Abstract
Intermediate wheatgrass (IWG, Thinopyrum intermedium [Host] Barkworth & D. R. Dewey) has been developed as a perennial grain crop for human consumption along with providing environmental benefits and ecosystem services. Grain and products derived from IWG cultivars improved for food production have been marketed under the registered trademark, Kernza. Development of IWG as a perennial grain crop began in 1980s with a phenotypic recurrent selection program as the Rodale Institute (RI) and the Big Flats Plant Material Center (BFPMC) used IWG plant introductions (PI) from the National Plant Germplasm System (NPGS) to improve populations of IWG. Initial selections were provided to The Land Institute (TLI) where they were subsequently improved for grain production, yet the identity of the founder material of improved, food-grade IWG has not been publicly documented. Recently recovered original documents have been used to reconstruct the early breeding program to identify the most likely 20 PIs that form the founders of modern food-grade IWG. Molecular data using genotyping-by-sequencing in current elite breeding material, and remnant seed and plant material from the initial RI selections have provided supporting evidence for the historical records. The genetic origin for food-grade IWG is focused between the Black Sea and Caspian Sea in the Stavropol region of Russia, with smaller contributions likely from collections as distant as Kazakhstan in the east to Turkey in the west. This work connects the flow of germplasm and utility of NPGS PIs to present day IWG grain cultivars being developed in multiple breeding programs around the world.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Inspired by Wes Jackson’s ideas (1980) for a diverse, herbaceous perennial polyculture, the Rodale Research Center (now known as Rodale Institute [RI]) in the early 1980s evaluated approximately 300 different species for their utility in perennial agriculture similar to natural ecosystems. Following evaluation of more than 100 different grass species, intermediate wheatgrass (IWG, Thinopyrum intermedium (Host) Barkworth & D.R. Dewey) was selected in 1985 as having the highest potential to be developed into a perennial grain crop based on its plant structure, seed production characteristics, perennialism, food use potential (Wagoner 1990a), and the fact that it is related to important Triticeae grain crops (Wagoner 1990b) likely indicating a good nutritional profile and lack of anti-nutritive compounds. Prior to selection for domestication, IWG had a history within the United States for erosion control, forage production (Asay and Jensen 1996), and utilization as a tertiary gene pool for annual wheat (Triticum aestivum) improvement (Li and Wang 2009; Pototskaya et al. 2022).
Since the identification of IWG as a target of domestication, a dedicated and expanding research effort has been conducted to bring the idea of perennial grains to fruition. Research on IWG for perennial grain use has focused on all facets of plant breeding, agronomic practices, and potential consumer utilization. Some of this research has shown the environmental benefits posited by perennial crops such as reduced nitrate leaching (Culman et al. 2013; Jungers et al. 2019; Huddell et al. 2023), net carbon accumulation (de Oliveira et al. 2018), increased soil particulate organic matter (van der Pol et al. 2022), and reduced surface runoff of particulate matter (poultry litter) (Katuwal et al. 2023) compared to annual crops. Supplementing positive environmental benefits of perennial grain, agronomic research has led to a better understanding of plant development (Duchene et al. 2021), management (Jungers et al. 2017, 2022), and harvesting (Heineck et al. 2022; Tautges et al. 2023) of IWG as a perennial grain and forage crop. A comprehensive grower guide that details many planting, management, and harvesting methods used for IWG was released in 2023 with particular emphasis on the differences between perennial IWG management and annual cereals (Tautges et al. 2023). Finally, many food applications have been evaluated including flour quality and processing methods (Marti et al. 2015, 2016; Zhang et al. 2015; Rahardjo et al. 2018; Banjade et al. 2019) as well as research looking at malting applications (Marcus and Fox 2023). While current IWG yields are estimated to be about 25–30% of annual wheat yields (Cassman and Connor 2022; Bajgain et al. 2023), IWG grain quality has higher protein, bran, and essential amino acid percentages than annual wheat (Becker et al. 1991). Additionally, research has shown that equal mixtures of IWG and wheat flour can maintain baking quality while increasing nutritional content (Marti et al. 2015).
A central tenet to the environmental, agronomic, and end use research has been that plant breeding will play a pivotal role in developing higher yielding cultivars. Bolstered by early breeding successes in IWG that showed up to 77% increase in grain yield in two cycles of selection (DeHaan et al. 2014, TLI Cycle 0–2, see Discussion) and molecular breeding methods (Moose and Mumm 2008), plant breeding programs are working to develop better performing cultivars with a consequential moment in 2019 with the release of ‘MN-Clearwater’ as the world’s first IWG developed for food consumption (Bajgain et al. 2020b). Kernza refers to the grain that is marketed from IWG cultivars that have been developed for human consumption of the grain, such as MN-Clearwater, and currently there are a number of commercial Kernza products available to consumers. We use “improved IWG” to signify Thinopyrum intermedium germplasm bred for human consumption that could produce Kernza grain to clearly distinguish it from seeds of forage-type IWG. Following the early breeding efforts of RI, there are now breeding programs in Canada, Ukraine, Sweden, and the United States actively breeding for improved IWG. In an effort to speed genetic gain, some of these programs have actively utilized genomic selection (GS) (Zhang et al. 2016; Crain et al. 2021a) to reduce breeding cycle time. Implementation of GS has required molecular methods like genotyping-by-sequencing (GBS) (Elshire et al. 2011; Poland et al. 2012) that have also provided data needed to dissect genetic architecture through biparental quantitative trait loci (QTL) mapping (Zhang et al. 2017; Larson et al. 2019) and association mapping (Bajgain et al. 2019; Altendorf et al. 2021a, b; Bajgain and Anderson 2021; Crain et al. 2022) Taken together, breeding programs have leveraged these newer tools and methodologies to both enhance the agronomic performance of IWG as well as identify causal variants of traits to both assist in breeding and increase our scientific understanding of IWG genetics.
The basic evaluation and development of improved IWG since the 1980s was previously reported (Cox et al. 2002; Zhang et al. 2016) as well as the development of breeding programs and selection methods in programs established after 2001 (DeHaan et al. 2018; Bajgain et al. 2023). As many of the breeding programs have a combination of pedigree or molecular data along with laboratory records, most improved IWG lineages can be traced back to their programs’ beginning material which subsequently traces back to RI selections. For reference, IWG was first introduced into the United States in 1932 (Musil 1948). It is also known that RI evaluated many of the publicly available plant introduction (PI) accessions from the United States Department of Agriculture (USDA) National Plant Germplasm System (NPGS) gene bank (Wagoner 1990a), so there is a clear connection between improved IWG (Kernza) cultivars and PIs although specific PIs have never been published. As most of the IWG PI accessions in the gene bank have been characterized phenotypically and genotypically (Crain et al. 2023), and include at least some source documentation, there is an opportunity to fill in the missing link between the geographic origin of the PI founders and improved IWG cultivars. We strive to identify the likely 20 PI accessions used to initiate the first cycle of the RI breeding program (Polycross-1) along with the potential origins of additional germplasm included in the second cycle (Polycross-2). Selections from Polycross-2 formed the basis of subsequent IWG breeding programs (DeHaan et al. 2018; Bajgain et al. 2023). Using molecular data, remnant seed and plant material, and recently recovered field and laboratory documents, our objective is to solidify the reported early breeding lineages (Zhang et al. 2016; Bajgain et al. 2023) to the NPGS PI material used to develop improved, food-grade IWG.
Materials and methods
Historical records and experiments
In an effort to trace the origins of improved IWG for Kernza grain production, we consulted a number of historical records. Listed in Supplementary File 1, these records included field and laboratory notebooks, program reports, principal investigators, and presentations that helped document IWG evaluation and breeding activities conducted by RI. These records, detailed in the results section, provide an overview of the first three cycles of phenotypic recurrent selection by RI in collaboration with Big Flats Plant Materials Center (BFPMC), which we refer to as Polycross-1, Polycross-2, and Polycross-3.
Genomic analysis
Plant material
To complement historical records, we leveraged existing genotype data from The Land Institute (TLI) IWG breeding program (DeHaan et al. 2018; Crain et al. 2021a, b), the NPGS IWG PI collection (Crain et al. 2023), 306 genotyped genets from remnant seed of RI Polycross-2, and remnant stems, leaves, and inflorescence material from the 14 selected Polycross-2 parents (collectively referred to as “Remnant Rodale”) for a total of 10,456 unique genotyped genets. As each individual genet will have its own unique genetic makeup due to IWG’s outcrossing nature (Jensen et al. 1990), we use the term genet (Zhang et al. 2016) to refer to individual plants of an IWG accession. Using the terminology of Zhang et al. (2016) a single genet can be cloned into multiple ramets (plants) having the same genetic makeup. Within the TLI breeding program, we sampled several germplasm pools including TLI Cycle-6 (n = 3072), TLI Cycle-12 (n = 4032), and Breeding Parents from TLI Cycle-6–13 (n = 803, approximately 100 each cycle) as they represent a direct link from TLI Cycle-6 to TLI Cycle-12. Cycle-6 from TLI represents six generations (genetic recombination and selection) between germplasm that was shared with TLI from RI (Polycross-2 and Selection Nursery 2, see Results). Likewise, another six generations beyond TLI Cycle-6 were represented by TLI Cycle-12. We used TLI Cycle-12 and the breeding parents to increase the number of source assignments to NPGS PI accessions. As this is a mostly closed breeding program (Bajgain et al. 2023), TLI Cycle-6 & 12, and the breeding parents also represent a check that PI assignments were not vastly different between germplasm pools, a potential issue that would likely indicate incorrect model choice. A total of 2329 unique genets representing 371 NPGS accessions (approximately 6 genets per accession) were genotyped previously (Crain et al. 2023) and received from NPGS web request 23,159 (October 26, 2017). Known off-types and PI accessions that did not appear to morphologically resemble IWG were removed from further analysis (Crain et al. 2023), leaving a total of 338 NPGS PI accessions available for further analysis.
Genomic profiling and bioinformatics
DNA was extracted using a range of products including MagMAX (ThermoFisher Scientific, Waltham, MA), BioSprint (QIAGEN, Venlo, The Netherlands), and CTAB (for older tissues from Polycross-2). Across all genets, we used a two-enzyme restriction digest genotyping-by-sequencing (GBS) protocol following the methods of Poland et al. (2012). Multiplexed libraries ranging from 96 to 384 plexing were sequenced on Illumina sequencing platforms. As the samples represent a range of time and locations, sequencing platforms and output increased as technology improved. To call single nucleotide polymorphism (SNP) markers, we used the TASSEL-GBSv2 pipeline (Glaubitz et al. 2014) using the IWG genome V3.1. We filtered the marker data for strictly biallelic SNPs, a minor allele frequency greater than 0.05, and for SNPs to be called in a minimum of 30% of the genets (up to 70% missing data). Individual genets with more than 95% missing data were removed from further analysis. We required a minimum read depth of four to call a homozygote, otherwise the SNP call was set to missing if there were less than four identical reads. Heterozygous calls were allowed with a read depth of two contrasting reads. For the NPGS PI accessions where multiple genets were genotyped from each accession (Crain et al. 2023), we called SNPs on single genets as well as combining genets of a single accession to create a composite genomic profile. After filtering, a total of 25,773 SNPs and 8242 genets were used for further analysis.
We created a pairwise genetic relationship matrix using the stats package (R Core Team 2022), where smaller distance values represented more SNPs in common between individuals and larger values indicated fewer SNPs in common between compared individuals. From this matrix, the first and second most closely related PI accessions were determined for each improved IWG genet from the TLI Cycle-6, TLI Cycle-12, breeding parents, and Remnant Rodale evaluations (n = 7904 of which 97% of these were from advanced generations of the TLI breeding materials). Of the 139 accessions evaluated by RI and available for use in Polycross-1, a subset of 114 have been genotyped. Using the genetic relationship matrix to ascertain population membership (Manel et al. 2005), we first assigned the 7904 improved IWG genets to a potential 114 NPGS source populations (PI accessions). To identify other potential founders that did not have any known genotypic or historic context, namely RI accession #31, we repeated the population assignment by expanding from 114 to 338 possible germplasm sources that represented the majority of the IWG NPGS collection (Crain et al. 2023). We excluded PIs that were collected (not donated to NPGS) ex situ after 1990 as they would have been unavailable to RI, for a total 331 PI accessions. Because written records were quite specific that 20 accessions were used to form Polycross-1 (although accession names were not provided), we chose the 20 NPGS PI accessions that accounted for the most founder assignments of the improved IWG genets (or total number of PI accessions if less than 20) as the most likely genomic progenitors of current improved IWG germplasm for Kernza grain production. To evaluate our ability to discriminate between relationships within and among the NPGS accessions, we tested genetic assignments of 1,997 individuals representing 338 NPGS accessions used in this study. The ggplot2 (Wickham 2016) and VennDiagram (Chen 2022) R packages were used for data visualization.
Results
Historical records
Original material evaluation
Over the course of nearly two decades, RI was actively involved in all aspects of perennial grain development including species evaluation and selection, germplasm enhancements, agronomic practices, and product utilization. Beginning in 1983, an evaluation nursery (hereafter referred to as RI Herbary) of nearly 300 perennial species was established at RI to evaluate plant accessions for their potential use in perennial herbaceous polyculture. Each accession was grown in a single, one-meter-long row separated from other accessions by mowed grass alleyways. Once IWG was selected as the most promising candidate for domestication in 1985, RI obtained more accessions of IWG from a variety of sources including the USDA NPGS gene bank in Pullman, WA, researchers in the United States, seed companies, and from foreign seed banks. Each evaluation row consisted of ten genets from the same PI accession. By 1987, there were 43 unique IWG accessions in the RI Herbary and each accession was evaluated for plant structure and vigor, as well as seed production characteristics including ease of threshing, synchrony of maturity, and shatter resistance. In addition, 100-seed weight, as a yield component trait, was determined for each accession by averaging the weight of 3 randomly selected sets of 100 seeds dried to 12% moisture. From 1988 to 1993, a larger number of IWG accessions were more intensively evaluated annually as described above with the inclusion of two more yield components including seed set rating (Trupp and Slinkard 1965) and seed yield (g) per 10 heads. The seed set rating trait was calculated as the weight (g) of clean, naked seed from 10 seed heads divided by the weight (g) of the unthreshed seed heads. Naked seed was obtained by using a fabricated threshing board and manually processing samples (Supplementary Fig. 1). Evaluations at RI emphasized seed size (as measured by 100-seed weight) and seed head fertility (measured by seed set rating) as these measurements were both highly heritable in IWG and a direct correlation had been established between seed yield and seed head fertility (Slinkard 1965).
Polycross-1
In the fall of 1987, another 99 IWG accessions consisting of 10 genets each were added to the RI Herbary for a total of 139 unique IWG accessions completing the panel of potential candidates for the first polycross (Polycross-1) breeding cycle. Of the evaluated material, 116 accessions were PIs from the USDA NPGS gene bank in Pullman, WA. Selection of parents from the 139 accessions evaluated both in 1988 and 1989 was based on favorable seed yield characteristics including 100-seed weight, seed set rating, and seed yield per 10 heads. This multi-year evaluation included both a drought (1988) and above average precipitation (1989). Twenty accessions which represented favorable phenotypic traits were selected. Segments of each selected accession row were dug from the RI Herbary in November 1989. Each segment was divided into three subsegments, which were placed into pots and vernalized during December 1989. It is possible that subsegments from each accession were clonally derived from one plant or from multiple genets as the rhizomes of individual genets (10 per row) would be intermingled and indistinguishable from each other within these selected accession rows. Thus, we deduce that the minimum number of selected genets was 20 (one per accession) and the maximum number of selected genets was 60 (three per accession) for Polycross-1. In January 1990, the pots were placed in the RI greenhouse for intercrossing to produce Syn-0 seeds. Pots were rearranged frequently to ensure that pollination could occur between all accessions. Syn-0 seeds, produced by this intercrossing, were bulked and then planted into market packs, one seed per cell in June 1990. A total of 360 seedlings (genets) were transplanted in September 1990 at the USDA Natural Resource Conservation Service (NRCS) BFPMC, Big Flats, NY, establishing Selection Nursery-1. The collaboration with BFPMC resulted from both an institutional reorganization of RI and the expertise of BFPMC in selection and varietal development of undomesticated species.
Polycross-2
From 1991 to 1994, Selection Nursery-1 was evaluated jointly by RI and BFPMC for the same traits that were emphasized in initial RI Herbary accession evaluations. The 360 genets grown from Syn-0 seed produced in Polycross-1 were maintained as individual genets with bare ground around each plant. Additionally, seed heads were removed from each plant at maturity to prevent field contamination by shattered seed. Of the 360 genets, 11 genets were chosen to be included in Polycross-2.
While evaluations of Selection Nursery-1 were occurring, RI was actively continuing evaluations and expanding their IWG germplasm collection. In the fall 1988, another 111 IWG accessions were added to the RI Herbary, planted in rows (as described in Original Material Evaluations). Of these 111 accessions, 102 were comprised of half-sib families planted from seeds collected from each of 102 open-pollinated plants selected by RI researchers in 1987 from John Berdahl’s breeding nurseries at the USDA Northern Great Plains Research Laboratory (NGPRL) in Mandan, ND. From 1989 to 1994, all of these new NGPRL-derived RI accessions were evaluated in RI Herbary rows, each containing four half-sib plants. Along with the 11 genets from Selection Nursery-1, three RI accessions originating from NGPRL were selected in 1994 for inclusion in Polycross-2. From each of the three selected accession rows, a segment of plant material was dug up with each selection originating from at least one genet and up to three genets due to the possible intermingling of rhizomes among the half-sib progenies within each RI Herbary accession row.
Polycross-2 consisted of 11 Selection Nursery-1 genets that were cloned to produce 3 ramets of each individual genet and at least one targeted genet (up to a maximum of 3 potential genets) from each of the subsegments (clones) of three NGPRL-derived RI accessions. The 3 clones of each of 14 selections were planted in the field at BFPMC in a pattern to allow maximum cross pollination among all selections in the crossing block. From a breeding point of view, we deduce that Polycross-2 contained at least 14 distinct parental genotypes and a maximum of 20 distinct parental genotypes. Intermating of clones in the field occurred in 1996 producing Syn-1 seeds which would include genetic material from the original 20 selected accessions used in Polycross-1 and the introduced genetic material from the 3 RI Herbary accessions that were originally collected from NGPRL. In fall 1997, 400 individual genets grown from Syn-1 seeds were planted at BFPMC, Big Flats, NY to form Selection Nursery-2. Selection Nursery-2 was evaluated jointly by RI and BFPMC from 1998 to 2001 and only by BFPMC from 2002 to 2005. Seed and materials from Polycross-2 and Selection Nursery-2 were distributed to several scientists and institutions (Fig. 1) which formed the basis of the TLI breeding program in the early 2000’s. Subsequent work by BFPMC selected a total of 16 genets from Selection Nursery-2 to form Polycross-3. Each genet was cloned four times and planted into a crossing block in 2006. Even though BFPMC evaluated Polycross-3 between 2007 and 2016, seed produced from Polycross-3 was not used in successive breeding efforts by TLI.
Recreating polycross-1 from historical records
Although the identity of the original 20 accessions selected for use in Polycross-1 would have been recorded at the time, this information is no longer available. To reconstruct the most likely selected accessions, recently recovered field and laboratory records have been reviewed to determine the most likely accessions used in Polycross-1 (Supplementary File 1). These records provide details of yield component traits in 1988 and 1989 and specifically state that 20 accessions with favorable yield characteristics were selected for Polycross-1; however, the identities of these accessions are now obscure. There was evidence of genotype-by-environment interaction as a list of 22 accessions that had high yield in 1988 (drought year) were often not the highest yielding in 1989 (wet year). Additionally, the average values combined across the two years for the three yield components of the 20 selected accessions were given as 100-seed weight (0.495 g), seed set rating (45%), and yield per 10 heads (2.5g). These averages provided a target value in searching the records to identify the most likely combination of selected accessions. Using this information, we have identified the most likely accessions for these 20 parents (Table 1). Recovered records of the 139 accessions evaluated in 1988 and 1989 show 100-seed weight and seed set rating for both years but yield per 10 head data are limited to 1989 and only a few accessions from 1988. Therefore, calculated values for yield per 10 heads do not perfectly align with the target values. The estimated yield per 10 heads for the most likely selected accessions is 2.67 g compared to a target value of 2.5 g. As there were genotype-by-environment interactions, inclusion of missing data would most likely lower the calculated value closer to the target. Based on historical records, 19 of the 20 selected accessions were from NPGS. The one accession not from NPGS was given to RI in 1982 by Wes Jackson and TLI, but no other information is available about this accession. In terms of geographical origin, USDA records indicate that 13 accessions were from Russia with the majority of these collected between the Black Sea and Caspian Sea (Caspian-Pontic Steppe, Stavropol and Svetlograd regions). Passport data from NPGS suggest that three of the accessions were cultivated when collected, raising the possibility that they may all come from the same cultivar “Rostov(sky) 31” (Table 1).
Genomic analysis
Genomic profiling of plant material
We compared genomic data from several germplasm pools including TLI Cycle-6, TLI Cycle-12, breeding parents, Remnant Rodale, and NPGS PIs. As genotyping was conducted on different sequencing platforms and times, we evaluated the number of reads and number of SNPs called per genet. The germplasm pool affected the number of SNPs with an overall average of 21,698 SNPs called per genet. Single genets from the NPGS PIs had the fewest called SNPs with an average of 9867 SNPs followed by TLI Cycle-6 with an average of 19,597 SNPs per genet while TLI Cycle-12 have the highest average of 23,043 SNPs per genet. When the NPGS PI accessions were pooled (approximately six genets per accession), the average number of SNPs called per accession increased to 23,086. Along with number of SNPs called, we also examined the read depth of each SNP to verify that there was no apparent bias due to sequencing differences in the data sets. Cycle-6 from TLI had the lowest read depth per SNP with an average of 2.6 reads per SNP. Cycle-12 from TLI had a mean read depth of 7.6, the breeding parents were higher at 8.4, and both Remnant Rodale and NPGS PI accessions had a mean read depth of 11. When individual genets of the NPGS PIs are considered, the average read depth per genet was 1.9 (approximately 6 genets were combined per accession). These values most likely reflect older sequencing technology in TLI Cycle-6 along with different program objectives that targeted higher read depths in the breeding parents, Remnant Rodale and NPGS accessions to ensure a greater amount of data compared to the TLI breeding program that balances practical objectives with cost.
Validation test of genet assignments
To test the sensitivity of our assignments using the relationship matrix, we used the 1997 genets that represented 338 unique NPGS PI IWG accessions. For each individual genet, we tested whether it was most related to other individuals of the same accession based strictly on DNA genotypes. Across 1997 iterations, only one genet was assigned to the incorrect PI accession (99% accuracy).
Likely founders of improved IWG from genomic analysis
Genomic comparison to RI evaluated material
Leveraging the genetic resources that have been developed for implementing genomics assisted breeding in IWG, we investigated the most likely founders of improved IWG using empirical molecular genetic data. Based on historical knowledge of 114 genotyped accessions evaluated by RI, a total of 30 NPGS PI accessions were identified by genomic analysis as the most likely founders across 7904 improved IWG genets (Fig. 2). Of these potential matches, there was a very skewed distribution where 10 NPGS PI accessions accounted for over 98% of the assigned individuals, and 11 NPGS PI accessions had 10 or fewer descendants (Fig. 2). In assessing model fit, we looked at the average increase in distance between the top two NPGS PI candidates (Supplementary Table 1). Across all tested genets, there was an average distance increase of 0.45 between the first and second most related NPGS accession compared to an average distance increase of 0.21 between any other comparisons. The 14 Polycross-2 parents (11 of which were only one generation away from original selected PI accessions) had an average distance increase of 3.8 between the first and second most related NPGS accessions. TLI Cycle-6 and 12 had much smaller average increases of 0.53 and 0.37 between the first and second most likely NPGS accessions. This result is likely based on breeding progress and genetic recombination across cycles obscuring the original founder haplotypes as well as providing evidence to support these assignments.
Bar graph of the number of times each National Plant Germplasm System plant introduction (NPGS PI) intermediate wheatgrass (IWG) accessions were assigned as a founder of 7904 improved IWG genets (from Remnant Rodale, TLI Cycle-6 & 12 and Breeding Parents) for Kernza production. Assignment was based on the 114 genotyped PI accessions originally evaluated by RI
When considering NPGS PI assignment by test subject (Remnant Rodale, TLI Cycle-6, Breeding Parents, TLI Cycle-12) the assignments of the newer Breeding Parents and TLI Cycle-12 test subjects were a subset of assignments identified in the older TLI Cycle-6 test subjects. This result might be expected as both the Breeding Parents and TLI Cycle-12 were developed from a mostly closed population from TLI Cycle-6. Thus, further analyses will be focused on the older and more diverse Remnant Rodale and TLI Cycle-6 test subjects.
When comparing the 20 most likely selections used in RI Polycross-1 inferred from historical records to the genetic assignments made using molecular data from Remnant Rodale and TLI Cycle-6 test subjects, seven NPGS PIs were identified in all data sources (Fig. 3 and Tables 1 and 2). These seven NPGS PIs (PI 273732, PI 286118, PI 314054, PI 315353, PI 316122, PI 440004, PI 440015) were the closest NPGS PI accession for over 40% of the tested genets indicating they are quite likely founders of current IWG germplasm being improved for Kernza grain production. Ten NPGS PI accessions that were identified as possible selections based on historical records for Polycross-1 did not show any descendants based on genetic assignments; however, these accessions were often part of a series of adjacent PI accessions (i.e. PI440004-18).
National Plant Germplasm System plant introduction (NPGS PI) source assignments (maximum 20) for Rodale Institute (RI) Polycross-1 historical records, Remnant Rodale DNA genotypes, and The Land Institute (TLI) Cycle-6 DNA genotypes. Remnant Rodale is 227 unique genets resulting from RI Polycross-2 which includes genetic contributions from RI Polycross-1 and additional RI selections originating from the Northern Great Plains Research Laboratory breeding program. TLI Cycle-6 is 3072 unique genets from TLI IWG breeding program
Genomic comparison to the broader IWG NPGS collection
As not all the initial RI accessions were genotyped (i.e. RI Accession 31 and 3 accessions selected for inclusion in Polycross-2 which would be included as founders of improved IWG), we expanded the search of potential founders to the broader 331 NPGS IWG accessions that were collected prior to 1990. This broader search identified 47 possible NPGS sources of improved IWG germplasm. However, only 10 potential NPGS accessions accounted for 95% of all assignments as the most likely founders of the tested germplasm, and just 20 NPGS PI accessions were considered founders to more than 10 IWG genets. From written records, an upper limit of 20 NPGS PI founders in Polycross-1 was placed on all analysis (historical records, assignment to NPGS PI evaluated by RI, and assignment to broader NPGS accessions), and a total of 38 NPGS PI accessions were identified as being the potential founders of improved IWG that is currently being used for Kernza grain production (Tables 1, 2, 3).
Repeating the founder assignment to a broader set of 331 NPGS PI accessions, also allowed us to evaluate the strength of assignment by comparing assignments to the 114 RI accessions to the expanded set. Of the 7904 genets assigned to a source, over 53% were assigned to the same source. One NPGS accession, PI 502351, accounted for another 37% of assignments. According to NPGS Germplasm Resources Information Network passport data, this accession was cultivated material and from Elista, Russia. This region, between the Black Sea and Caspian Sea, is also the origin of other accessions identified as sources of improved IWG. If this material was cultivated, it would likely have superior agronomic traits, like other cultivated material in Table 1, and share large genetic similarities leading to the high rate of assignment to this PI accession.
Inferred geographic origin of improved IWG
Finally, we investigated the most likely geographic origin of germplasm used to develop improved IWG cultivars used for Kernza grain production using NPGS Germpasm Resources Information Network passport data. A total of 27 NPGS PI accessions identified in the recreated RI Polycross-1 and molecular data had reliable location data from which 20 were from Russia, two each from Turkey and Kazakhstan, and one each from Afghanistan, Iran, and Uzbekistan. Seven other accessions either had missing data or location data that was deemed insufficient. For example, PI 286118 was listed with a country of origin of Denmark, yet the sample came from a botanical garden. Given this information, it is very likely this accession had been collected elsewhere before arriving in Denmark, but records of its natural origin are not available to our knowledge. The overlapping NPGS PIs identified as possible sources of improved IWG from our historical and genetic analyses, originated from the Pontic-Caspian steppe between the Black Sea and the Caspian Sea (Fig. 4) likely indicating a primary geographic origin of improved IWG germplasm currently used for Kernza grain production.
Geographic location of 27 National Plant Germplasm System intermediate wheatgrass (IWG) plant introduction (PI) accessions that are inferred to be the founders of improved IWG germplasm used for current Kernza grain production. Orange circles represent common PIs inferred through historical and molecular methods. Green diamonds are PI accessions inferred through historic records only, and purple triangles are PI inferred through molecular methods only
Discussion
Rodale research activities
In the 1980s, RI initiated an effort to identify potential perennial grains and develop them into a crop. In many ways, RI developed some of the first empirical blueprints of pipeline perennial grain development that has been expanded on by later research (Wagoner 1990b; Cox et al. 2002; DeHaan et al. 2016, 2023). Additionally, the whole systems approach (breeding, agronomy, economics, end use) used by RI is still used today. Some examples of this integrated approach are the Kernza coordinated agriculture project (KernzaCAP, https://kernza.org/kernzacap/), the reflective plant breeding paradigm (Runck et al. 2014), and organized efforts through TLI (https://landintitute.org).
Using two years of phenotypic data RI launched a phenotypic recurrent selection program that would result in the improved germplasm used in modern Kernza grain production. Even though RI would have preferred to have had more years of evaluation for a perennial crop with an anticipated five-year crop lifetime, breeding efforts were balanced between time and institutional support. Seed shared with TLI came from Polycross-2 and Selection Nursery-2, which have also been referred to as BFPMC Cycle 1 and 2 (Fig. 1), served as the basis of improved IWG (DeHaan et al. 2018; Bajgain et al. 2023). Much like Cox et al. suggested in 2002 that new molecular tools would aid in perennial grain development, the advent of genomic selection (Meuwissen et al. 2001) and subsequent next generation sequencing methods have allowed plant (IWG) breeding programs to harness molecular technology for improved plant breeding (Zhang et al. 2016; Bajgain et al. 2020a; Crain et al. 2021a, b). Current breeding programs harnessing genomic selection can make yearly breeding selection with numerous years of phenotypic data informing the models (Crain et al. 2021a, b). Emerging technologies like speed breeding (Watson et al. 2018) have the potential to further reduce breeding cycle time and increase the rate of genetic gain. Much like the initial RI selections that included extreme drought and wet years, genomic selection models can utilize data across multiples years and cycles (Crain et al. 2020) incorporating a range of climatic conditions into the selection information.
Linkage between PI accessions and Kernza Grain production
Overall, the genomic data provide strong support to clarify the partial historical records, which indicate that improved IWG that is currently used for Kernza grain production is primarily descended from a limited number of NPGS PI accessions mainly originating between the Black Sea and Caspian Sea. Using the genomic data, we inferred the most likely source of the 20 accessions in RI Polycross-1; however, most of the material we sampled was after Polycross-2 which had another severe bottleneck of only 14 accessions comprised of at least 14 and not more than 20 genets, Fig. 1. The bottleneck in Polycross-2 included additional genetic material of unknown origin from the three RI selections that had originally been collected from NGPRL.
Potential confounding factors
While the historical records and genomic data had large areas of overlap, there was not complete consistency between the methods. This should not be surprising given the number of accessions evaluated, time span of the programs, and even the different institutions and staff that have been involved. Within the TLI breeding program, most of the germplasm was obtained directly from RI (Polycross-2 and Selection Nursery-2) and was initially referred to as TLI Cycle-0 (DeHaan et al. 2014). However, there was evaluation and incorporation of other NPGS PI accessions before TLI Cycle-6 (Bajgain et al. 2023). Within our analysis, assignment of single NPGS PI genets to their respective source NPGS PI accession was very high demonstrating our ability to discriminate NPGS accessions by DNA genotype. Even though assignments of improved IWG genets to NPGS PI accessions appears plausible, it does not appear to be as precise as evidenced by up to 47 NPGS PI founders for improved IWG genets. Even though this number is higher than reported records, the skewed distribution suggests a number complementary with the recorded data as the source for improved IWG. Both the nature and structure of the data could be influencing these results. Perhaps the most obvious reason is that intermating and genetic recombination has blended the original NPGS accessions in such a way that it is more difficult to match the improved IWG genets to any one specific NPGS accession. As we tried to identify the most likely founders in Polycross-1, most of the material genotyped had at least two cycles of genetic recombination (Remnant Rodale) and up to 14 cycles of genetic recombination (TLI Cycle-12) from the original founders as well as contributions from additional genetic material included in Polycross-2 from the NGPRL breeding program. As each recombination occurred there would have been reshuffling and breaking of the original haplotypes, and additionally there would have been selection pressure applied to obtain agronomically superior plants.
In previous analysis of IWG, we have noted that more than 70% of the genetic variation observed is within accessions (Crain et al. 2023), suggesting that random sampling of seed could influence our observations. In our analysis, we have assumed that field records and that sampling were completely accurate, with similar assumptions for genetic profiling. If errors occurred, after the passage of time it would be almost impossible to identify or correct. Along with potential field errors, the IWG accessions evaluated by RI mainly came from the NPGS system after having been collected in the 1970s and earlier. Until the 1990s there was no isolation protocol in accession regenerations (Johnson et al. 1996) and IWG pollen dispersal has been documented to mainly occur within 10 m (Bajgain et al. 2022) indicating that there could have been the possibility of admixture among the NPGS accessions before they were received for phenotypic evaluation by RI in the 1980s and later genomic profiling. However, for many of the PIs used in Polycross-1, records from NPGS indicate that there had been only one seed increase since collection and distribution to RI therefore RI received NPGS PI accessions as close to the original collection as possible.
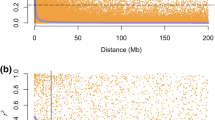
Notably, both historic records and genomic data often indicate many NPGS accessions come from a series of adjacent collections. For example, the PI 4400xx series formed a large portion of the selected 20 accessions for Polycross-1. As many of the accessions were collected during the same expeditions and most likely in chronological or spatial order, it is not surprising that these accessions would often be similar to each other. Work by Crain et al. (2023) showed a strong correlation between geographic distance and genetic distance within the IWG NPGS collections, suggesting that accessions collected near each other are more likely to share alleles. Even with potential germplasm, field, and laboratory errors, the data clearly indicate a small subset of IWG NPGS accessions that are the most likely primary founders of improved IWG germplasm that is currently used for Kernza grain production.
Conclusion
This work identifies direct linkages between NPGS accessions and improved IWG germplasm that is currently grown for Kernza grain production, showcasing how plant germplasm collections and repositories can be utilized for breeding and the development of new crops. By identifying the most likely genetic origins of food-grade IWG, plant breeders can continue to utilize the NPGS accessions for additional genetic diversity for enhanced crop production as well as better understand the domestication effort behind this grain. Confirming the small number of genets (14–20) within the RI breeding program that have been used to improve IWG helps indicate likely effective population size within this new crop and could guide future genetic research. Finally, identifying the primary geographic origin between the Black and Caspian Seas provides areas for future germplasm collections, in-situ conservation initiatives, and identifying accessions that can be used to broaden the current germplasm pool for improved IWG.
Data availability
DNA sequence data generated for this study was uploaded to the NCBI Sequence Read Archive (SRA) (https://www.ncbi.nlm.nih.gov/bioproject) as BioProject PRJNA968038. Data from BioProjects PRJNA609325 and PRJNA866171 were also used for analysis. All data scripts and data files have been placed in the Zenodo digital repository doi at https://doi.org/10.5281/zenodo.10126848.
Abbreviations
- ARS:
-
Agriculture Research Service
- BFPMC:
-
Big Flats Plant Materials Center, United States Department of Agriculture, Natural Resources Conservation Service, Big Flats, NY
- GBS:
-
Genotyping-by-sequencing
- GS:
-
Genomic selection
- IWG:
-
Intermediate wheatgrass
- NGPRL:
-
Northern Great Plains Research Laboratory, United States Department of Agriculture, Agriculture Research Service, Mandan, ND
- NPGS:
-
National plant germplasm system
- PI:
-
Plant introduction
- RI:
-
Rodale Institute
- SNP:
-
Single nucleotide polymorphism
- TLI:
-
The Land Institute
References
Altendorf KR, DeHaan LR, Larson SR, Anderson JA (2021a) QTL for seed shattering and threshability in intermediate wheatgrass align closely with well-studied orthologs from wheat, barley, and rice. Plant Genome. https://doi.org/10.1002/tpg2.20145
Altendorf KR, Larson SR, DeHaan LR et al (2021b) Nested association mapping reveals the genetic architecture of spike emergence and anthesis timing in intermediate wheatgrass. G3 Genes Genom Genet 11:1–14. https://doi.org/10.1093/g3journal/jkab025
Asay K, Jensen K (1996) Wheatgrasses. Cool Season Forage Grasses 34:691–724
Bajgain P, Anderson JA (2021) Multi-allelic haplotype-based association analysis identifies genomic regions controlling domestication traits in intermediate wheatgrass. Agriculture 11:667. https://doi.org/10.3390/agriculture11070667
Bajgain P, Zhang X, Anderson JA (2019) Genome-wide association study of yield component traits in intermediate wheatgrass and implications in genomic selection and breeding. G3 Genes Genom Genet 9:2429–2439. https://doi.org/10.1534/g3.119.400073
Bajgain P, Zhang X, Anderson JA (2020a) Dominance and G×E interaction effects improve genomic prediction and genetic gain in intermediate wheatgrass (Thinopyrum intermedium). Plant Genome 13:1–13. https://doi.org/10.1002/tpg2.20012
Bajgain P, Zhang X, Jungers JM et al (2020b) ‘MN-Clearwater’, the first food-grade intermediate wheatgrass (Kernza perennial grain) cultivar. J Plant Regist 14:288–297. https://doi.org/10.1002/plr2.20042
Bajgain P, Brandvain Y, Anderson JA (2022) Influence of pollen dispersal and mating pattern in domestication of intermediate wheatgrass, a novel perennial food crop. Front Plant Sci 13:871130. https://doi.org/10.3389/fpls.2022.871130
Bajgain P, Crain JL, Cattani DJ et al (2023) Breeding intermediate wheatgrass for grain production. In: Plant breeding reviews. Wiley, pp 119–217
Banjade JD, Gajadeera C, Tyl CE et al (2019) Evaluation of dough conditioners and bran refinement on functional properties of intermediate wheatgrass (Thinopyrum intermedium). J Cereal Sci 86:26–32. https://doi.org/10.1016/j.jcs.2019.01.001
Becker R, Wagoner P, Hanners GD, Saunders RM (1991) Compositional, nutritional and functional evaluation of intermediate wheatgrass (Thinopyrum intermedium). J Food Process Preserv 15:63–77
Cassman KG, Connor DJ (2022) Progress towards perennial grains for prairies and plains. Outlook Agric 51:32–38. https://doi.org/10.1177/00307270211073153
Chen H (2022) VennDiagram: generate high-resolution Venn and Euler plots. R package version 1.7.3. https://CRAN.R-project.org/package=VennDiagram
Cox TS, Bender M, Picone C et al (2002) Breeding perennial grain crops. Crit Rev Plant Sci 21:59–91. https://doi.org/10.1080/0735-260291044188
Crain J, Bajgain P, Anderson J et al (2020) Enhancing crop domestication through genomic selection, a case study of intermediate wheatgrass. Front Plant Sci 11:319. https://doi.org/10.3389/fpls.2020.00319
Crain J, DeHaan L, Poland J (2021a) Genomic prediction enables rapid selection of high-performing genets in an intermediate wheatgrass breeding program. Plant Genome. https://doi.org/10.1002/tpg2.20080
Crain J, Haghighattalab A, DeHaan L, Poland J (2021b) Development of whole-genome prediction models to increase the rate of genetic gain in intermediate wheatgrass (Thinopyrum intermedium) breeding. Plant Genome. https://doi.org/10.1002/tpg2.20089
Crain J, Larson S, Dorn K et al (2022) Genetic architecture and QTL selection response for Kernza perennial grain domestication traits. Theor Appl Genet. https://doi.org/10.1007/s00122-022-04148-2
Crain J, Larson S, Sthapit S et al (2023) Genomic insights into the NPGS intermediate wheatgrass germplasm collection. Crop Sci 2:20944. https://doi.org/10.1002/csc2.20944
Culman SW, Snapp SS, Ollenburger M et al (2013) Soil and water quality rapidly responds to the perennial grain Kernza wheatgrass. Agron J 105:735–744. https://doi.org/10.2134/agronj2012.0273
de Oliveira G, Brunsell NA, Sutherlin CE et al (2018) Energy, water and carbon exchange over a perennial Kernza wheatgrass crop. Agric for Meteorol 249:120–137. https://doi.org/10.1016/j.agrformet.2017.11.022
DeHaan LR, Van Tassel DL, Anderson JA et al (2016) A pipeline strategy for grain crop domestication. Crop Sci 56:917–930. https://doi.org/10.2135/cropsci2015.06.0356
DeHaan L, Christians M, Crain J, Poland J (2018) Development and evolution of an intermediate wheatgrass domestication program. Sustainability 10:1499. https://doi.org/10.3390/su10051499
DeHaan LR, Anderson JA, Bajgain P et al (2023) Discussion: prioritize perennial grain development for sustainable food production and environmental benefits. Sci Total Environ 895:164975
DeHaan LR, Wang S, Larson SR et al (2014) Current efforts to develop perennial wheat and domesticate Thinopyrum intermedium as a perennial grain. In: Perennial crops for food security proceedings of the FAO expert workshop, pp 72–89
Duchene O, Dumont B, Cattani DJ et al (2021) Process-based analysis of Thinopyrum intermedium phenological development highlights the importance of dual induction for reproductive growth and agronomic performance. Agric for Meteorol 301–302:108341. https://doi.org/10.1016/j.agrformet.2021.108341
Elshire RJ, Glaubitz JC, Sun Q et al (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6:e19379. https://doi.org/10.1371/journal.pone.0019379
Glaubitz JC, Casstevens TM, Lu F et al (2014) TASSEL-GBS: a high capacity genotyping by sequencing analysis pipeline. PLoS ONE 9:e90346. https://doi.org/10.1371/journal.pone.0090346
Heineck GC, Schlautman B, Law EP et al (2022) Intermediate wheatgrass seed size and moisture dynamics inform grain harvest timing. Crop Sci 62:410–424. https://doi.org/10.1002/csc2.20662
Huddell A, Ernfors M, Crews T et al (2023) Nitrate leaching losses and the fate of 15N fertilizer in perennial intermediate wheatgrass and annual wheat—a field study. Sci Total Environ 857:159255. https://doi.org/10.1016/j.scitotenv.2022.159255
Jackson W (1980) New roots for agriculture. U of Nebraska Press
Jensen KB, Dewey DR, Zhang YF (1990) Mode of pollination of perennial species of the Triticeae in relation to genomically defined genera. Can J Plant Sci 70:215–225. https://doi.org/10.4141/cjps90-024
Johnson R, Bradley V, Knowles R (1996) Genetic contamination by windborne pollen in germplasm-regeneration plots of smooth bromegrass. Bulletin des Ressources Phytogenetiques (IPGRI/FAO); Noticiario de Recursos Fitogeneticos (IPGRI/FAO), pp 30–34
Jungers JM, DeHaan LR, Betts KJ et al (2017) Intermediate wheatgrass grain and forage yield responses to nitrogen fertilization. Agron J 109:462–472. https://doi.org/10.2134/agronj2016.07.0438
Jungers JM, DeHaan LH, Mulla DJ et al (2019) Reduced nitrate leaching in a perennial grain crop compared to maize in the Upper Midwest, USA. Agric Ecosyst Environ 272:63–73. https://doi.org/10.1016/j.agee.2018.11.007
Jungers JM, Schiffner S, Sheaffer C et al (2022) Effects of seeding date on grain and biomass yield of intermediate wheatgrass. Agron J 114:2342–2351. https://doi.org/10.1002/agj2.21083
Katuwal S, Ashworth AJ, Moore PA, Owens PR (2023) Characterization of nutrient runoff from perennial and annual forages following broiler litter application. J Environ Qual 52:88–99. https://doi.org/10.1002/jeq2.20425
Larson S, DeHaan L, Poland J et al (2019) Genome mapping of quantitative trait loci (QTL) controlling domestication traits of intermediate wheatgrass (Thinopyrum intermedium). Theor Appl Genet 132:2325–2351. https://doi.org/10.1007/s00122-019-03357-6
Li H, Wang X (2009) Thinopyrum ponticum and Th. intermedium: the promising source of resistance to fungal and viral diseases of wheat. J Genet Genom 36:557–565. https://doi.org/10.1016/S1673-8527(08)60147-2
Manel S, Gaggiotti O, Waples R (2005) Assignment methods: matching biological questions with appropriate techniques. Trends Ecol Evol 20:136–142. https://doi.org/10.1016/j.tree.2004.12.004
Marcus A, Fox G (2023) Malting and wort production potential of the novel grain Kernza (Thinopyrum intermedium). J Am Soc Brew Chem 81:308–318. https://doi.org/10.1080/03610470.2022.2026662
Marti A, Qiu X, Schoenfuss TC, Seetharaman K (2015) Characteristics of perennial wheatgrass (Thinopyrum intermedium) and refined wheat flour blends: impact on rheological properties. Cereal Chem J 92:434–440. https://doi.org/10.1094/CCHEM-01-15-0017-R
Marti A, Bock JE, Pagani MA et al (2016) Structural characterization of proteins in wheat flour doughs enriched with intermediate wheatgrass (Thinopyrum intermedium) flour. Food Chem 194:994–1002. https://doi.org/10.1016/j.foodchem.2015.08.082
Meuwissen THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829. https://doi.org/10.1093/genetics/157.4.1819
Moose SP, Mumm RH (2008) Molecular plant breeding as the foundation for 21st century crop improvement. Plant Physiol 147:969–977. https://doi.org/10.1104/pp.108.118232
Musil AF (1948) The taxanomic status of Agropyron intermedium as indicated by a comparative study of its seed with that of A. Trichophorum and A. elongatum. Proc Assoc off Seed Anal 38:79–83
Poland JA, Brown PJ, Sorrells ME, Jannink JL (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS ONE 7:e32253. https://doi.org/10.1371/journal.pone.0032253
Pototskaya IV, Shamanin VP, Aydarov AN, Morgounov AI (2022) The use of wheatgrass (Thinopyrum intermedium) in breeding. Vestn VOGiS 26:413–421. https://doi.org/10.18699/VJGB-22-51
R Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rahardjo CP, Gajadeera CS, Simsek S et al (2018) Chemical characterization, functionality, and baking quality of intermediate wheatgrass (Thinopyrum intermedium). J Cereal Sci 83:266–274. https://doi.org/10.1016/j.jcs.2018.09.002
Runck BC, Kantar MB, Jordan NR et al (2014) The reflective plant breeding paradigm: a robust system of germplasm development to support strategic diversification of agroecosystems. Crop Sci 54:1939–1948. https://doi.org/10.2135/cropsci2014.03.0195
Slinkard A (1965) Fertility in intermediate wheatgrass Agropyron intermedium (Host) Beauv. Crop Sci 5:363–365
Tautges N, Detjens A, Jungers JM (2023) Kernza grower guide. Retrieved from the University of Minnesota Digital Conservancy. https://hdl.handle.net/11299/255778
Trupp C, Slinkard A (1965) Seedset rating as a measure of fertility in grasses. Crop Sci 5:599–600
van der Pol LK, Nester B, Schlautman B et al (2022) Perennial grain Kernza® fields have higher particulate organic carbon at depth than annual grain fields. Can J Soil Sci 102:1005–1009. https://doi.org/10.1139/cjss-2022-0026
Wagoner P (1990a) Perennial grain new use for intermediate wheatgrass. J Soil Water Conserv 45:81–82
Wagoner P (1990b) Perennial grain development: past efforts and potential for the future. Crit Rev Plant Sci 9:381–408. https://doi.org/10.1080/07352689009382298
Watson A, Ghosh S, Williams MJ et al (2018) Speed breeding is a powerful tool to accelerate crop research and breeding. Nat Plants 4:23–29. https://doi.org/10.1038/s41477-017-0083-8
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, New York
Zhang X, Ohm J-B, Haring S et al (2015) Towards the understanding of end-use quality in intermediate wheatgrass (Thinopyrum intermedium): high-molecular-weight glutenin subunits, protein polymerization, and mixing characteristics. J Cereal Sci 66:81–88. https://doi.org/10.1016/j.jcs.2015.10.008
Zhang X, Sallam A, Gao L et al (2016) Establishment and optimization of genomic selection to accelerate the domestication and improvement of intermediate wheatgrass. Plant Genome. https://doi.org/10.3835/plantgenome2015.07.0059
Zhang X, Larson SR, Gao L et al (2017) Uncovering the genetic architecture of seed weight and size in intermediate wheatgrass through linkage and association mapping. Plant Genome. https://doi.org/10.3835/plantgenome2017.03.0022
Acknowledgements
We thank the Department of Energy Joint Genome Institute, the Perennial Agriculture Project and The Land Institute for prepublication access to the Thinopyrum intermedium Genome sequence. We want to acknowledge the valuable contributions of Martin van der Grinten, retired NRCS BFPMC, for his collaboration during the RI breeding program, his continued efforts beyond RI involvement, and for his recent assistance in locating and sharing historical documents and germplasm used to document this research. Shuangye Wu and Ethan Faryna provided valuable laboratory assistance. The high-performance computing for this project was performed on the Beocat Research Cluster at Kansas State University, which was funded in part by NSF grants CNS1006860, EPS-1006860, EPS-0919443, ACI-1440548, CHE1726332, and NIH P20GM113109. Keygene N.V. owns patents and patent applications protecting its Sequence Based Genotyping technologies. Contribution no. 24-044-J from the Kansas Agricultural Experiment Station.
Funding
This project was partially supported by the Perennial Agriculture Project and the KernzaCAP, an AFRI Sustainable Agricultural Systems Coordinated Agricultural Project (SAS-CAP) grant no. 2020-68012-31934 from the USDA National Institute of Food and Agriculture.
Author information
Authors and Affiliations
Contributions
P.W. Conceptualization; Investigation; Formal analysis; Methodology; Writing-original draft; Writing-review & editing. J.C. Formal analysis; Methodology; Writing-original draft; Writing-review & editing. S.L. Conceptualization; Formal analysis; Writing-review & editing. L.D. Conceptualization; Investigation; Writing-review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest or competing interest in the completion of this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10722_2024_1952_MOESM1_ESM.docx
Primary source references that were consulted in reconstructing the Rodale Institute intermediate wheatgrass breeding program (DOCX 17 kb)
10722_2024_1952_MOESM3_ESM.csv
Table 1. Improved intermediate wheatgrass (IWG) genets that have been assigned to a source population of 114 IWG accessions evaluated by Rodale Institute using a distance matrix made from molecular markers. Columns include tested genet, first and second highest source population (parent_1, parent_2), distance between tested genet and source accession (p1_value, p2_value), and germplasm group (TLI-Cycle 6, TLI-Cycle 12, Breeding Parents, and Remnant Rodale). (CSV 525 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Crain, J., Wagoner, P., Larson, S. et al. Origin of current intermediate wheatgrass germplasm being developed for Kernza grain production. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01952-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01952-1