Abstract
Chamomile (Matricaria chamomilla L., Asteraceae) is one of the most important medicinal plants with antiphlogistic, spasmolytic, carminative, antibacterial and antimycotic properties. Thirty-one chloroplast markers were developed, optimised for high-resolution melting analysis. Subsequently, 23 M. chamomilla accessions (247 individuals) complemented with 2 individuals of the closely related species M. discoidea were analysed with this marker set. The marker set can practically be reduced to 20 markers without information loss due to the linkage of 16 markers. In total, 20 chlorotypes (multilocus genotypes) were identified, organised in three evolutionary main lineages. Only 8 accessions were monomorphic, the other 15 accessions had between 2 and 6 chlorotypes per accession. The high number of polymorphic accessions and the high number of chlorotypes within many accessions indicate already a high degree of variability within accessions, confirmed by the 66% variation within by AMOVA. Gene diversity of the polymorphic accessions ranged between 0.069 and 0.261. Since many cultivars had their origin in wild populations of the countries where breeding took place, a first hypothetical indication of the geographic origin of chamomile could point to Albania and adjacent countries. The ‘ancient chlorotype’ was identified by comparing published plastome sequences of Artemisia annua and Lactuca sativa. The ‘ancient chlorotype’ was present only in the closely related M. discoidea but not in any of the M. chamomilla accessions. Two commercially important tetraploid cultivars proofed to be unrelated to their presumed diploid parents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Matricaria L. is a small genus of 25 species in the family of Asteraceae, of which Matricaria chamomilla L. (German chamomile) (WFO) is an important medicinal plant with a trade volume of about 8,000t per year (Franke and Hannig 2012). The species origins are suggested to be in Europe and West Asia. Today, German chamomile is present in many temperate regions worldwide (Franke and Schilcher 2007). German chamomile is an annual, herbaceous plant of 10 to 90 cm height and a natively diploid species (2n = 2x = 18) (Albrecht and Otto 2020). Combined pharmacological and biochemical effects of several chamomile compounds, mainly from the essential oil and flavonoids, are responsible for the therapeutic effectiveness of chamomile products, which possess antiphlogistic, spasmolytic, carminative, antibacterial and antimycotic properties (Schilcher et al. 2005).
Breeding activities started approximately 50 years ago from natural European populations in former Czechoslovakia, Poland, Hungary and Germany and led to the development of diploid and autotetraploid cultivars (Albrecht and Otto 2020; Franke and Schilcher 2007). Breeding focussed on the improvements of crop yield (flower heads), essential oil content and composition (correct chemotype), height adapted to harvesting and disease resistance (Albrecht and Otto 2020). Autopolyploidisation of chamomile resulted in higher weight of developing chamomile anthodia and ligulate flowers (Glücknerova et al. 1965) and in a higher cuticular flavonoid content (Repčák et al. 1999). Therefore, tetraploid varieties are important in commercial chamomile production.
Several molecular marker studies were performed in German chamomile to study genetic variability (Ahmadi et al. 2014; Mehdikhani et al. 2014; Mežaka et al. 2020; Okoń et al. 2013; Otto et al. 2017; Pirkhezri et al. 2010; Ruzicka et al. 2021; Ruzicka and Novak 2020; Solouki et al. 2008; Wagner et al. 2005). Only a few of them (Otto et al. 2017; Ruzicka et al. 2021; Ruzicka and Novak 2020) were using SNP markers and only two (Ruzicka et al. 2021; Ruzicka and Novak 2020) focused on extranuclear (mitochondrial) DNA markers.
Extranuclear or cytoplasmic DNA (cpDNA), located outside the cell nucleus in mitochondria and plastids, are essential tools in molecular systematics (Timmis and Ayliffe 1990), but also helpful for certain scientific intraspecific problems of e.g. phylogeography (Avise et al. 1987; Shaal et al. 1998). Extranuclear DNA is inherited only maternally (in rare cases paternally) without recombination. Therefore, cpDNA markers are usually of a lower variability than nuclear DNA markers. In case of hypervariability of nuclear markers, chloroplast markers can be more informative and provide sufficient variability (Osterberger et al. 2021).
HRM (high resolution melting) is a ‘post-PCR’ method able to detect SNPs by differences of the melting curves of double stranded DNA recorded at high resolution (Chatzidimopoulos et al. 2019). The method allows a high throughput at low costs per reaction and is widely used for mutation scanning and SNP detection in plant breeding and genotyping (Borna et al. 2017, Sorkheh et al. 2017, Kim and Kim 2019).
In this research, chloroplast markers were used to study genetic relationships between and variability within different accessions of chamomile. This study complements the mitochondrial marker previous research by Ruzicka and Novak (2020) and Ruzicka et al. (2021).
Material and methods
Plant material
Chamomile were grown in the greenhouse and young leaves of 249 individuals from 24 accessions were harvested, dried and stored on silica until DNA extraction (Table 1).
DNA extraction
Approximately 1 cm2 of dried leaf material was grounded with glass beads in a swing mill (Mixer Mill MM301, Retsch GmbH, Germany) and total genomic DNA isolated using a modified CTAB extraction protocol (Schmiderer et al. 2013). Quantity and quality of the DNA were determined on a spectrophotometer (NanoDrop™ 2000, Thermo Fisher Scientific Inc., USA) and via gel electrophoresis on a 1.4% agarose gel stained with peqGREEN (VWR, Austria). The DNA was dissolved in TE buffer and stored at − 20 °C until further usage.
SNP detection and primer design
NGS data from a previous study (Submission ID: SUB5046906/BioProject ID: PRJNA515664) were de novo assembled to source chloroplast SNP markers. Some chloroplast consensus sequences were blasted and Artemisia annua was identified as the closest related species with a complete published plastome (KY085890) and therefore used as reference. From the three individuals of cultivar ‘Promyk’ a complete chloroplast was assembled and submitted to GenBank (accession no. OM911930, haplotype B3). Reads of the 11 accessions were mapped to the chloroplast reference and SNP identified with Geneious® 11.1.5 (Biomatters Ltd., New Zealand). All primers were developed in Geneious® 11.1.5 and were synthesised by Sigma-Aldrich GmbH (Austria) (Supplementary Table 1).
To identify the ‘ancient’ chlorotype, the chamomile chloroplast was aligned with Artemisia annua (KY085890) and Lactuca sativa (AP007232) plastomes and marker polymorphisms compared and two samples of Matricaria discoidea DC. (syn. M. matricarioides Porter ex Britton) were integrated in HRM analysis as ‘ancient marker type’ (confirmed by sequencing).
High resolution melting analysis (HRM)
HRM was performed in duplicates on a Rotor-Gene™ 6000 with the Rotor-Gene Q Series Software 2.1.0 (Qiagen, Germany). The final reaction volume of 10 µl contained 1 × HRM mastermix (HOT FIREPol® EvaGreen® HRM Mix, Solis BioDyne, Estonia), 150 nM of the forward and reverse primer each, and 2 ng genomic DNA. PCR started with an initial phase of 14 min at 95 °C, 45 cycles of 95 °C/57 °C to 60 °C or to 72 °C (depending on the primer pair). After PCR, a hold at 95 °C for 1 min guaranteed complete denaturation of all nucleotide strands. HRM was done in increments of 0.1 °C and 1 s hold before each temperature step.
Evaluation of the HRM analysis
qPCR results were checked for average fluorescence levels and average Cq values of the samples. Products with Cq values over 30, with a standard deviation of the duplicates ≥ 0.25 and those with an end fluorescence level of less than 80% of the overall average were repeated. Previously sequenced samples served as reference samples for the classification of curve types and were added to each run of the respective marker. The ∆TM of the curve types (Table 2) were calculated for each primer combination and samples were assigned to the respective curve type with a deviation of 30% from the calculated TM of the reference.
Statistical analysis
Gene diversity (GD), Evenness (E.5), the number of multilocus genotypes (MLG) standardized by sample numbers, the Shannon–Wiener Index (H), the Index of association (Brown et al. 1980) corrected for the number of markers (ṝd) (Agapow and Burt 2001) and AMOVA were calculated. The number of multilocus genotypes (MLG) and Nei’s genetic distance were used for construction of a minimum spanning network. These calculations were performed in R 3.3.0 with the package poppr 2.2.0 (Kamvar et al. 2014, 2015). Matrix correlation between chloroplast and mitochondrial distance matrices were calculated according to the modified RV (= RV2) as implemented in the R package MatrixCorrelation (Indahl et al. 2018).
Results
A set of 31 chloroplast markers was developed and 247 M. chamomilla individuals from 24 accessions were analysed with this marker set. Additionally, two individuals of M. discoidea, the Matricaria species closest related to M. chamomilla were analysed to gain information beyond the species level.
Development of a core SNP marker set
NGS sequences of whole genome samples (NCBI accession no. SRX5257894 to SRX5257926 (Ruzicka et al. 2021; Ruzicka and Novak 2020)) were mapped to the closest related chloroplast available (Artemisia annua, accession no. KY085890). Amongst the polymorphisms found between the cultivars, a primer set of 31 markers was selected suitable for high-resolution melting analysis. In total 24 accessions (249 samples) were analysed with this set.
Characterisation of markers
The mean temperature difference of the melting curves within a marker was 0.45 °C ranging from 0.2 °C in mrcp39 to 0.71 °C in mrcp64 (Table 2). The 31 markers showed low to medium diversity with a mean gene diversity of 0.285. Gene diversity ranged from 0.016 (mrcp66) to 0.498 (mrcp04, mrcp27, mrcp33, mrcp38 and mrcp39). Evenness (mean value of 0.703) ranged from 0.339 (mrcp66) to 0.993 (mrcp27). In eleven markers the genotypes were almost equally distributed (Evenness > 0.9), while in five markers the genotypes were uneven distributed (Evenness < 0.5).
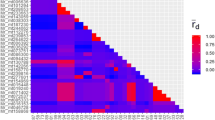
For estimating the linkage disequilibrium between the markers, the Index of association (ṝd) was calculated across all markers. The average index of association corrected for the number of markers (ṝd) was 0.20. Five groups with complete linkage of some markers were identified (Table 2). Sixteen markers belonged to one of the linkage groups. Recalculation of ṝd without the completely linked markers lowered the index of association only to 0.18 (p < 0.05).
Chlorotypes (= multilocus genotypes)
Twenty-one chlorotypes (= multilocus genotypes on chloroplast DNA, MLG) were identified. The minimum spanning network shows the presence of three evolutionary main lineages (‘A’, ‘B’, ‘C’) and with increasing index mostly a higher differentiation from the origin (Fig. 1). Comparisons with the plastomes of Artemisia annua and Lactuca sativa (both Asteraceae) showed that all 31 mutations occurred within Matricaria. Analysis of two individuals of M. discoidea—the closely related Matricaria species—restricted all these mutations to the time after the branching of the two species from their common ancestor.
Minimum spanning network of the 20 chlorotypes (left) and VENN diagram of the distribution of accessions amongst the three major chlorotype lineages A, B and C (right). ‘ac’ is the ancient chlorotype present in M. discoidea, Artemisia annua and Lactuca sativa but not in any samples of M. chamomilla. The network clearly shows three major evolutionary lines A, B and C. Thicker lines represent closer related nodes
Eight of the 23 M. chamomilla accessions were monomorphic with only one chlorotype (Table 3). Five of the eight monomorphic accessions originated from natural stands. The other four accessions from natural stands were polymorphic. Seven accessions consisted of two chlorotypes, five accessions of three chlorotypes. Three accessions were highly variable, reaching four, five and six chlorotypes, respectively. In ‘Argenmilla’, ‘Bodegold’, ‘Margaritar’ and the cultivated Croatian accession all three main lineages (‘A’, ‘B’ and ‘C’) were present (Table 3). ‘Degumille’, ‘Bulgaria1’ and ‘Soroksári 40’ consisted of two chlorotypes from the same lineage.
This high degree of cp variability within chamomile accessions was corroborated by AMOVA which allocated 66% of the variation within populations.
The Shannon Wiener Index H ranged in the polymorphic accessions from 0.41 to 1.33 (Table 4). The degree of this index is influenced by the number of chlorotypes as well as by the evenness of chlorotypes as can be seen exemplarily in the cultivated Croatian accession (MLG = 5, H = 0.95, E.5 = 0.505) and ‘Bona’ (MLG = 4, H = 1.332, E.5 = 0.922). In five polymorphic accessions the chlorotypes were evenly distributed (‘Bohemia’, ‘Bona’, ‘Bulgaria1’, ‘PNOS’, ‘Soroksári 40’, E.5 close to 1), while in ‘Argenmilla’, ‘M121’, the cultivated Croatian accession and the accession from Albania one chlorotype dominated (E.5 < 0.6). Gene diversity (overall mean = 0.285) ranged from 0.069 in ‘Bulgaria1 to 0.261 in ‘PNOS’.
Three accessions had private alleles. The accession Bulgaria1 had seven individuals with a private allele in marker mrcp02 (‘C’), responsible for chlorotype B7. ‘Argenmilla’ had two individuals with a private allele of marker mrcp66 (‘A’), responsible for chlorotype C2 and all individuals from the accession from North Korea shared the private allele in marker mrcp98 (‘T’), responsible for chlorotype B6.
155 samples of this dataset were overlapping with the mitochondria marker set published previously (Ruzicka et al. 2021; Ruzicka and Novak 2020). Both marker sets showed a very high correlation of their respective genetic distance matrices (RV2 = 0.9918) (Smilde et al. 2009).
Discussion
From NGS assemblies of several chamomile genotypes polymorphic sites could be identified in the chloroplast DNA. Markers were developed that worked well with high-resolution melting analysis (HRM). HRM is a powerful method to elegantly identify differences in DNA sequences in a one-tube/one machine system with mutation detection following immediately the PCR step in the same tube and in the same PCR machine with integrated HRM measurement module (Mader et al. 2010; Ruzicka et al. 2021; Ruzicka and Novak 2020).
Of the 31 markers selected for HRM, 5 groups of markers with complete linkage were identified finally reducing the number of non-redundant markers to 20. With the marker set 20 chlorotypes could be identified, of which three chlorotypes were based on private alleles. In the same species, 20 mitotypes (mitochondrial haplotypes) were identified with 36 SNP markers (Ruzicka et al. 2021).
The 20 chlorotypes could be easily grouped in three major chlorotype lineages, named A, B and C with chlorotypes A1 and A2 as evolutionary most ancient chlorotypes from where the three major lineages radiated from. The close relative M. discoidea formed an own chlorotype that was identical to Artemisia annua and Lactuca sativa and hence can be regarded as starting point of all evolutionary developments in M. chamomilla seen by our marker set. The increasing number of genomes facilitates the comparison to closely related species or genera to identify the ancient status of SNP variation, to ‘root’ the haplotype tree and possibly identify the geographical origin of a species as demonstrated for Cannabis sativa (Osterberger et al. 2021). A further deduction is that all mutations in M. chamomilla at the positions examined occurred after branching of the two species M. chamomilla and M. discoidea from their common ancestor.
Most of the accessions were composed of more than one chlorotype, only about one third of the accessions were monomorphic. However, this number was a bit higher than the 23% of monomorphic accessions regarding mitochondrial variation (Ruzicka et al. 2021). In the chloroplast data set presented here, a higher number of genotypes was used including more wild populations (40% in contrast to 15% in Ruzicka et al. (2021)) to support geographical information about the breeding history of cultivars.
Nevertheless, four out of the nine wild populations were polymorphic in chlorotypes, whereas only four out of 15 accessions from cultivation were monomorphic. In a breeding process many accessions are growing in the beginning side by side, and unintentional or intentional mixtures may occur leading to the high degree of polymorphisms in breeding material. Most of the cultivars of German chamomile are the product of crossings and selections of promising plant lines originally gained from natural populations. In the beginning of chamomile breeding selection was done from geographically close natural stands (Albrecht and Otto 2020).
The Albanian population showed the least differentiation from the ancient chlorotype (A1, B2, B3), possibly pointing to the Balkan Peninsula as place of origin of M. chamomilla. The differentiation into the main lineages is so clear cut that natural populations with mixed lineages most likely origin from immigration of a lineage into an existing population of different lineage. The Croatian accession (A1, A2, B3, B5, C3), although from cultivation, had probably its origin from that region. The accession from the UK (A2, A4) contradicts the hypothesis of the origin on the Balkans or has been quite a fast migration from there. Italy exhibits two main lineages, A2 in the North and C1 in the South indicating two independent colonisation events. The Southern Italian population is the only one with C1, so also close to the origin. A wild population from Central Germany (A5) shows already a higher differentiation from the origin by distance and the Alps as geographical barrier. The Hungarian population (A5, B3, B4) is probably a mix of a meeting of an immigration from West (A5) and South (B3, B4). Bulgaria would also make an interesting case (Bulgaria1 with B3 and B7 and Bulgaria2 with C5). Unfortunately, the exact geographical origin of the two samples is unknown. Also unknown is if the North Korean population (B6) was indeed spontaneous or sub-spontaneous from seed imports. The low differentiation from other B-types would rather suggest the second option. However, with the low number of wild populations and important geographic gaps (e.g., samples from the Iberian Peninsula) all deductions presented here are hypothesis rather than assured knowledge.
Amongst the cultivars, also interesting conclusions can be drawn based on chlorotypes. The Romanian ‘Margaritar’ (A5, B4), selected from the Polish cultivar ‘Zloti Lan’, and ‘PNOS’ (A5, C5) both go back to Polish landraces (Das 2014), a close relationship also seen with the mitotypes (Ruzicka et al. 2021). The close relationship found with AFLP between ‘Margaritar’ and ‘Bona’ (Wagner et al. 2005) could not be verified by the chloroplast markers.
The cultivar ‘Bohemia’ (A2, B3) goes back to wild chamomile from Bohemia and Moravia and was introduced in field production in the 1950’s (Salamon 1994). ‘Bona’ (A2, A3, A6, B3) was selected in Slovakia from 1975 onwards and registered in 1984. ‘Bona’ is a cross between ‘Bohemia’ and wild populations from Spain because of the favourable chemotype composition of the Spanish populations (Salamon 1994). Interestingly, the chlorotype of ‘Bona’ is essentially the same as ‘Bohemia’ (A2, B3) complemented with A3 and A6. Unfortunately, no Spanish wild populations were included in our study. The tetraploid ‘Goral’ (B4, C5), registered in 1990, was bred by polyploidisation of ‘Bona’ (Salamon 1994) but is unrelated with ‘Bona’ as demonstrated by chlorotypes (this work) or mitotypes (Ruzicka et al. 2021). ‘Manzana’ (pure B3) is a tetraploid chamomile variety produced by polyploidisation of the diploid ‘Degumille’ (A2, A6) developed in the former GDR (Franz and Isaac 1984). Interestingly, both cultivars are completely unrelated. Therefore, either breeding history is incorrect or further breeding in the last decades changed the chlorotype of one of the partners. ‘Bodegold’ (A2, B1) was bred in the 1950s in Quedlinburg, Germany, from wild populations of this area.
The results from the cultivars contradict partially documented breeding history, but even more so the deductions about the origin or geographical distribution from (sub-)spontaneous populations. They would rather point to an introduction of Mediterranean breeding materials into the breeding process much earlier than documented.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Agapow PM, Burt A (2001) Indices of multilocus linkage disequilibrium. Mol Ecol Notes 1:101–102
Ahmadi H, Rahimmalek M, Zeinali H (2014) Assessment of the genetic variation of chamomile (Matricaria chamomilla L.) populations using phytochemical, morphological and ISSR markers. Biochem Syst Ecol 54:190–197. https://doi.org/10.1016/j.bse.2014.01.010
Albrecht S, Otto L-G (2020) Matricaria recutita L.: true chamomile. In: Novak J, Blüthner W-D (eds) Medicinal, aromatic and stimulant plants, vol 12, 1st edn. Springer, Cham, pp 313–331
Avise JC, Arnold J, Ball RM, Bermingham E, Lamb T, Neigel JE, Reeb CA, Saunders NC (1987) Intraspecific phylogeography: the mitochondrial DNA bridge between population genetics and systematics. Annu Rev Ecol Syst 18:489–522. https://doi.org/10.1146/annurev.es.18.110187.002421
Brown A, Feldman MW, Nevo E (1980) Multilocus structure of natural populations of Hordeum spontaneum. Genetics 96:523–536
Chatzidimopoulos M, Ganopoulos I, Moraitou-Daponta E, Lioliopoulou F, Ntantali O, Panagiotaki P, Vellios EK (2019) High-resolution melting (HRM) analysis reveals genotypic differentiation of Venturia inaequalis populations in Greece. Front Ecol Evol. https://doi.org/10.3389/fevo.2019.00489
Das M (2014) Chamomile. CRC Press
Franke R, Hannig HJ (2012) Kamille (Matricaria recutita L.). In: Hoppe B (ed) Arznei- und Gewürzpflanzen A-K, vol 4. Saluplanta, Bernburg, pp 618–648
Franke R, Schilcher H (2007) Relevance and use of Chamomile (Matricaria recutita L.). Acta Hortic 25:29–43. https://doi.org/10.17660/ActaHortic.2007.749.2
Franz C, Isaac O (1984) Verfahren zur Herstellung der neuen Kamillensorte Manzana: Erfindungspatent für die Schweiz und Liechtenstein (CH 667 180 A5)
Glücknerova E, Blazek Z, Stary F (1965) Charakteristika Ceskoslovensk’ych Povolen’ych Odrud He RM’anku Prav’eho (Matricaria chamomilla L.). Cesk Farm 14:105–112
Indahl UG, Naes T, Liland KH (2018) A similarity index for comparing coupled matrices. J Chemom 32:e3049. https://doi.org/10.1002/cem.3049
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2013:1–14. https://doi.org/10.7717/peerj.281
Kamvar ZN, Brooks JC, Grünwald NJ (2015) Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Front Genet 6:208. https://doi.org/10.3389/fgene.2015.00208
Mader E, Lohwasser U, Börner A, Novak J (2010) Population structures of genebank accessions of Salvia officinalis L. (Lamiaceae) revealed by high resolution melting analysis. Biochem Syst Ecol 38:178–186. https://doi.org/10.1016/j.bse.2010.01.001
Mehdikhani H, Soolooki M, Zeinali H (2014) Study of genetic diversity in several scentless chamomile landraces (Matricaria inodora L.) based on morphological traits and RAPD molecular markers. Iran J Rangel Forests Plant Breed Genet Res 21:242–256. https://doi.org/10.22092/ijrfpbgr.2014.5001
Mežaka I, Kronberga A, Nakurte I, Taškova I, Jakovels D, Primavera A (2020) Genetic, chemical and morphological variability of chamomile (Chamomilla recutita L.) populations of Latvia. Ind Crops Prod 154:112614. https://doi.org/10.1016/j.indcrop.2020.112614
Okoń S, Surmacz-Magdziak A, Paczos-Grzęda E (2013) Genetic diversity among cultivated and wild chamomile germplasm based on ISSR analysis. asphc 12
Osterberger E, Lohwasser U, Jovanovic D, Ruzicka J, Novak J (2021) The origin of the genus Cannabis. Genet Resour Crop Evol. https://doi.org/10.1007/s10722-021-01309-y
Otto L-G, Mondal P, Brassac J, Preiss S, Degenhardt J, He S, Reif JC, Sharbel TF (2017) Use of genotyping-by-sequencing to determine the genetic structure in the medicinal plant chamomile, and to identify flowering time and alpha-bisabolol associated SNP-loci by genome-wide association mapping. BMC Genom 18:599. https://doi.org/10.1186/s12864-017-3991-0
Pirkhezri M, Hassani ME, Hadian J (2010) Genetic Diversity in different populations of Matricaria chamomilla L. growing in southwest of iran, based on morphological and RAPD markers. Res J Med Plant 4:1–13. https://doi.org/10.3923/rjmp.2010.1.13
Repčák M, Švehlı́ková V, Imrich J, Pihlaja K (1999) Jaceidin and chrysosplenetin chemotypes of Chamomilla recutita (L.) Rauschert. Biochem Syst Ecol 27:727–732. https://doi.org/10.1016/S0305-1978(98)00124-0
Ruzicka J, Novak J (2020) Mitochondrial genome variation between different accessions of Matricaria chamomilla L. (Asteraceae) based on SNP mutation analysis. Genet Resour Crop Evol 67:853–864. https://doi.org/10.1007/s10722-020-00881-z
Ruzicka J, Hacek M, Novak J (2021) Mitochondrial relationships between various chamomile accessions. J Appl Genet 62:73–84. https://doi.org/10.1007/s13353-020-00602-3
Salamon I (1994) Growing condition and breeding of chamomile (Chamomilla recutita (L.) Rauschert) regarding the essential oil qualitative-quantitative characteristics in Slovakia. Herba Polonica 40:68–74
Schilcher H, Imming P, Goeters S (2005) Pharmacology and toxicology. In: Franke R, Schilcher H (eds) Chamomile. CRC Press, pp 245–264
Schmiderer C, Lukas B, Novak J (2013) Der Einfluss verschiedener DNS-Extraktionsmethoden und Verdünnungen auf den Amplifikationserfolg in der PCR von verschiedenen Arznei- und Gewürzpflanzen. Zeitschrift Für Azrnei- Und Gewürzpflanzen 18:65–72
Shaal BA, Hayworth DA, Rauscher JT, Smith WA (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7:465–474. https://doi.org/10.1046/j.1365-294x.1998.00318.x
Smilde AK, Kiers HAL, Bijlsma S, Rubingh CM, van Erk MJ (2009) Matrix correlations for high-dimensional data: the modified RV-coefficient. Bioinformatics 25:401–405. https://doi.org/10.1093/bioinformatics/btn634
Solouki M, Mehdikhani H, Zeinali H, Emamjomeh AA (2008) Study of genetic diversity in Chamomile (Matricaria chamomilla) based on morphological traits and molecular markers. Sci Hortic 117:281–287. https://doi.org/10.1016/j.scienta.2008.03.029
Timmis JN, Ayliffe MA (1990) Extranuclear DNA and its use in systematics. Aust Syst Bot 3:137. https://doi.org/10.1071/SB9900137
Wagner C, Friedt W, Marquard RA, Ordon F (2005) Molecular analyses on the genetic diversity and inheritance of (−)-α-bisabolol and chamazulene content in tetraploid chamomile (Chamomilla recutita (L.) Rausch.). Plant Sci 169:917–927. https://doi.org/10.1016/j.plantsci.2005.06.010
WFO Matricaria chamomilla L. http://www.worldfloraonline.org/taxon/wfo-0000003146. Accessed 16 February 2022
Acknowledgements
This work was initiated by a collaboration project with the Leibniz Institute of Plant Genetics and Crop Plant Research (IPK), Department Plant Breeding Research, Apomixis Research Group, Gatersleben, Germany and the Pharmaplant GmbH, Artern, Germany. The authors would like to thank Lars-Gernot Otto (IPK) for his advice in the project and Marlis Sonnenschein, Bartholome Plocharski (Pharmaplant GmbH, Germany), Eva Nemeth (University of Agriculture and Life Sciences, Budapest, Hungary), Alban Ibraliu (Agricultural University, Tirana, Albania), Richard Pickering (UK) and Ina Bein-Lobmaier (University of Veterinary Medicine Vienna, Austria) for providing seed material.
Funding
Open access funding provided by University of Veterinary Medicine Vienna. This work was financed based upon a resolution of the Federal Parliament of Germany (Deutscher Bundestag) by the Bundesministerium für Ernährung und Landwirtschaft (BMEL, support code 14NR063) via the Fachagentur Nachwachsende Rohstoffe e.V. (FNR) as project executing organisation for the funding programme ‘Renewable Resources’. The funding organisation did not execute any influence on the study nor the manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Laboratory work was conducted by JR, GB and DJ. Statistical evaluations by JR, GB and JN. The manuscript was written by JN and all authors commented on the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ruzicka, J., Baumschlager, G., Jovanovic, D. et al. Three major chlorotype lineages in chamomile (Matricaria chamomilla L., Asteraceae). Genet Resour Crop Evol 71, 331–340 (2024). https://doi.org/10.1007/s10722-023-01625-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01625-5