Abstract
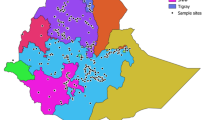
Knowing the extent of genetic variability among and within barley accessions provide bases of selection for breeders. However, given the heterogeneous nature of the germplasm, genetic information about barley landraces stored in gene bank of Ethiopia is not well archived. Therefore, genetic diversity study was conducted among and within 20 barley accessions represented by 194 barley plants. The accessions originate from 10 different geographic regions of Ethiopia covering a wide range of altitudes. The accessions were tested using 15 simple sequence repeat markers (SSRs). The SSR markers detected 57 alleles ranging from two (GBM1042) to seven (Bmac0040) alleles per marker indicating the genetic diversity in the barley plants. The allelic diversity across geographic regions and altitude reveals that barley accessions collected from mid and low altitudes of Ethiopia show high allelic diversity compared to accessions originating from high altitudes. Analysis of molecular variance results based on geographic regions (64.39%) and altitude classes (64.04%) also showed the highest variation within accessions compared to among accessions and groups. Both geographic regions and altitude-based genetic structure analysis detected two sub-populations. The individual assignment following the altitude of collection revealed the role of altitude in shaping the genetic diversity observed in barley populations. In general, the result of this study shows that each single barley line within the accessions should be considered as a potential source of genetic material and must be treated distinctly during germplasm collection and utilization.





Similar content being viewed by others
References
Abebe TD (2010) Genetic diversity and population differentiation analysis of Ethiopian barley (Hordeum vulgare L.) landraces using morphological traits and SSR markers. Dissertation, University Bonn
Abebe TD, Leon J (2013) Spatial and temporal genetic analyses of Ethiopian barley (Hordeum vulgare L.) landraces reveal the absence of a distinct population structure. Genet Resour Crop Evol 60:1547–1558. https://doi.org/10.1007/s10722-012-9941-4
Abebe TD, Bauer AM, Leon J (2010) Morphological diversity of Ethiopian barley (Hordeum vulgare L.) in relation to geographic regions and altitudes. Hereditas 147:154–164. https://doi.org/10.1111/j.1601-5223.2010.02173.x
Abebe TD, Mathew B, Leon J (2013) Barrier analysis detected genetic discontinuity among Ethiopian barley (Hordeum vulgare L.) landraces due to landscape and human mobility on gene flow. Genet Resour Crop Evol 60:297–309. https://doi.org/10.1007/s10722-012-9834-6
Abebe TD, Naz AA, Leon J (2015) Landscape genomics reveal signatures of local adaptation in barley (Hordeum vulgare L.). Front Plant Sci 6:813. https://doi.org/10.3389/fpls.2015.00813
Adugna A (2008) Assessment of yield stability in sorghum using univariate and multivariate statistical approaches. Hereditas 145:28–37. https://doi.org/10.1111/j.0018-0661.2008.2023.x
Afolayan G, Deshpande SP, Aladele SE, Kolawole AO, Angarawai I, Nwosu DJ, Michael C, Blay ET, Danquah EY (2019) Genetic diversity assessment of sorghum (Sorghum bicolor (L.) Moench) accessions using single nucleotide polymorphism markers. Plant Genet Resour 17(5):412–420. https://doi.org/10.1017/S1479262119000212
Anderson J, Churchill G, Autrique J, Tanksley S, Sorrells M (1993) Optimizing parental selection for genetic linkage maps. Genome 36:181–186. https://doi.org/10.1139/g93-024
Asfaw Z (1988) Variation in the morphology of the spike within Ethiopian barley, Hordeum vulgare L. (Poaceae). Acta Agric Scand B 38:277–288. https://doi.org/10.1080/00015128809437989
Asfaw Z (2000) The Barleys of Ethiopia. In: Brush SB (ed) Genes in the field: on-farm conservation of crop diversity. IDRC, Ottawa, pp 77–107
Bellucci E, Bitocchi E, Rau D, Nanni L, Ferradini N, Giardini A, Rodriguez M, Attene G, Papa R (2013) Population structure of barley landrace populations and gene-flow with modern varieties. PLoS ONE 8(12):e83891. https://doi.org/10.1371/journal.pone.0083891
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphism. Am J Hum Genet 32:314–331
Brown AHD (2000) The genetic structure of crop landraces and the challenge to conserve them in situ on farms. In: Brush SB (ed) Genes in the field: on-farm conservation of crop diversity. IDRC, Ottawa, pp 29–48
Caccone A (1985) Gene flow in cave arthropods: a qualitative and quantitative approach. Evolution 39:1223–1235. https://doi.org/10.2307/2408779
Cavatassi R, Lipper L, Hopkins J (2006) The role of crop genetic diversity in coping with agricultural production shocks: insights from Eastern Ethiopia. Agricultural development economics division, working paper no. 06-17. Food and Agriculture Organization of the United Nations, Rome, Italy. https://doi.org/10.22004/ag.econ.289050
Ceccarelli S (1996) Positive interpretation of genotype by environment interactions in relation to sustainability and biodiversity. In: Cooper M, Hammer GL (eds) Plant adaptation and crop improvement. CABI International, Manila, pp 467–486
Ceccarelli S, Grando S (2000) Barley landraces from the Fertile Crescent: a lesson for plant breeders. In: Brush SB (ed) Genes in the field: on-farm conservation of crop diversity. IDRC, Ottawa, pp 51–76
Climate Risk Profile: Ethiopia (2020) The World Bank Group
Collins NC, Paltridge NG, Ford CM, Symons RH (1996) The Yd2 gene for barley yellowdwarf virus resistance maps close to the centromere on the long arm of barley chromosome 3. Theor Appl Genet 92:858–864. https://doi.org/10.1007/BF00221898
Demissie A, Bjernstad A, Kleinhofs A (1998) Restriction fragment length polymorphisms in landrace barleys from Ethiopia in relation to geographic, altitude, and agro-ecological factors. Crop Sci 38(1):237–243. https://doi.org/10.2135/cropsci1998.0011183X003800010040x
Dido AA, Singh BJK, Ermias A, Krishna MSR, Dawit D, Kassahun T (2022) Genetic diversity, population structure and relationship of Ethiopian barley (Hordeum vulgare L.) landraces as reveled by simple sequence repeat (SSR) markers. J Genet 101:9. https://doi.org/10.1007/s12041-021-01346-7
Dziurdziak J, Gryziak G, Groszyk J, Podyma W, Boczkowska M (2021) DArTseq genotypic and phenotypic diversity of barley landraces originating from different countries. Agronomy 11(11):2330. https://doi.org/10.3390/agronomy11112330
Earl DA, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Engels JMM (1994) Genetic diversity in Ethiopian barley in relation to altitude. Genet Resour Crop Evol 42:761–768. https://doi.org/10.1007/BF00053050
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587. https://doi.org/10.1093/genetics/164.4.1567
FASOSTAT (2021) Crops and livestock products. http://www.fao.org/faostat/en/#data/QCL . Retrieved the data on Ocober 2021
Ge C, Wentzel E, D’souza N, Chen K, Oliver RP, Ellwood SR (2021) Adult resistance genes to barley powdery mildew confer basal penetration resistance associated with broad-spectrum resistance. Plant Genome 14(3):e20129. https://doi.org/10.1002/tpg2.20129
Global Crop Diversity Trust (2008) Global strategy for the ex situ conservation and use of barley germplasm [online]. https://www.croptrust.org/wp/wp-content/uploads/2014/12/Barley_Strategy_FINAL_27Oct08.pdf. Accessed 02 Oct 2021
Hartl D, Clark A (1997) Population sub-strucutre. In: Hartel D, Clark A (eds) Principles of population genetics. Sinauer Associates, Sunderland, pp 111–161
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. https://doi.org/10.1093/bioinformatics/btm233
Jørgensen JH (1992) Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 63:141–152. https://doi.org/10.1007/BF00023919
Lakew B, Assefa A (2011) Advances and experiences in barley landrace improvement in Ethiopia. In: Mulatu B, Grando S (eds) Barley research and development in Ethiopia. Proceedings of the 2nd National Barley research and development review workshop. 28–30 November 2006, HARC, Holetta, Ethiopia. ICARDA, PO Box 5466, Aleppo, Syria, pp xiv + 391
Lakew B, Gebre H, Alemayehu F (1996) Barley production and research. In: Gebre H, Leur JV (eds) Barley research in Ethiopia: past work and future prospects. Proceeding of the first Barley resaerch review workshop. IAR/ICARDA, Addis Ababa
Marame F, Desalegne L, Fininsa C, Sigvald R (2009) Genetic analysis for some plant and fruit traits, and its implication for a breeding program of hot pepper (Capsicum annuum var. annuum L.). Hereditas 146:131–140. https://doi.org/10.1111/j.1601-5223.2009.02101.x
Marone D, Russo MA, Mores A, Ficco DBM, Laidò G, Mastrangelo AM, Borrelli GM (2021) Importance of landraces in cereal breeding for stress tolerance. Plants 10:1267. https://doi.org/10.3390/plants10071267
Mengistu D, Kidane Y, Fadda C, Pè M (2018) Genetic diversity in Ethiopian Durum Wheat (Triticum turgidum var durum) inferred from phenotypic variations. Plant Genet Resour 16(1):39–49. https://doi.org/10.1017/S1479262116000393
Munck L, Karlsson KE, Hagberg A, Eggum BO (1970) Gene for Improved nutritional value in barley seed protein. Science 168:985–987
Ould Med Mahmoud M, Hamza S (2009) Genetic diversity in local barley accessions collected from different geographical regions of Tunisia. Plant Genet Resour 7(2):169–176. https://doi.org/10.1017/S1479262108162086
Peakall R, Smouse PE (2012) GENALEX 6: genetic analysis in Excel. Population genetics software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1093/bioinformatics/bts460
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1534/genetics.116.195164
Qualset CO (1975) Sampling germplasm in a centre of diversity: an example of disease resistance in Ethiopian barley. In: Frankel OH, Hawkes JG (eds) Crop genetic resources for today and tomorrow. Cambridge University Press, Cambridge, pp 81–96
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci 81:8014–8018. https://doi.org/10.1073/pnas.81.24.8014
Samberg LH, Fishman L, Allendorf FW (2013) Population genetic structure in a social landscape: barley in a traditional Ethiopian agricultural system. Evol Appl 6:1133–1145. https://doi.org/10.1111/eva.12091
Slatkin M (1981) Estimating levels of gene flow in natural populations. Genetics 99:323–335. https://doi.org/10.1093/genetics/99.2.323
Slatkin M (1993) Isolation by distance in equilibrium and nonequilibrium populations. Evolution 47:264–279. https://doi.org/10.2307/2410134
Soriano JM, Villegas D, Sorrells ME, Royo C (2018) Durum wheat landraces from east and west regions of the mediterranean basin are genetically distinct for yield components and phenology. Front Plant Sci 9:80. https://doi.org/10.3389/fpls.2018.00080
Tadesse D, Derso B (2019) The status and constraints of food barley production in the North Gondar highlands, North Western Ethiopia. Agric Food Secur 8(1):1–7. https://doi.org/10.1186/s40066-018-0248-3
Tanto HT, Rau D, Bitocchi E, Papa R (2010) Adaptation and diversity along an altitudinal gradient in Ethiopian barley (Hordeum vulgare L.) landraces revealed by molecular analysis. BMC Plant Biol 10:121. https://doi.org/10.1186/1471-2229-10-121
USDA (2012) http://www.fas.usda.gov/pecad2/highlights/2002/10/ethiopia/baseline/Eth_Crop_Production.htm Accessed 14 Nov 2012
Vavilov NI (1951) Phytogeographic basis of plant breeding. In: Verdooran F (ed) The origin, variation, Immunity and breeding cultivated plants, vol 13, no 1/6. Chron Bot, pp 13–54
Waples R (1987) A multispecies approach to the analysis of a gene flow in marine shore fishes. Evolution 41:385–400. https://doi.org/10.2307/2409146
Weir BS (1996) Diversity. In: Weir BS (ed) Genetic data analysis II: methods for discrete population genetic data. Sinauer Associates, Sunderland, pp 141–160
Wiberg A (1974) Sources of resistance to powdery mildew in barley. Hereditas 78:1–40. https://doi.org/10.1111/j.1601-5223.1974.tb01426.x
Wright S (1978) Genetic variability in natural populations: methods. In: Wright S (ed) Evolution and the genetics of population: variability within and among natural populations. The University of Chicago Press, Chicago, pp 79–103
Yitbarek S, Berhane L, Fikadu A, van Leur JAG, Grando S, Ceccarelli S (1998) Variation in Ethiopian barley landrace populations for resistance to barley leaf scald and net blotch. Plant Breed 117:419–423. https://doi.org/10.1111/j.1439-0523.1998.tb01966.x
Zohary D, Hopf M, Weiss E (2012) Cereals. In: Zohary D, Hopf M, Weiss E (eds) Domestication of plants in the old world. Oxford Univeristy Press, New York, pp 20–74
Funding
The laboratory research was funded by Faculty of Agriculture, University of Bonn, Germany.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by TA, AA, and JL The first draft of the manuscript was written by TA and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abebe, T.D., Abate, A. & Leon, J. Genetic diversity within landraces of barley (Hordeum vulgare L.) and its implications on germplasm collection and utilization. Genet Resour Crop Evol 70, 1985–1998 (2023). https://doi.org/10.1007/s10722-023-01549-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01549-0