain weight.
Principal component analysis (PCA) revealed a high level of variation among landraces. The variation studied with principal component analysis showed that two principal components having greater than one eigenvalue contributed 77.22% of the total variance among the eight variable agro-morphological traits (Table 4).
At P < 0.05, spike grain weight, number of grains per spike, plant height, and row number of spikelets were, respectively, the main contributors to the first principle component: grain weight per spike, single plant weight, number of grains per spike, plant height, row number per spike, thousand grain weight, and spike length, which made up 60.70% of the total variation at P < 0.05. However, days to 50% flowering time had the lowest contribution to the observed variations in PC1. On the contrary, the main contributor of the second principal component, responsible for 16.52% of the total variation, was the number of days to 50% flowering time. Thousand grain weight, grain weight per spike, and plant height contributed negatively to the PC2. The first two principal components, which accounted for 77.22% of the total variance, were graphically plotted to display the similarities among genotypes (Fig. 3). Each feature was separately computed to distinguish the genotypes, and the resulting biplot graph also exhibited the variation of genotypes in relation to the eight morphological traits studied. Based on the distribution model of the genotypes on the biplot, it can be inferred that there is a vast amount of genetic variability among the genotypes.
Ebrahim et al. (2015) found that the initial two principal components were responsible for 68.57% of the complete variation and that the complete variation was accounted for by four principal components using agronomic characteristics data of 20 Ethiopian barley genotypes. We revealed that the first two principal components made up 79.59% of the total variation, and the entire variation was explained by eight principal components. It is thought that this difference may be attributed to the small sample size of genotypes analyzed. Hagenblad et al. (2019) morphologically characterized 42 barley landraces originated from the Canary Islands and found that the first two principal components accounted for 34.18% of the total variation. They reported that the first principal component was negatively influenced by plant height and spike length. The discrepancy between their study and our results is probably due to the large number of samples in our study and the high genetic diversity among the accessions. Demissie and Bjørnstad (1996) also used principal component analysis to detect variation in 49 barley populations and reported that the first four principal components accounted for 63% of the total variation. In contrast, this study found that the first four principal components accounted for 90.96% of the total variation. The difference is supposed to be due to the genetic diversity in the current study and the high number of genotypes characterized. Abebe et al. (2010) analyzed the morphological diversity of the 199 Ethiopian barley genotypes. They stated that based on 9 quantitative characteristics, the first 3 principal components accounted for 72.85% of the total variation. These findings are in agreement with our study, since both investigations analyzed a higher number of genotypes than previous research. Similarly, to the current study, Enyew et al. (2019) said that the first two principal components covered 65.93% of the total variation when they classified barley landraces according to their morphological characteristics. Desta et al. (2024) reported that the effect of principal components with eigenvalue greater than 1 on total variation was 74.96%. In our study, we found this value to be 77.22%.
Cluster analysis was performed to establish the genetic relationships among genotypes and determine suitable ones for future breeding programs. Accurate characterization of parental diversity is critical for developing superior cultivars in breeding programs. Cluster analysis based on eight quantitative traits for 262 genotypes was categorized hierarchically with the Wards linkage clustering method into six separate and well-defined clusters (Table 5). According to cluster analysis, genotypes with the least genetic diversity distance between them in terms of morphological traits examined were determined as OTGB514 and OTGB1972. These genotypes are included in Cluster 1. The genotypes exhibiting the highest genetic diversity distance between them were OTGB6 and OTGB261. Cluster 2 was the largest, consisting of 85 genotypes (Table 5; Fig. 4), while Cluster 5 was the smallest, with eight genotypes.
The genotypes in Clusters 1 and 2 were located on the biplot graph with a narrow angle to the vectors, representing the agronomic traits, except the vector represents the number of days to 50% flowering. This result indicated that these genotypes within Clusters 1 and 2 carry more desired agronomic traits than the other genotypes in Clusters 3, 4, and 5, located at a wider angle to the vectors on the biplot graph.
One of the main purposes of this study is to determine the genotypes with early flowering characters. The mean of the number of days to 50% flowering in Cluster 1 was 210.63 days, while 216.42 days in Cluster 2. According to this result, among the clusters, Cluster 1 consists of genotypes with early flowering characteristics as well as desirable agronomic traits. The genotypes grouped in Cluster 1 attract attention because of these characteristics. On the other hand, the genotypes within Cluster 1 still showed a wide variation in terms of morphological characteristics, including days to 50% flowering (204–219 days), plant height (70.0–113.6 cm), spike length (100.8–131.8 mm), row number of spikelet (11.8–15.6), number of grains per spike (20.2–28.8), spike grain weight (1.4–2.1 g), single plant weight (3.1–5.3 g), and thousand grain weight (60.1–76.9 g) (Table 6).
In order to evaluate and identify the genotypes with superior traits, the data in cluster one were subjected to re-clustering. Classifying genotypes into phenotypically similar clusters of different groups is useful for selecting parents for hybridization. The cluster analysis grouped the barley genotypes with greater phenotypic similarity; however, not all accessions from the same or adjacent location were included in the cluster. The cultivars used in this study, for instance, were all grouped into separate clusters. This finding is consistent with Ahmad et al. (2008) and Abebe et al. (2010). They reported that clustering of accessions based on morphological characteristics did not reveal any distinct geographic grouping patterns, as accessions from the same or adjacent regions were found in different clusters. In terms of late maturity, shorter plant height, and low thousand-grain weight, the clusters with the least participation in the classification consisted of 16 genotypes. Mekonnon et al. (2015) found similar results when clustering barley genotypes based on their morphological traits, as those with late maturity, shorter plant height, and low thousand grain weight were in the cluster with the least number of participants.
The genotypes within Cluster 1 were further divided into six new groups by reapplying cluster analysis (Table 7; Fig. 5). Upon evaluation of the new clusters (sub-clusters), it was found that C1-4 exhibited the desired values for the examined traits. C1-4 comprises seven genotypes, all of which are landraces (Table 8). C1-1 has the highest number of accessions and mainly consists of early maturing genotypes. The cluster with the lowest number of accessions was C1-6, and it consisted of genotypes with short plant height, long spike length, and high thousand grain weight (Table 7). According to re-cluster analysis, genotypes with the least genetic diversity distance between them in terms of morphological traits examined were determined as OTGB437 and OTGB1972. These genotypes are included in C1-2. The genotypes with the furthest genetic distance between them were OTGB6 and OTGB589.
The highest contributor to genetic diversity is the variation between accessions (Dido et al. 2022). This suggests that even a single accession can be a resource for breeding material due to the genetic variability that exists within each accession (Dziurdziak et al. 2021; Abebe et al. 2023).
Conclusion
This study provides new data on the characterization of 262 two-row barley genotypes conserved in the Osman Tosun Gene Bank, mainly collected from diverse agroecological environments of Anatolia. We, for the first time, evaluated the genetic diversity and phenotypic traits of two-row barley landraces and unveiled notable disparities in the agro-morphological traits of the genotypes.
This characterization has led to the identification of promising landraces that demonstrate traits of considerable agronomic importance. Among the promising landraces, for example, compared to those in control cultivars, OTGB 955 reached heading 12 days earlier, OTGB 316 had 34% longer spikes and 53% heavier grain weight per spike, OTGB 417 and OTGB 32 genotypes had 12% more spikelets per spike and 28% more grains per spike, respectively. The thousand grain weight of the OTGB 1970 genotype was determined to be 23% higher than the average of the controls. This characteristic highlights OTGB 1970 as a large-grained and high-yielding genotype.
The study also underscored the significance of Turkish landraces, which function as valuable genetic resources for increased adaptability to changing climate conditions and end-user demands. These landraces, collected from a wide range of environments in terms of climate, soil, hydrography, and topography, contain valuable genetic diversity that can be used to breed new barley cultivars that are more resilient to these challenges. Moreover, creating a flexible core collection and trait-spesific germplasm will easily access diversity and facilitate the integration of the existing barley gene pool into enhancement programs.
Ongoing research aims to delve deeper into the barley accessions, employing a combination of phenotyping and genotyping techniques to identify advantageous genes. Future endeavors will include comprehensive genome-wide association research, with the core set cultivated under diverse agro-ecological conditions. This approach is expected to unveil a more nuanced understanding of the genetic factors, alleles, and markers contributing to the resilience and adaptability of these barley cultivars. The insights gained from such research endeavors are crucial for developing targeted breeding strategies that address the dynamic challenges posed by changing environmental conditions and evolving agricultural areas.
Referencess
Abebe TD, Bauer AM, Léon J (2010) Morphological diversity of Ethiopian barleys (Hordeum vulgare L.) in relation to geographic regions and altitudes. Hereditas 147(4):154–164
Abebe TD, Abate A, Leon J (2023) Genetic diversity within landraces of barley (Hordeum vulgare L.) and its implications on germplasm collection and utilization. Genet Resour Crop Evol 70:1–14
Ahmad Z, Ajmal SU, Munir M, Zubair M, Masood MS (2008) Genetic diversity for morpho-genetic traits in barley germplasm. Pak J Bot 40(3):1217–1224
Akar T, Avci M, Dusunceli F (2004) Barley: post-harvest operations. Food and Agriculture Organization (FAO) of the United Nations, The Central Research Institute for Field Crops, p 64
Akdeniz H, Keskin B, Yılmaz I, Oral E (2004) A research on yield and yield components of some barley cultivars. J Agric Sci 14:119–125
Alemayehu F, Parlevliet JE (1997) Variation between and within Ethiopian barley landraces. Euphytica 94:183–189
Allel D, Ben-Amar A, Badri M, Abdelly C (2016) Salt tolerance in barley originating from harsh environment of North Africa. Aust J Crop Sci 10:438–451
Badr A, Rabey HE, Effgen S, Ibrahim HH, Pozzi C, Rohde W, Salamini F (2000) On the origin and domestication history of barley (Hordeum vulgare). Mol Biol Evol 17(4):499–510
Bengtsson T, Manninen O, Jahoor A, Orabi J (2017) Genetic diversity, population structure and linkage disequilibrium in Nordic spring barley (Hordeum vulgare L. subsp. vulgare). Genet Resour Crop Evol 64:2021–2033
Bhutta WM, Barley T, Ibrahim M (2005) Path-coefficient analysis of some quantitative characters in husked barley. Caderno de Pesquisa Sér Biologia Santa Cruz do Sul 17(1):65–70
Czembor JH, Czembor E (2021) Mlo resistance to powdery mildew (Blumeria graminis f. sp. hordei) in barley landraces collected in Yemen. Agronomy 11(8):1582–82
Degu HD, Tehelku TF, Kalousova M, Sato K (2023) Genetic diversity and population structure of barley landraces from Southern Ethiopia’s Gumer district: utilization for breeding and conservation. PLoS ONE 18(1):e0279737. https://doi.org/10.1371/journal.pone.0279737
Demissie A, Bjørnstad Å (1996) Phenotypic diversity of Ethiopian barleys in relation to geographical regions, altitudinal range, and agro-ecological zones: as an aid to germplasm collection and conservation strategy. Hereditas 124(1):17–29
Derbew S, Elias U, Hussein M (2013) Genetic variability in Barley (Hordeum vulgare (L.)) landrace collections from Southern Ethiopia. Int J Sci Res 12(2):125–131
Desta KT, Choi YM, Yoon H, Lee S, Yi J, Jeon YA, Shin MJ (2024) Comprehensive characterization of global Barley (Hordeum vulgare L.) collection using agronomic traits, β-glucan level, phenolic content, and antioxidant activities. Plants 13:169
Dido AA, Singh BJK, Ermias A, Krishna MSR, Dawit D, Kassahun T (2022) Genetic diversity, population structure and relationship of Ethiopian barley (Hordeum vulgare L.) landraces as reveled by simple sequence repeat (SSR) markers. J Genet. https://doi.org/10.1007/s12041-021-01346-7
Dodig D, Kandić V, Zorić M, Nikolić-Đorić E, Nikolić A, Tančić Živanov S, Perović D (2020) Response of kernel growth of barley genotypes with different row type to climatic factors before and after inflection point of grain filling. Field Crop Res 255:107864
Dondup D, Yang Y, Xu D, Namgyal L, Wang Z, Shen X, Dorji T, Kyi N, Drolma L, Gao L, Ga Z, Sang Z, Ga Z, Mu W, Zhuoma P, Taba X, Jiao G, Liao W, Tang Y, Zeng X, Luobu Z, Wu Y, Wang C, Zhang J, Qi Z, Guo W, Guo G (2023) Genome diversity and highland-adaptative variation in Tibet barley landrace population of China. Front Plant Sci 14:1189642. https://doi.org/10.3389/fpls.2023.1189642
Dziurdziak J, Gryziak G, Groszyk J, Podyma W, Boczkowska M (2021) DArTseq genotypic and phenotypic diversity of barley landraces originating from diferent countries. Agronomy 11(11):2330. https://doi.org/10.3390/agronomy11112330
Ebrahim S, Shiferaw E, Hailu F (2015) Evaluation of genetic diversity in barley (Hordeum vulgare L.) from Wollo high land areas using agromorphological traits and hordein. Afr J Biotechnol 14:1886–1896
Enyew M, Dejene T, Lakew B, Worede F (2019) Clustering and principal component analysis of barley (Hordeum volugare L.) landraces for major morphological traits from North Western Ethiopia. Int J Agric Sci Food Technol 5:058–063
Ergun N, Aydogan S, Sayım I, Karakaya A, Celik Oguz A (2017) Investigation of yield and some agronomic Traits on barley (Hordeum vulgare L.) landraces. Tarla Bitkileri Merkez Araştırma Enstitüsü Dergisi 26:180–189
Fantahun B, Woldesemayate T, Fadda C, Gebrehawaryat Y, Pe E, Dell Acqua M (2023) Multivariate analysis in the dissection of phenotypic variation of Ethiopian cultivated barley (Hordeum vulgare ssp. Vulgare L.) genotypes. Cogent Food Agric 9(1):2157104
FAO (2018) Biodiversity of Turkey: contribution of genetic resources to sustainable agriculture and food systems. Ankara, p 222. Licence: CC BY-NC-SA 3.0 IGO
FAO (2024) Plant production statistics. FAO, Rome. http://www.fao.org/faostat/en/#data/QC
Hagenblad J, Leino MW, Hernández Afonso G, Afonso Morales D (2019) Morphological and genetic characterization of barley (Hordeum vulgare L.) landraces in the Canary Islands. Genet Resour Crop Evol 66:465–480
Harlan JR, de Wet JM (1971) Toward a rational classification of cultivated plants. Taxon 20(4):509–517
Hu X, Zuo J, Wang J, Liu L, Sun G, Li C, Sun D (2018) Multi-locus genome-wide association studies for 14 main agronomic traits in barley. Front Plant Sci 9:1683–9
Karagöz A (2017) Agro-morphological variation among an ancient world barley collection. J Agric Sci 23(4):444–452
Kaur V, Kumari J, Manju Jacob SR, Panwar BS (2018) Genetic diversity of indigenous and exotic germplasm of barley (Hordeum vulgare L.) and identification of trait specific superior accessions. Wheat Barley Res 10:190–197. https://doi.org/10.25174/2249-4065/2018/83620
Kaur V, Aravind J, Jacob SR, Kumari J, Panwar BS, Pal N, Rana C, Pandy A, Kumar A (2022) Phenotypic characterization, genetic diversity assessment in 6778 accessions of barley (Hordeum vulgare L. ssp. vulgare) germplasm conserved in National Genebank of India and development of a core set. Front Plant Sci 13:771920
Khan A, Ihsan M, Nisar M, Hazrat A, Ali M, Ul-Haq R, Faisal S (2021) Evaluation of genetic diversity in barley landraces through agro-morphological and biochemical characterization. Sarhad Agric, p 37
Kisana NS, Ahmed Z, Tahir M, Mujahid MY, Majid IAA, Zahid Mustafa S (1999) Variability and relationship between morpho-phenological traits and grain yield in winter and facultative barley under stress environments. Pak J Biol Sci 2:767–771
Konishi T, Yano Y, Fukushima Y (1993) Genetic variation in barley landraces from Bhutan. Genet Resour Crop Evol 40:33–38
Lasa JM, Igartua E, Ciudad FJ, Codesal P, Garcia EV, Gracia MP, Medina B, Romagosa I, Molina-Cano JL, Montoya JL (2001) Morphological and agronomical diversity patterns in the Spanish barley correlation-. Hereditas 135:217–225. https://doi.org/10.1111/j.1601-5223.2001.00217.x
Mahajan G, Hickey L, Chauhan BS (2020) Response of barley genotypes to weed interference in Australia. Agronomy 10(1):99
Manju KV, Sharma K, Jacob SR (2019) Assessment of genetic diversity in cultivated and wild species germplasm of barley based on morphoagronomical and root architecture traits. Ind J Plant Genet Resour 32:360–367. https://doi.org/10.5958/0976-1926.2019.00039.1
Manjunatha T, Bisht IS, Bhat KV, Singh BP (2007) Genetic diversity in barley (Hordeum vulgare L. ssp. vulgare) landraces from Uttaranchal Himalaya of India. Genet Resour Crop Evol 54:55–65
Mansour E, Casas AM, Gracia MP, Molina-Cano JL, Moralejo M, Cattivelli L, Thomas WTB, Igartua E (2014) Quantitative trait loci for agronomic traits in an elite barley population for mediterranean conditions. Mol Breed 33:249–265
Marone D, Russo MA, Mores A, Ficco DB, Laidò G, Mastrangelo AM, Borrelli GM (2021) Importance of landraces in cereal breeding for stress tolerance. Plants 10(7):1267
Massman C, Meints B, Hernandez J, Kunze K, Hayes PM, Sorrells ME, Smith KP, Dawson JC, Gutierrez L (2022) Genetic characterization of agronomic traits and grain threshability for organic naked barley in the northern United States. Crop Sci 62(2):690–703
Mekonnon B, Lakew B, Dessalegn T (2015) Morphological diversity and association of traits in Ethiopian food barley (Hordeum vulgare L.) landraces in relation to regions of origin and altitudes. J Plant Breed Crop Sci 7:44–54
Mikel MA, Kolb FL (2008) Genetic diversity of contemporary North American barley. Crop Sci 48(4):1399–1407
Munoz-Amatriaín M, Cuesta-Marcos A, Endelman JB, Comadran J, Bonman JM, Bockelman HE, Chao S, Russell J, Waugh R, Haues PM, Muehlbauer GJ (2014) The USDA barley core collection: genetic diversity, population structure, and potential for genome-wide association studies. PLoS ONE 9(4):e94688
Newton AC, Akar T, Baresel JP, Bebeli PJ, Bettencourt E, Bladenopoulos KV, Patto MV (2010) Cereal landraces for sustainable agriculture. Sustain Agric 2:147–186
Newton AC, Flavell AJ, George TS et al (2011) Crops that feed the world 4. Barley: a resilient crop? Strengths and weaknesses in the context of food security. Food Secur 3:141–178
Pasam RK, Sharma R, Malosetti M, van Eeuwijk FA, Haseneyer G, Kilian B, Graner A (2012) Genome-wide association studies for agronomical traits in a world wide spring Barley collection. BMC Plant Biol 12:16. https://doi.org/10.1186/1471-2229-12-16
Pietrusińska A, Żurek M, Pıechota U, Słowackı P, Smolıńska K (2018) Searching for diseases resistance sources in old cultivars, landraces and wild relatives of cereals. A review. Agronomy 6:66
Razzaq A, Kaur P, Akhter N, Wani SH, Saleem F (2021) Next-generation breeding strategies for climate-ready crops. Front Plant Sci 12:620420. https://doi.org/10.3389/fpls.2021.620420
SAS Institute Inc (2017) JMP® Statistical Discovery Software, version 13.2.0. SAS Institute Inc., Cary
Sato K (2020) History and future perspectives of barley genomics. DNA Res. 27(4):dsaa023. https://doi.org/10.1093/dnares/dsaa023
Shakhatreh Y, Haddad N, Alrababah M, Grando S, Ceccarelli S (2010) Phenotypic diversity in wild barley (Hordeum vulgare L. ssp. spontaneum (C. Koch) Thell.) accessions collected in Jordan. Genet Resour Crop Evol 57:131–146
Steel RG, Torrie JH, Dickey DA (1980) Principles and procedures of statistics: a biometrical approach. McGraw-Hill, New York, p 481
Teshome A, Brown AHD, Hodgkin T (2001) Diversity in landraces of cereal and legume crops. In: Janick J (ed) Plant breeding reviews. Wiley, New York, pp 221–261
Ullrich SE (2011) Barley: production, improvement, and uses, vol 12. Wiley, New York
Verma RPS, Sarkar B (2010) Diversity for malting quality in barley (Hordeum vulgare) varieties released in India. Indian J Agric Sci 80(6):493
Verma S, Yashveer S, Rehman S, Gyawali S, Kumar Y, Chao S, Sarker A, Verma RPS (2021) Genetic and Agro-morphological diversity in global barley (Hordeum vulgare L.) collection at ICARDA. Genet Resour Crop Evol 68:1315–1330
Von Bothmer R, Sato K, Komatsuda T, Yasuda S, Fischbeck G (2003) The domestication of cultivated barley. In: von Bothmer R, van Hintum T, Knüpffer H, Sato K (eds) Diversity in barley. Elsevier, Amsterdam, pp 9–27
Witzel K, Weidner A, Surabhi GK, Börner A, Mock HP (2009) Salt stress-induced alterations in the root proteome of barley genotypes with contrasting response towards salinity. J Exp Bot 60(12):3545–3557
Wulff BB, Dhugga KS (2018) Wheat the cereal abandoned by GM. Science 361(6401):451–452
Žáková M, Benková M (2006) Characterization of spring barley accessions based on multivariate analysis. Commun Biometry Crop Sci 1(2):124–134
Zohary D, Weiss E, Hopf M (2012a) Current state of the art. In: Zohary D, Hopf M, Weiss E (eds) Domestication of plants in the old world: the origin and spread of domesticated plants in Southwest Asia, Europe, and the Mediterranean Basin, 4th edn. Oxford University Press, Oxford, pp 1–8. https://doi.org/10.1093/acprof:osobl/9780199549061.001.0001
Zohary D, Weiss E, Hopf M (2012b) Cereals. In: Zohary D, Hopf M, Weiss E (eds) Domestication of plants in the old world: the origin and spread of domesticated plants in Southwest Asia, Europe, and the Mediterranean Basin, 4th edn. Oxford University Press, Oxford, pp 20–74
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK). This study was financially supported by Tubitak. TUBITAK-1001, Project number: 119O092.
Author information
Authors and Affiliations
Contributions
G.A. and B.B. conceived of the presented idea. G.A. developed the theory and performed the computations. B.B. and S.A. verified the analytical methods. G.A. encouraged B.B., M.B., H.A.A.A. T.E. and N.E. to investigate characterize barley germplasm and supervised the findings of this work. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have not disclosed any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Benlioglu, B., Bilir, M., Akdogan, G. et al. Phenotypic characterization of two-row barley (Hordeum vulgare L. ssp. vulgare) germplasm conserved in Osman Tosun Genebank of Türkiye by multivariate analysis model. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-02032-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-02032-0