Abstract
Opuntia ficus-indica (L.) Mill. is one of the most recognisable agricultural crops that can withstand harsh environmental conditions. South Africa is one of the few countries that hosts a large germplasm of Opuntia cultivars, which represent an ex-situ conservation population. However, little is known about the genetic diversity in this population. Additionally, some genotypes are morphologically indistinct, and therefore, making it a challenge for novice farmers and researchers to recognise specimens in the germplasm. The current study aimed to differentiate and measure the genetic diversity in 44 cultivars that represent the South African Opuntia germplasm using eight simple sequence repeat (SSR) markers. Evidently, the cultivars comprised moderate levels of diversity (average polymorphic information content, PIC = 0.37, Nei’s unbiased gene diversity = 0.42) that discriminated 90% of the cultivars. Analysis of the cultivars with the unweighted pair-group method using arithmetic averages (UPGMA) method revealed three main clusters, whereas principal coordinate analysis (PCoA) showed indistinct clustering of cultivars based on their usage in the agricultural market. Overall, the used SSR markers were able to distinguish most cultivars in the South African Opuntia germplasm—a genetic resource that appears to hold sufficient genetic diversity to aid the conservation and breeding of novel cultivars for the agriculture market.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cactus pear is a widely used and most economically valuable member of Cactaceae—a plant family that contains approximately 1600 species of cacti (Wallace and Gibson 2002). The family is currently classed into four subfamilies, and thus, Maihuenioideae, Pereskioideae, Cactoideae and Opuntioideae (Simpson 2010). The genus Opuntia (subfamily Opuntioideae) comprises taxa that are predominantly used in the agriculture industry as livestock feed (Dubeux et al. 2021; Pastorelli et al. 2022; Sipango et al. 2022). However, humans also consume cactus pear fruits (also called ‘prickly pear’) and young cladodes as vegetables (Barba et al. 2022). Fruit is highly nutritious as it contains numerous bioactive compounds (including betalains, carotenoids, flavonoids and phenols) along with proteins, minerals, vitamins, fatty acids, sterols, carbohydrates and fibres (Daniloski et al. 2022). Apart from nutritional benefits, cactus pear may further serve an important socio-economic role in the livelihood of human societies (Moshobane et al. 2022) and, as such, make it a crop of agricultural significance.
The genus Opuntia is highly diverse and comprises up to 250 different species (da Silva et al. 2022a). Members of Opuntia show different levels of ploidy, ranging from 2n = 2 × to 2n = 8x (Segura et al. 2007). Polyploidy in plants has been construed as the main driver of diversity in addition to supporting geographic expansion, climatic niche exchange and lineage longevity (Clark and Donoghue 2018). Further, polyploid genomes may have significant implications for plant breeding efforts by increasing allelic diversity, heterozygosity fixation, larger quantitative traits and genomic organisational changes at the genetic and epigenetic levels (van de Peer et al. 2017, 2021; da Silva et al. 2022a, b). Polyploidy in Opuntia facilitates survival in hostile environments (Granados-Aguilar et al. 2022). Opuntia ficus-indica (L.) Mill. and O. robusta J.C. Wendl. are, for instance, usually octoploids (2n = 8x = 88, Segura et al. 2007). Yet, O. ficus-indica can endure temperatures up to 70 °C (Granados-Aguilar et al. 2022), whereas O. robusta tolerates frost much better (Snyman et al. 2007). The capacity to withstand severe environments has made Opuntia species some of the important production crops in the South African agriculture industry (de Wit and Fouché 2021).
Opuntia breeding over the years has produced many cultivars and South Africa hosts one of the largest germplasm collections in the world (Chapman et al. 2002). However, cultivar identification tends to be challenging when plants are in a vegetative state due to inadequate distinctive features (Felker et al. 2006). Therefore, novel methods are required for differentiating Opuntia cultivars and species too. Modern high-throughput sequencing techniques have facilitated the discovery of genome-wide DNA markers that can be used to characterise germplasm pools and assess genetic diversity therein. To this end, simple sequence repeats (SSRs) have been established as molecular markers of choice owing to their abundance in plant genomes (Feng et al. 2016; Lu et al. 2019; Bhattarai et al. 2021).
As molecular markers, SSRs are more informative than the biallelic single nucleotide polymorphism (SNP) markers as the former tend to have many alleles per locus (Hamblin et al. 2007). Further, being co-dominant means that SSRs are valuable over dominant molecular markers as they will be relatively variable and may easily differentiate between homozygous and heterozygous individuals (Csencsics et al. 2010; Dutta et al. 2011). In the past, SSRs have been used to assess the genetic diversity in many diploid (Wen et al. 2010; Lin et al. 2015; Feng et al. 2016) and polyploid (Pfeiffer et al. 2011; Ali et al. 2019; Cusaro et al. 2022) plant species. Additionally, SSRs can discern morphologically indistinct cultivars (Caruso et al. 2010) and even determine gene flow incidents among individuals of different populations or closely related species (Fava et al. 2020). Then, SSRs are ideal markers for discerning and assessing the diversity in species and cultivars that form the South African Opuntia germplasm.
In this study, we aimed to distinguish and evaluate the genetic diversity in cultivars that make up the South African Opuntia germplasm. This was done using SSR markers that were previously identified in the Opuntia ficus-indica genome (Maleka et al., unpublished data). Similar to other genotypes all over the world (Caruso et al. 2010), some cultivars in the South African germplasm are phenotypically indistinguishable. Therefore, the current study may assist in genetically resolving the local germplasm in addition to enabling assessments of genetic diversity within breeding pools across South Africa.
Materials and methods
Plant material and DNA extraction
In 2015, an orchard including 44 Opuntia cultivars was established at the University of the Free State in Bloemfontein, Free State Province, Republic of South Africa (29°6′27.08″S; 26°11′32.90″E). The plants were propagated clonally from a 10-year-old evaluation trial that was planted at the Waterkloof farm, which is located roughly 30 km outside Bloemfontein. The Waterkloof orchard was established in 2005 using cladodes collected from other conservation sites located all over South Africa.
For the current study, a single flower was harvested from each of the 44 Opuntia cultivars (Table 1) and used for extracting genomic DNA. Specifically, genomic DNA was extracted from 50–100 mg of liquid nitrogen-powdered samples following the NucleoSpin Plant II Kit (Macherey–Nagel, Düren, Germany). However, cell lysis was completed at 65 °C for 30 min. DNA samples were quantified with a spectrophotometer (NanoDrop ND-1000, Thermo Fisher Scientific, Waltham, USA), whereas DNA quality was analysed by electrophoresis via 1% (w/v) TAE agarose gels. Gels were stained with the GelRed Nucleic Acid Gel Stain (Biotium, Hayward, USA) and visualised under UV light using a G:Box Gel Documentation System (Syngene, Cambridge, UK).
Polymerase Chain Reaction (PCR) amplification and genotyping of SSR loci
In the past, we identified many SSR loci in the Opuntia ficus-indica genome after Illumina sequencing (Maleka et al., unpublished data). In this study, the 5ʹ ends of forward primers for eight SSR loci were labelled with the 6-FAM fluorescent dye and used during PCR. PCR was carried out with 40–50 ng template DNA, 0.2 μM of each primer, 1 × EmeraldAmp GT PCR Master Mix (Takara Bio Inc., Shiga, Japan) and filled to 10 μL volumes per reaction with nuclease-free water. Cycling conditions included the primary denaturation at 98 °C for 3 min, followed by 30 cycles of denaturation at 98 °C for 10 s, primer annealing at 55.7 °C or 62.5 °C (Table 2) for 20 s and primer extension at 72 °C for 20 s. The final extension was done at 72 °C for 1 min. All reactions were effected on an Applied Biosystems 2720 Thermal Cycler (Applied Biosystems, Foster City, CA, USA). Post-PCR, selected products were stained with GelRed (Biotium) and electrophoresed on 3% (w/v) TBE agarose gels that contained a 100–3000 bp PCR DNA ladder (Ampliqon, Odense, Denmark). Once verified by gel visualisation (G:Box Gel Doc. System), amplicons were analysed by capillary electrophoresis on an ABI3500xl Genetic Analyzer (Applied Biosystems). The polymer POP-7 (Thermo Fisher Sci.) was applied as a separation matrix, while fragment sizes were determined with a GeneScan 500 LIZ Size Standard (Applied Biosystems) during these analyses.
Allele scoring and genetic diversity estimation
SSR alleles at the eight loci were scored with the software package GeneMarker v1.6 (BioGene Ltd., Kimbolton, UK). After scoring, allelic data were exported into the R packages polysat v1.7–6 (Clark and Jasieniuk 2011) and poppr v2.9.2 (Kamvar et al. 2014) for calculating different genetic diversity statistics. Statistics included the numbers of alleles (NA), allelic frequencies, the number of multilocus genotypes (MLG, which signifies the unique combination of alleles at two or more loci), the expected MLG (eMLG) and the Hardy–Weinberg equilibrium (HWE). Moreover, the polymorphic information content (PIC) for each locus was calculated with PIC_Calc 0.6 (http://www.biosoft.net/dna/pic.htm).
Genotypic diversity was computed using various indices, and these included the Simpson’s index (λ; Simpson 1949), the Evenness index (E.5; Grünwald et al. 2003) and Nei’s unbiased gene diversity (Hexp; Nei 1978). The software poppr was also used to create a genotype accumulation curve, which shows the power of random loci to discern unique individuals. Linkage disequilibrium (LD) between pairs of loci was determined via the standardized index of association test (rbarD; Agapow and Burt 2001). Contrary to the traditional index of association (IA), calculations for the rbarD are independent of the number of loci included in the test. The programme DARwin v6.0.021 (https://darwin.cirad.fr/) was used to do principal coordinate analysis (PCoA) and make a dendrogram based on the unweighted pair-group method using arithmetic averages (UPGMA).
Results
DNA quantity and quality
The amount of DNA extracted from the 44 Opuntia flower tissues ranged from 13.90–207.09 ng/uL (Table 1), with an average yield of 91.79 ng/uL. Three cultivars (Fusicaulis, Nepgen and Ofer) yielded DNA amounts < 20 ng/μL, while another three (Mexican, Meyers, and Roedtan) produced < 40 ng/μL. Regarding DNA quality, the A260/280 ratios ranged from 1.25–1.87 (Table 1), with an average ratio of 1.51. Curiously, cultivars that produced low DNA concentrations presented DNA of ideal quality, with Ofer and Meyers having the highest A260/280 ratios of 1.79 and 1.87, respectively.
SSR genotyping and genetic diversity estimates
Barring some loci in particular individuals, the eight loci generally yielded reproducible profiles that permitted allele scoring. Six of the eight SSR loci were polymorphic (thus, 75% polymorphism rate) and two (L5354 and L86067) were monomorphic. In total, the eight loci consisted of 24 alleles in the 44 Opuntia samples, with a mean number of three alleles per locus (Table 2). The 24 alleles occurred at frequencies ranging from 0.01–1.00. With six alleles, locus L161092 yielded the highest number of alleles per locus, and this corresponded to a PIC of 0.56. The most informative locus in this study, however, was L38909 (PIC = 0.62), and L37320 (PIC = 0.35) was the least polymorphic. The mean PIC value across all eight loci was 0.37 (Table 2), implying that the studied loci are fairly informative.
According to the poppr package, a total of 40 MLGs were expected (SE = 0.0) and realised based on the current data. The genotype accumulation curve, however, showed that five loci were sufficient to distinguish 90% of the 40 MLGs (Fig. 1). Each of the four remaining MLGs (10%) occurred twice in the samples (data not shown). The Simpson’s index was high, and this indicated that most genotypes are unique (λ = 0.973), while the high Evenness score (E.5 = 0.959) suggested that the population has equally abundant genotypes. Genetic variation in Opuntia cultivars was assessed with Nei’s unbiased genetic diversity index. To this end, locus L38909 emerged as the most diverse (Hexp = 0.66) in the study, followed by both L23031 (0.62) and L161092 (0.62). The least diverse polymorphic locus was L37320, with Hexp = 0.46 (Table 2). Across all loci, the Hexp index = 0.422 and this shows that the South African Opuntia germplasm harbours modest levels of genetic variation. Indices of the two LD tests deviated significantly from the null hypothesis of individuals in the population mating randomly (IA = 0.763, p-value = 0.02; rbarD = 0.173, p-value = 0.02). Hence, the null hypothesis is rejected and the alternative hypothesis (that the germplasm is asexual or clonal) is deemed to be valid. Four of the eight loci diverged from the HWE significantly (Table 2).
Genetic relationships among Opuntia cultivars
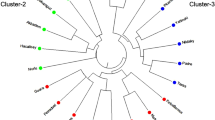
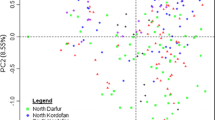
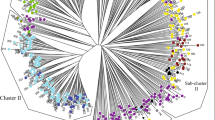
Relationships among the 44 Opuntia cultivars were analysed via clustering (UPGMA) and ordination (PCoA) methods. Clustering analysis produced a dendrogram with three main clades (Fig. 2). Overall, cultivars in clade I tended to be genetically farther distant from each other than those within clades II and III. However, the two cultivars of O. robusta (Monterey and Robusta) in clade I were genetically closer to each other as expected. Cultivars that historically came from the same country, surprisingly, did not cluster together. Also, the three cultivars from Botswana (R1251, R1259 and R1260) occurred in each of the three clades, while those from Israel (Ofer, Messina and Sharsheret) grouped into clades II and III. Clades II and III each comprised two branched tips relating to the four shared MLGs defined earlier. One of the divided tips showed Roedtan to be genetically related to Rossa, a cultivar of Italian origin (Fig. 2). As with the dendrogram, PCoA (Fig. 3) did not show any distinctive clustering, even when considering the commercial usage of cultivars as depicted in Table 1. Overall, the SSR loci in this study contain sufficient polymorphisms that effectively segregated cultivars in the South African Opuntia germplasm.
A UPGMA tree demonstrating the relationships among the 44 cultivars of Opuntia ficus-indica and O. robusta (Robusta and Monterey). The tree reveals three main clusters referred to as clades I–III. Clade II shows two branched tips that clustered Malta with American Giant and Meyers with Berg x Mexican. Rossa and Roedtan along with Postmasburg and Muscatel occupy the branched tips in clade III
Principal Coordinate Analysis (PCoA) showing the relationships among the 44 cultivars of Opuntia ficus-indica and O. robusta (Robusta and Monterey). The numbers refer to the cultivars as per Table 1. Overlaid numbers, marked with black arrows, relate to the branched tips in Fig. 2 as follows: 2 + 16, 5 + 19, 22 + 26 as well as 32 + 34. Coloured circles indicate cultivar usage as follows: orange (fruit), green (forage), grey (fruit/forage) and blue (unknown)
Discussion
Members of the Cactaceae attract research interest globally because of their specialised physiological traits that allow survival in severe habitats (Nobel 2002). Among such interests is the requirement to genetically characterise and differentiate species and cultivars, particularly within the genus Opuntia. Studies have been done on many ecotypes from diverse germplasms using various molecular markers (Omar et al. 2021). Opuntia cultivars forming the South African germplasm have, however, received little molecular characterisation despite this being one of the key genetic pools (Chapman et al. 2002). Notable studies include those of Oelofse (2002) and Mashope et al. (2011) that both applied amplified fragment length polymorphisms (AFLPs) as molecular markers to discern cultivars in the germplasm. Similarly, the current study sought to differentiate and measure the genetic diversity within the South African Opuntia germplasm using eight genomic SSR markers. Information on the relationships and genetic diversity in cultivars will assist in the selection of superior genotypes toward the development of new cultivars and preservation of cultivars as distinct genotypes. Overall, the studied SSR markers were able to distinguish 90% of the Opuntia cultivars, and evidently, the genotypes harbour moderate levels of genetic variability.
DNA extraction from cacti is infamously challenging due to the large quantity of mucilage, secondary metabolites and polysaccharides (Nobel et al. 1992). As such, several protocols have been developed and improved over the years to try overcoming this challenge (De La Cruz et al. 1997; Mondragon-Jacobo et al. 2000; Fehlberg et al. 2013; Martínez-González et al. 2017). Although the protocols have been effective, they tend to be laborious. Moreover, some protocols need large amounts (ca. 5–8 g) of tissue material to begin with (Mondragon-Jacobo et al. 2000). Interestingly, Fehlberg et al. (2013) developed a novel method that is based on relatively small amounts (ca. 40–100 mg) of cactus pear spines. Our study also sought to simplify DNA isolation from cactus pear by using small amounts of flower tissues because they are soft and amenable to crushing with minimal effort. While the amount of DNA extracted from flower tissues was below that obtained from cladodes using current protocols, the quality was, however, similar (Martínez-González et al. 2017). In retrospect, we suppose that the flower tissues were not crushed sufficiently, and this impacted DNA recovery. Even so, our approach establishes flowers as an alternative source of quality DNA for molecular studies in Opuntia. Tissue sampling, however, should be planned around the limited annual flowering season.
Of the eight SSR markers used in the current study, five were able to discriminate 90% of the 44 O. ficus-indica samples—meaning that ~ 20% of the samples (four MLGs) remained undifferentiated. In European plum (Prunus domestica L.), however, four genomic SSRs were sufficient to differentiate 44 MLGs and this was attributed to the high number of alleles detected at the loci (averaging 19.2 per locus; Manco et al. 2020). Elsewhere, Fiore et al. (2022) were able to distinguish 30 hazelnut (Corylus avellana L.) genotypes via nine SSR markers that had an average of 7.67 alleles per locus. Given that the Opuntia genotypes in the current study revealed low to moderate allelic diversity per locus, it can, therefore, be asserted that the genetic diversity in the germplasm was most likely underestimated. The use of additional markers to further distinguish the overlapped genotypes will offer an opportunity to better estimate the genetic diversity in this germplasm.
To our knowledge, this is the first study to describe estimates of genetic diversity in the South African Opuntia germplasm via SSRs. The moderate levels of polymorphism (average PIC = 0.37) observed in the studied loci were similar to those described by Mashope et al. 2011 (average PIC = 0.29) based on AFLP markers. PIC indicates the ability of a marker to identify polymorphisms among individuals in a population, and this depends on the number of alleles detected at each locus and the corresponding allelic distribution frequencies (Serrote et al. 2020). Markers with PIC values ≥ 0.50 are usually said to be very informative, while values in the range 0.25–0.50 are moderately informative. Under 0.25, the markers are deemed not to be very informative. Therefore, the current SSR markers and the AFLP markers used on Opuntia samples in the past (Mashope et al. 2011) are moderately informative.
Prior to this study, SSRs have been used several times to assess the genetic diversity in global Opuntia specimens (Helsen et al. 2009; Caruso et al. 2010; Erre et al. 2011; Chessa et al. 2013; Samah et al. 2016; Reis et al. 2018). Astonishingly, the Opuntia germplasm from Mexico presented low levels of polymorphisms (average PIC = 0.246; Samah et al. 2016), as did a global group of wild and cultivated genotypes that represented 16 different species of Opuntia (average PIC = 0.191; Caruso et al. 2010). Asserted reasons for the low polymorphic rate included the background of researched germplasms as well as the choice and number of tested SSR loci (Samah et al. 2016). Overall, that the South African Opuntia germplasm is more polymorphic than specimens used by Caruso et al. (2010) and Samah et al. (2016) was remarkable and encouraging.
In contrast to the above studies, Chessa et al. (2013) and Bendhifi Zarroug et al. (2015) reported high levels of polymorphism (average PIC > 0.71) within global genotypes of Opuntia species and cultivars having diverse histories. Similarly, Reis et al. (2018) found high levels of polymorphisms (average PIC = 0.711) in Opuntia populations from Portugal. In other plants, the average PIC from genomic SSRs ranged from 0.25 in global accessions of Spinacia oleracea L. (Bhattarai et al. 2021); 0.29–0.43 in cultivars of five Prunus species (Dettori et al. 2015); 0.61 in worldwide accessions of Olea europaea L. (Li et al. 2020) and 0.75 in Chinese genotypes of Citrus sinesis L. Osbeck (Biswas et al. 2014). In addition, Ali et al. (2019) reported an average PIC of 0.92 in species and hybrids of Saccharum and Erianthus. Nevertheless, the high levels of polymorphisms and diversity observed in Opuntia samples were attributed to the varied origins of cultivars along with the prevalence of asexual propagation and limited artificial selection (Chessa et al. 2013; Bendhifi Zarroug et al. 2015). The Opuntia germplasm in South Africa seemingly derive from the Burbank collection, which was produced through extensive crossings and selections among accessions from Mexico, Africa, Australia and several other countries (Chapman et al. 2002). Thus, their histories are convoluted. Also, the modest levels of polymorphisms and diversity in the South African Opuntia germplasm may suggest the adaptation of cultivars to the local climate. Increased genetic diversity and divergence are known indicators of organisms adapting to new or extreme environments (Chen et al. 2018; Wang et al. 2018). Indeed, the Opuntia population at the Waterkloof orchard was founded recently in relation to other cactus pear conservation sites in South Africa (Potgieter and Mashope 2009). Cultivars in this study may, therefore, be adapting to the local environment—a notion that can only be confirmed by analysing the genetic diversity in Opuntia cultivars grown at other sites in South Africa. Overall, our findings affirm the South African Opuntia germplasm as one of the diverse and key breeding pools of cactus pear in the world.
The 44 Opuntia cultivars differentiated into three main clades, unlike the nine identified by Mashope et al. (2011) using AFLPs on 38 cultivars. Moreover, there was no distinct grouping of cultivars based on their main usage (either as fruit or fodder) in the agriculture industry. Such grouping was, however, noted when using randomly amplified polymorphic DNA (RAPD) markers to describe a few Opuntia samples (Wang et al. 1998). Mashope et al. (2011) also discovered some clustering in relation to plant growth habit (upright or bushy), whereas Opuntia ficus-indica samples from Portugal clustered based on fruit pulp colour (Reis et al. 2018). While it would have been interesting to detect some reasonable clustering in the South African Opuntia germplasm, the lack of it is also significant as it suggests the presence of sufficient levels of diversity in the cultivars. Future studies can try to identify and develop molecular markers associating with traits desired in the agriculture market (e.g., fruit quality, de Wit et al. 2010; Coetzer et al. 2019). Similar studies have already been done on fruit traits in sweet cherry (Ganopoulos et al. 2011) and kiwifruit (Liao et al. 2019). The South African Opuntia germplasm, in essence, appears to not only support the ex-situ conservation of cactus pear, but also holds the genetic capacity for breeding novel cultivars that will benefit the agriculture industry.
Conclusions
South Africa hosts one of the major germplasms of Opuntia ficus-indica in the world. The germplasm understandably derives from the Burbank collection that was established using material sourced from all over the world. Consequently, the diverse history and possibly adaptation to the local environment over many years may have influenced the moderate genetic variation found in South African cultivars. Genetic variability in the South African Opuntia cultivars means that the germplasm may be used for breeding new cultivars for the agriculture industry. Alternatively, the cultivars may serve as a source population for deriving new alleles that can rescue less diverse Opuntia breeding programmes around the world. Last, the SSR markers used in this study will be useful toward distinguishing and verifying unknown samples used by farmers and researchers all over South Africa.
References
Agapow P-M, Burt A (2001) Indices of multilocus linkage disequilibrium. Mol Ecol Notes 1:101–102. https://doi.org/10.1046/j.1471-8278.2000.00014.x
Ali A, Pan Y-B, Wang Q-N et al (2019) Genetic diversity and population structure analysis of Saccharum and Erianthus genera using microsatellite (SSR) markers. Sci Rep 9:395. https://doi.org/10.1038/s41598-018-36630-7
Barba FJ, Garcia C, Fessard A et al (2022) Opuntia ficus-indica edible parts: A food and nutritional security perspective. Food Rev Int 38:930–952. https://doi.org/10.1080/87559129.2020.1756844
Bendhifi Zarroug M, Salhi Hannachi A, Souid S et al (2015) Molecular research on the genetic diversity of cactus (Opuntia spp.) using the SSR method. Acta Hortic 1067:53–58. https://doi.org/10.17660/ActaHortic.2015.1067.6
Bhattarai G, Shi A, Kandel DR et al (2021) Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions. Sci Rep 11:9999. https://doi.org/10.1038/s41598-021-89473-0
Biswas MK, Xu Q, Mayer C, Deng X (2014) Genome wide characterization of short tandem repeat markers in sweet orange (Citrus sinensis). PLoS ONE 9:e104182
Caruso M, Currò S, Las CG et al (2010) Microsatellite markers help to assess genetic diversity among Opuntia ficus-indica cultivated genotypes and their relation with related species. Plant Syst Evol 290:85–97. https://doi.org/10.1007/s00606-010-0351-9
Chapman B, Mondragon Jacobo C, Bunch RA, Paterson AH (2002) Breeding and Biotechnology. In: Nobel PS (ed) Cacti: Biology and uses. University of California Press, Los Angeles, pp 255–271
Chen Y, Shenkar N, Ni P et al (2018) Rapid microevolution during recent range expansion to harsh environments. BMC Evol Biol 18:187. https://doi.org/10.1186/s12862-018-1311-1
Chessa I, Erre P, Barbato M et al (2013) Polymorphic microsatellite DNA markers in Opuntia spp. collections. Acta Hortic 995:35–42. https://doi.org/10.17660/actahortic.2013.995.3
Clark JW, Donoghue PCJ (2018) Whole-genome duplication and plant macroevolution. Trends Plant Sci 23:933–945. https://doi.org/10.1016/j.tplants.2018.07.006
Clark LV, Jasieniuk M (2011) polysat: an R package for polyploid microsatellite analysis. Mol Ecol Resour 11:562–566. https://doi.org/10.1111/j.1755-0998.2011.02985.x
Coetzer GM, De Wit M, Fouché HJ, Venter SL (2019) Climatic influences on fruit quality and sensory traits of cactus pear (Opuntia ficus-indica): A 5-year evaluation. Acta Hortic 1247:23–30
Csencsics D, Brodbeck S, Holderegger R (2010) Cost-effective, species-specific microsatellite development for the endangered dwarf bulrush (Typha minima) using next-generation sequencing technology. J Hered 101:789–793
Cusaro CM, Grazioli C, Zambuto F et al (2022) An improved method for assessing simple sequence repeat (SSR) variation in Echinochloa crus-galli (L.) P. Beauv (Barnyardgrass). Diversity 14:3. https://doi.org/10.3390/d14010003
da Silva RT, Nollet F, dos Santos DC et al (2022a) Heterochromatin and polyploidy in Opuntia Mill. (Cactaceae: Opuntioideae): Implications for plant breeding. New Zeal J Crop Hortic Sci. https://doi.org/10.1080/01140671.2022.2028861
da Silva RT, RdeLA B, Nascimento RGS et al (2022b) Physiological quality of forage palm seeds and seedlings associated with the level of ploidy. Acta Sci Agron 44:e55711. https://doi.org/10.4025/actasciagron.v44i1.55711
Daniloski D, D’Cunha NM, Speer H et al (2022) Recent developments on Opuntia spp., their bioactive composition, nutritional values, and health effects. Food Biosci 47:101665. https://doi.org/10.1016/j.fbio.2022.101665
De La Cruz M, Ramirez F, Hernandez H (1997) DNA isolation and amplification from cacti. Plant Mol Biol Report 15:319–325
de Wit M, Nel P, Osthoff G, Labuschagne MT (2010) The effect of variety and location on cactus pear (Opuntia ficus-indica) fruit quality. Plant Foods Hum Nutr 65:136–145. https://doi.org/10.1007/s11130-010-0163-7
de Wit M, Fouché H (2021) South African perspective on Opuntia spp.: Cultivation, human and livestock food and industrial applications BT–Opuntia spp: Chemistry, bioactivity and industrial applications. In: Ramadan MF, Ayoub TEM, Rohn S (eds). Springer International Publishing, Cham, pp 13–47
Dettori MT, Micali S, Giovinazzi J et al (2015) Mining microsatellites in the peach genome: development of new long-core SSR markers for genetic analyses in five Prunus species. Springerplus 4:337. https://doi.org/10.1186/s40064-015-1098-0
Dubeux JCB, Santos MVFD, Cunha MVD et al (2021) Cactus (Opuntia and Nopalea) nutritive value: a review. Anim Feed Sci Technol 275:114890. https://doi.org/10.1016/j.anifeedsci.2021.114890
Dutta S, Kumawat G, Singh BP et al (2011) Development of genic-SSR markers by deep transcriptome sequencing in pigeonpea [Cajanus cajan (L.) Millspaugh]. BMC Plant Biol 11:17–29
Erre P, Nieddu G, Chessa I (2011) Identification of microsatellite loci in Opuntia spp. and their characterization in cultivars and species. Acta Hortic 918:327–332. https://doi.org/10.17660/ActaHortic.2011.918.42
Fava WS, Gomes VGN, Lorenz AP, Paggi GM (2020) Cross-amplification of microsatellite loci in the cacti species from Brazilian chaco. Mol Biol Rep 47:1535–1542. https://doi.org/10.1007/s11033-019-05064-3
Fehlberg SD, Allen JM, Church K (2013) A novel method of genomic DNA extraction for cactaceae. Appl Plant Sci 1:apps.1200013. https://doi.org/10.3732/apps.1200013
Felker P, Paterson A, Jenderek M (2006) Forage potential of Opuntia clones maintained by the USDA, National Plant Germplasm System (NPGS) Collection. Crop Sci 46:2161–2168
Feng S, He R, Lu J et al (2016) Development of SSR markers and assessment of genetic diversity in medicinal Chrysanthemum morifolium cultivars. Front Genet 7:113. https://doi.org/10.3389/fgene.2016.00113
Fiore MC, Marchese A, Mauceri A et al (2022a) Diversity assessment and DNA-based fingerprinting of Sicilian hazelnut (Corylus avellana L) germplasm. Plants 11:631. https://doi.org/10.3390/plants11050631
Ganopoulos IV, Kazantzis K, Chatzicharisis I et al (2011) Genetic diversity, structure and fruit trait associations in Greek sweet cherry cultivars using microsatellite based (SSR/ISSR) and morpho-physiological markers. Euphytica 181:237–251. https://doi.org/10.1007/s10681-011-0416-z
Granados-Aguilar X, Palomino G, Martínez-Ramón J, Arias S (2022) Genome evolution and phylogenetic relationships in Opuntia tehuacana (Cactaceae, Opuntioideae). Brazilian J Bot 45:957–969. https://doi.org/10.1007/s40415-022-00821-4
Grünwald NJ, Goodwin SB, Milgroom MG, Fry WE (2003) Analysis of genotypic diversity data for populations of microorganisms. Phytopathology 93:738–746. https://doi.org/10.1094/PHYTO.2003.93.6.738
Hamblin M, Warburton M, Buckler E (2007) Empirical comparison of simple sequence repeats and single nucle otide polymorphisms in assessment of maize diversity and relatedness. PLoS ONE 2:e1367
Helsen P, Verdyck P, Tye A, Van Dongen S (2009) Low levels of genetic differentiation between Opuntia echios varieties on Santa Cruz (Galapagos). Plant Syst Evol 279:1–10. https://doi.org/10.1007/s00606-008-0064-5
Kamvar ZN, Tabima JF, Gr̈unwald NJ, (2014) poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014:1–14. https://doi.org/10.7717/peerj.281
Li D, Long C, Pang X et al (2020) The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions. PeerJ 8:e8573. https://doi.org/10.7717/peerj.8573
Liao G, Li Z, Huang C et al (2019) Genetic diversity of inner quality and SSR association analysis of wild kiwifruit (Actinidia eriantha). Sci Hortic (amsterdam) 248:241–247. https://doi.org/10.1016/j.scienta.2019.01.021
Lin YS, Kuan CS, Weng IS, Tsai CC (2015) Cultivar identification and genetic relationship of pineapple (Ananas comosus) cultivars using SSR markers. Genet Mol Res 14:15035–15043. https://doi.org/10.4238/2015.November.24.11
Lu Q, Hong Y, Li S et al (2019) Genome-wide identification of microsatellite markers from cultivated peanut (Arachis hypogaea L). BMC Genomics 20:799. https://doi.org/10.1186/s12864-019-6148-5
Manco R, Chiaiese P, Basile B, Corrado G (2020) Comparative analysis of genomic- and EST-SSRs in European plum (Prunus domestica L.): implications for the diversity analysis of polyploids. Biotech 10:543. https://doi.org/10.1007/s13205-020-02513-w
Martínez-González CR, Ramírez-Mendoza R, Jiménez-Ramírez J et al (2017) Improved method for genomic DNA extraction for Opuntia Mill (Cactaceae). Plant Methods 13:82. https://doi.org/10.1186/s13007-017-0234-y
Mashope BK, Herselman L, Labuschagne MT (2011) Genetic diversity among South African cactus pear genebank accessions using AFLP markers. Bradleya. https://doi.org/10.25223/brad.n29.2011.a12
Mashope BK (2007) Characterization of cactus pear germplasm in South Africa, PhD thesis, Department of Plant Science, University of the Free State.
Mondragon-Jacobo C, Doudareva N, Bordelon BP (2000) DNA extraction from several cacti. HortScience 35:1124–1126. https://doi.org/10.21273/hortsci.35.6.1124
Moshobane MC, Olowoyo JO, Middleton L (2022) The influence of Opuntia ficus-indica on human livelihoods in Southern Africa. Plants People Planet 4:451–462. https://doi.org/10.1002/ppp3.10278
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590. https://doi.org/10.1093/genetics/89.3.583
Nobel PS (2002) Cacti: biology and uses. University of California Press, Los Angeles, First Edit
Nobel PS, Cavelier J, Andrade JL (1992) Mucilage in cacti: Its apoplastic capacitance, associated solutes, and influence on tissue. J Exp Bot 43:641–648. https://doi.org/10.1093/jxb/43.5.641
Oelofse RM (2002) Characterization of Opuntia ficus-indica cultivars in South Africa. MSc dissertation, Department of Plant Science, University of the Free State.
Omar AA, ElSayed AI, Mohamed AH (2021) Genetic diversity and ecotypes of Opuntia spp. BT—Opuntia spp.: Chemistry, bioactivity and industrial applications. In: Ramadan MF, Ayoub TEM, Rohn S (eds). Springer International Publishing, Cham, pp 181–199
Pastorelli G, Serra V, Vannuccini C, Attard E (2022) Opuntia spp. as alternative fodder for sustainable livestock production. Animals 12:1597. https://doi.org/10.3390/ani12131597
Pfeiffer T, Roschanski AM, Pannell JR et al (2011) Characterization of microsatellite loci and reliable genotyping in a polyploid plant, Mercurialis perennis (Euphorbiaceae). J Hered 102:479–488. https://doi.org/10.1093/jhered/esr024
Potgieter JP, Mashope BK (2009) Cactus pear (Opuntia spp.) germplasm conservation in South Africa. Acta Hortic 811:47–54
Reis CMG, Raimundo J, Ribeiro MM (2018) Assessment of genetic diversity in Opuntia spp Portuguese populations using SSR molecular markers. Agronomy 8:55. https://doi.org/10.3390/agronomy8040055
Samah S, De Teodoro Pardo CV, Serrato Cruz MA, Valadez-Moctezuma E (2016) Genetic diversity, genotype discrimination, and population structure of Mexican Opuntia sp., determined by SSR markers. Plant Mol Biol Report 34:146–159. https://doi.org/10.1007/s11105-015-0908-4
Segura S, Scheinvar L, Olalde G et al (2007) Genome sizes and ploidy levels in Mexican cactus pear species Opuntia (Tourn.) Mill. series Streptacanthae Britton et Rose, Leucotrichae DC., Heliabravoanae Scheinvar and Robustae Britton et Rose. Genet Resour Crop Evol 54:1033–1041
Serrote CML, Reiniger LRS, Silva KB et al (2020) Determining the polymorphism information content of a molecular marker. Gene 726:144175. https://doi.org/10.1016/j.gene.2019.144175
Simpson EH (1949) Measurement of diversity. Nature 163:688. https://doi.org/10.1038/163688a0
Simpson MG (2010) Diversity and classification of flowering plants: Eudicots. In: Second E (ed) Simpson MGBT-PS. Academic Press, San Diego, pp 275–448
Sipango N, Ravhuhali KE, Sebola NA et al (2022b) Prickly pear (Opuntia spp) as an invasive species and a potential fodder resource for ruminant animals. Sustain 14:3719. https://doi.org/10.3390/su14073719
Snyman HA, Fouché HJ, Avenant PL, Ratsele C (2007) Frost sensitivity of Opuntia ficus-indica and O. robusta in a semiarid climate of South Africa. J Prof Assoc Cactus Dev 9:1–21
van de Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18:411–424. https://doi.org/10.1038/nrg.2017.26
van de Peer Y, Ashman TL, Soltis PS, Soltis DE (2021) Polyploidy: an evolutionary and ecological force in stressful times. Plant Cell 33:11–26. https://doi.org/10.1093/plcell/koaa015
Wallace RS, Gibson AC (2002) Evolution and Systematics: Cacti: Biology and uses. Park PS. University of California Press, Los Angeles, pp 1–21
Wang X, Felker P, Burow MD, Paterson AH (1998) Comparison of RAPD marker patterns to morphological and physiological data in the classification of Opuntia accessions. J Prof Assoc Cactus Dev 3:98–109
Wang X, Zhao W, Li L et al (2018) Clonal plasticity and diversity facilitates the adaptation of Rhododendron aureum Georgi to alpine environment. PLoS ONE 13:e0197089. https://doi.org/10.1371/journal.pone.0197089
Wen M, Wang H, Xia Z et al (2010) Developmenrt of EST-SSR and genomic-SSR markers to assess genetic diversity in Jatropha curcas L. BMC Res Notes 3:42. https://doi.org/10.1186/1756-0500-3-42
Acknowledgements
The authors wish to thank the Department of Science & Innovation and the Human Science Research Council of South Africa for funding Tsholofelo J. Modise’s internship position.
Funding
Open access funding provided by University of the Free State. The authors certify that they have no affiliations or involvement in any organization/entity that has financial or non-financial interest in the subject matter or materials discussed in this manuscript.
Author information
Authors and Affiliations
Contributions
All authors were involved in the writing, reviewing and editing of the manuscript, whereas Tsholofelo Modise conducted all experiments and data analysis during the study. The datasets produced and/or analysed during the current study are available from the corresponding author on reasonable request.
Corresponding author
Ethics declarations
Competing interests
The authors declare that no external funding was received to help with the execution of this study or preparation of this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Modise, T.J., Maleka, M.F., Fouché, H. et al. Genetic diversity and differentiation of South African cactus pear cultivars (Opuntia spp.) based on simple sequence repeat (SSR) markers. Genet Resour Crop Evol 71, 373–384 (2024). https://doi.org/10.1007/s10722-023-01629-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01629-1